Abstract
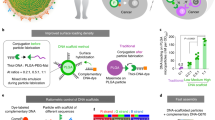
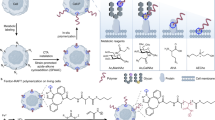
The biofunctionalization of synthetic materials has extensive utility for biomedical applications, but approaches to bioconjugation typically show insufficient efficiency and controllability. We recently developed an approach by building synthetic DNA scaffolds on biomaterial surfaces that enables the precise control of cargo density and ratio, thus improving the assembly and organization of functional cargos. We used this approach to show that the modulation and phenotypic adaptation of immune cells can be regulated using our precisely functionalized biomaterials. Here, we describe the three key procedures, including the fabrication of polymeric particles engrafted with short DNA scaffolds, the attachment of functional cargos with complementary DNA strands, and the surface assembly control and quantification. We also explain the critical checkpoints needed to ensure the overall quality and expected characteristics of the biological product. We provide additional experimental design considerations for modifying the approach by varying the material composition, size or cargo types. As an example, we cover the use of the protocol for human primary T cell activation and for the identification of parameters that affect ex vivo T cell manufacturing. The protocol requires users with diverse expertise ranging from synthetic materials to bioconjugation chemistry to immunology. The fabrication procedures and validation assays to design high-fidelity DNA-scaffolded biomaterials typically require 8 d.
Key points
-
The protocol describes the fabrication of DNA scaffolds, the bioconjugation of biomolecules with complementary DNAs, conjugate assembly onto the DNA scaffolds and their immunomodulatory effect on primary human T cells in culture.
-
Steric hindrance typically limits the use of orthogonal chemistry and covalent surface attachment strategies, whereas this DNA hybridization-based approach maintains control over the loading of each biomolecule species.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Any raw data that supports the plots within this paper are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Mitchell, M. J. et al. Engineering precision nanoparticles for drug delivery. Nat. Rev. Drug Discov. 20, 101–124 (2021).
Blanco, E., Shen, H. & Ferrari, M. Principles of nanoparticle design for overcoming biological barriers to drug delivery. Nat. Biotechnol. 33, 941–951 (2015).
Dellacherie, M. O., Seo, B. R. & Mooney, D. J. Macroscale biomaterials strategies for local immunomodulation. Nat. Rev. Mater. 4, 379–397 (2019).
Wang, H. & Mooney, D. J. Biomaterial-assisted targeted modulation of immune cells in cancer treatment. Nat. Mater. 17, 761–772 (2018).
Waldman, A. D., Fritz, J. M. & Lenardo, M. J. A guide to cancer immunotherapy: from T cell basic science to clinical practice. Nat. Rev. Immunol. 20, 651–668 (2020).
Tan, S., Li, D. & Zhu, X. Cancer immunotherapy: pros, cons and beyond. Biomed. Pharmacother. 124, 109821 (2020).
Sterner, R. C. & Sterner, R. M. CAR-T cell therapy: current limitations and potential strategies. Blood Cancer J. 11, 1–11 (2021).
Romano, M., Fanelli, G., Albany, C. J., Giganti, G. & Lombardi, G. Past, present, and future of regulatory T cell therapy in transplantation and autoimmunity. Front. Immunol. 10, 43 (2019).
Ferreira, L. M. R., Muller, Y. D., Bluestone, J. A. & Tang, Q. Next-generation regulatory T cell therapy. Nat. Rev. Drug Discov. 18, 749–769 (2019).
Herold, K. C. et al. An anti-CD3 antibody, teplizumab, in relatives at risk for type 1 diabetes. N. Engl. J. Med. 381, 603–613 (2019).
Balcerek, J. et al. Polyclonal regulatory T cell manufacturing under cGMP: a decade of experience. Front. Immunol. 12, 744763 (2021).
Dong, S. et al. The effect of low-dose IL-2 and Treg adoptive cell therapy in patients with type 1 diabetes. JCI Insight 6, e147474 (2021).
Lin, M. J. et al. Cancer vaccines: the next immunotherapy frontier. Nat. Cancer 3, 911–926 (2022).
Larson, R. C. & Maus, M. V. Recent advances and discoveries in the mechanisms and functions of CAR T cells. Nat. Rev. Cancer 21, 145–161 (2021).
Tang, L. et al. Enhancing T cell therapy through TCR-signaling-responsive nanoparticle drug delivery. Nat. Biotechnol. 36, 707–716 (2018).
Bonati, L. & Tang, L. Cytokine engineering for targeted cancer immunotherapy. Curr. Opin. Chem. Biol. 62, 43–52 (2021).
Hwang, J.-R., Byeon, Y., Kim, D. & Park, S.-G. Recent insights of T cell receptor-mediated signaling pathways for T cell activation and development. Exp. Mol. Med. 52, 750–761 (2020).
Veerman, R. E., Güçlüler Akpinar, G., Eldh, M. & Gabrielsson, S. Immune cell-derived extracellular vesicles—functions and therapeutic applications. Trends Mol. Med. 25, 382–394 (2019).
Chang, J. T., Wherry, E. J. & Goldrath, A. W. Molecular regulation of effector and memory T cell differentiation. Nat. Immunol. 15, 1104–1115 (2014).
Huang, X. et al. DNA scaffolds enable efficient and tunable functionalization of biomaterials for immune cell modulation. Nat. Nanotechnol. 16, 214–223 (2021).
Majedi, F. S. et al. Augmentation of T-cell activation by oscillatory forces and engineered antigen-presenting cells. Nano Lett. 19, 6945–6954 (2019).
Cheung, A. S., Zhang, D. K. Y., Koshy, S. T. & Mooney, D. J. Scaffolds that mimic antigen-presenting cells enable ex vivo expansion of primary T cells. Nat. Biotechnol. 36, 160–169 (2018).
Rhodes, K. R., Meyer, R. A., Wang, J., Tzeng, S. Y. & Green, J. J. Biomimetic tolerogenic artificial antigen presenting cells for regulatory T cell induction. Acta Biomater. 112, 136–148 (2020).
Kim, J. V., Latouche, J.-B., Rivière, I. & Sadelain, M. The ABCs of artificial antigen presentation. Nat. Biotechnol. 22, 403–410 (2004).
Boozer, C., Ladd, J., Chen, S. & Jiang, S. DNA-directed protein immobilization for simultaneous detection of multiple analytes by surface plasmon resonance biosensor. Anal. Chem. 78, 1515–1519 (2006).
Liu, Y. & Yu, J. Oriented immobilization of proteins on solid supports for use in biosensors and biochips: a review. Microchim. Acta 183, 1–19 (2016).
Bilal, M., Asgher, M., Cheng, H., Yan, Y. & Iqbal, H. M. N. Multi-point enzyme immobilization, surface chemistry, and novel platforms: a paradigm shift in biocatalyst design. Crit. Rev. Biotechnol. 39, 202–219 (2019).
Smith, M. R., Tolbert, S. V. & Wen, F. Protein-scaffold directed nanoscale assembly of T cell ligands: Artificial antigen presentation with defined valency, density, and ratio. ACS Synth. Biol. 7, 1629–1639 (2018).
Wang, X. & Rivière, I. Clinical manufacturing of CAR T cells: foundation of a promising therapy. Mol. Ther. Oncolytics 3, 16015 (2016).
Arcangeli, S. et al. CAR T cell manufacturing from naive/stem memory T lymphocytes enhances antitumor responses while curtailing cytokine release syndrome. J. Clin. Invest. 132, e150807 (2022).
López-Cantillo, G., Urueña, C., Camacho, B. A. & Ramírez-Segura, C. CAR-T cell performance: how to improve their persistence? Front. Immunol. 13, 878209 (2022).
Arcangeli, S. et al. Next-generation manufacturing protocols enriching TSCM CAR T cells can overcome disease-specific T cell defects in cancer patients. Front. Immunol. 11, 1217 (2020).
Elmowafy, E. M., Tiboni, M. & Soliman, M. E. Biocompatibility, biodegradation and biomedical applications of poly(lactic acid)/poly(lactic-co-glycolic acid) micro and nanoparticles. J. Pharm. Investig. 49, 347–380 (2019).
Roybal, K. T. et al. Precision tumor recognition by T cells with combinatorial antigen-sensing circuits. Cell 164, 770–779 (2016).
Maynard, S. A., Winter, C. W., Cunnane, E. M. & Stevens, M. M. Advancing cell-instructive biomaterials through increased understanding of cell receptor spacing and material surface functionalization. Regen. Eng. Transl. Med. 7, 533–547 (2021).
Zhong, J. X., Raghavan, P. & Desai, T. A. Harnessing biomaterials for immunomodulatory-driven tissue engineering. Regen. Eng. Transl. Med. https://doi.org/10.1007/s40883-022-00279-6 (2022).
Mertgen, A.-S. et al. Multifunctional biomaterials: combining material modification strategies for engineering of cell-contacting surfaces. ACS Appl. Mater. Interfaces 12, 21342–21367 (2020).
Bao, G., Mitragotri, S. & Tong, S. Multifunctional nanoparticles for drug delivery and molecular imaging. Annu. Rev. Biomed. Eng. 15, 253–282 (2013).
Saminathan, A., Zajac, M., Anees, P. & Krishnan, Y. Organelle-level precision with next-generation targeting technologies. Nat. Rev. Mater. 7, 355–371 (2022).
Lagreca, E. et al. Recent advances in the formulation of PLGA microparticles for controlled drug delivery. Prog. Biomater. 9, 153–174 (2020).
Yu, W., Liu, R., Zhou, Y. & Gao, H. Size-tunable strategies for a tumor targeted drug delivery system. ACS Cent. Sci. 6, 100–116 (2020).
Pradal, J. et al. Effect of particle size on the biodistribution of nano- and microparticles following intra-articular injection in mice. Int. J. Pharm. 498, 119–129 (2016).
Adakkattil, R., Thakur, K. & Rai, V. Reactivity and selectivity principles in native protein bioconjugation. Chem. Rec. 21, 1941–1956 (2021).
Trads, J. B., Tørring, T. & Gothelf, K. V. Site-selective conjugation of native proteins with DNA. Acc. Chem. Res. 50, 1367–1374 (2017).
Saha, B., Songe, P., Evers, T. H. & Prins, M. W. J. The influence of covalent immobilization conditions on antibody accessibility on nanoparticles. Analyst 142, 4247–4256 (2017).
Zamecnik, C. R., Lowe, M. M., Patterson, D. M., Rosenblum, M. D. & Desai, T. A. Injectable polymeric cytokine-binding nanowires are effective tissue-specific immunomodulators. ACS Nano 11, 11433–11440 (2017).
Makaraviciute, A., Jackson, C. D., Millner, P. A. & Ramanaviciene, A. Considerations in producing preferentially reduced half-antibody fragments. J. Immunol. Methods 429, 50–56 (2016).
Sapsford, K. E. et al. Functionalizing nanoparticles with biological molecules: developing chemistries that facilitate nanotechnology. Chem. Rev. 113, 1904–2074 (2013).
Khandare, J. & Minko, T. Polymer–drug conjugates: progress in polymeric prodrugs. Prog. Polym. Sci. 31, 359–397 (2006).
Martínez-Jothar, L. et al. Insights into maleimide-thiol conjugation chemistry: conditions for efficient surface functionalization of nanoparticles for receptor targeting. J. Control. Release 282, 101–109 (2018).
Chiodi, E., Marn, A. M., Geib, M. T. & Ünlü, M. S. The role of surface chemistry in the efficacy of protein and DNA microarrays for label-free detection: an overview. Polymers 13, 1026 (2021).
Pei, X. et al. Putting precision and elegance in enzyme immobilisation with bio-orthogonal chemistry. Chem. Soc. Rev. 51, 7281–7304 (2022).
Wasserberg, D., Cabanas-Danés, J., Subramaniam, V., Huskens, J. & Jonkheijm, P. Orthogonal supramolecular protein assembly on patterned bifunctional surfaces. Chem. Commun. 54, 1615–1618 (2018).
Meder, F., Kaur, S., Treccani, L. & Rezwan, K. Controlling mixed-protein adsorption layers on colloidal alumina particles by tailoring carboxyl and hydroxyl surface group densities. Langmuir 29, 12502–12510 (2013).
Wongrakpanich, A., Khunkitchai, N., Achayawat, Y. & Suksiriworapong, J. Ketorolac-loaded PLGA-/PLA-based microparticles stabilized by hyaluronic acid: effects of formulation composition and emulsification technique on particle characteristics and drug release behaviors. Polymers 15, 266 (2023).
Makadia, H. K. & Siegel, S. J. Poly lactic-co-glycolic acid (PLGA) as biodegradable controlled drug delivery carrier. Polymers 3, 1377–1397 (2011).
Vivek, K., Harivardhan Reddy, L. & Murthy, R. S. R. Comparative study of some biodegradable polymers on the entrapment efficiency and release behavior of etoposide from microspheres. Pharm. Dev. Technol. 12, 79–88 (2007).
Ghasemiyeh, P. & Mohammadi-Samani, S. Polymers blending as release modulating tool in drug delivery. Front. Mater. 8, 752813 (2021).
Fu, J. et al. DNA-scaffolded proximity assembly and confinement of multienzyme reactions. Top. Curr. Chem. 378, 38 (2020).
Wiener, J., Kokotek, D., Rosowski, S., Lickert, H. & Meier, M. Preparation of single- and double-oligonucleotide antibody conjugates and their application for protein analytics. Sci. Rep. 10, 1457 (2020).
van der Sleen, L. M. & Tych, K. M. Bioconjugation strategies for connecting proteins to DNA-linkers for single-molecule force-based experiments. Nanomaterials 11, 2424 (2021).
von Witting, E., Hober, S. & Kanje, S. Affinity-based methods for site-specific conjugation of antibodies. Bioconjugate Chem. 32, 1515–1524 (2021).
Manning, M. C., Chou, D. K., Murphy, B. M., Payne, R. W. & Katayama, D. S. Stability of protein pharmaceuticals: an update. Pharm. Res. 27, 544–575 (2010).
Frokjaer, S. & Otzen, D. E. Protein drug stability: a formulation challenge. Nat. Rev. Drug Discov. 4, 298–306 (2005).
Wang, W. Protein aggregation and its inhibition in biopharmaceutics. Int. J. Pharm. 289, 1–30 (2005).
Themeli, M., Rivière, I. & Sadelain, M. New cell sources for T cell engineering and adoptive immunotherapy. Cell Stem Cell 16, 357–366 (2015).
Stock, S., Schmitt, M. & Sellner, L. Optimizing manufacturing protocols of chimeric antigen receptor T cells for improved anticancer immunotherapy. Int. J. Mol. Sci. 20, 6223 (2019).
Eskandari, S. K. et al. Regulatory T cells engineered with TCR signaling–responsive IL-2 nanogels suppress alloimmunity in sites of antigen encounter. Sci. Transl. Med. 12, eaaw4744 (2020).
Ghaffari, S. et al. Optimizing interleukin-2 concentration, seeding density and bead-to-cell ratio of T-cell expansion for adoptive immunotherapy. BMC Immunol. 22, 43 (2021).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Wang, Y. et al. Phase transitions in human IgG solutions. J. Chem. Phys. 139, 121904 (2013).
Wang, S. S., Yan, Y. S. & Ho, K. US FDA-approved therapeutic antibodies with high-concentration formulation: summaries and perspectives. Antib. Ther. 4, 262–272 (2021).
Vincent, M. P., Navidzadeh, J. O., Bobbala, S. & Scott, E. A. Leveraging self-assembled nanobiomaterials for improved cancer immunotherapy. Cancer Cell 40, 255–276 (2022).
Danaei, M. et al. Impact of particle size and polydispersity index on the clinical applications of lipidic nanocarrier systems. Pharmaceutics 10, 57 (2018).
Fass, D. & Thorpe, C. Chemistry and enzymology of disulfide cross-linking in proteins. Chem. Rev. 118, 1169–1198 (2018).
Delcassian, D., Sattler, S. & Dunlop, I. E. T cell immunoengineering with advanced biomaterials. Integr. Biol. 9, 211–222 (2017).
Wang, C., Sun, W., Ye, Y., Bomba, H. N. & Gu, Z. Bioengineering of artificial antigen presenting cells and lymphoid organs. Theranostics 7, 3504–3516 (2017).
Si, X., Xiao, L., Brown, C. E. & Wang, D. Preclinical evaluation of CAR T cell function: in vitro and in vivo models. Int. J. Mol. Sci. 23, 3154 (2022).
Acknowledgements
The authors thank Z. Gartner and B. Moser for relevant DNA synthesis, S. Douglas and K. Shen for gel imaging, W. Lim for sonication equipment, the University of California, San Francisco (UCSF) Nikon Imaging center for confocal microscopy, and V. Nguyen and the UCSF Flow Cytometry Core (RRID: SCR_018206) for their flow cytometry expertise and equipment use—funded in part by the Diabetes Research Center via the National Institutes of Health (NIH) grant P30 DK063720. P.H. was supported by the UCSF Medical Scientist Training Program training grant T32GM141323 via the National Institute of General Medical Sciences (NIGMS). X.H. acknowledges the start-up fund provided by the School of Biomedical Engineering, Science, and Health Systems at Drexel University. This work was also partially funded by the UCSF Diabetes Center Pilot and Feasibility award funded in part by NIH grant P30 DK063720 (P.H., T.D. and Q.T.), NIH grant 1U54CA244438 (T.D.), the Northern California JDRF Center of Excellence 5-COE-2019-860-S-B (Q.T.) and the JDRF grant 2-SRA-2022-1221-S-B (T.D.). We thank E. Ronin and P. Ho for helpful discussion and trainings. We also thank E. Hansen for help with protocol validation and assistance.
Author information
Authors and Affiliations
Contributions
P.H. and X.H. designed the experiments and drafted the manuscript. P.H. and L.C. contributed to the experiments in Figs. 2–5. Y.C. and Z.H. contributed to the experiments in Fig. 2. P.H. analyzed the data. All authors contributed to the editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
T.D. and X.H. are inventors of a pending patent related to the technology described within this manuscript (WO2020014270A1). Q.T. is a co-founder, shareholder and scientific advisor of Sonoma Biotherapeutics. P.H., Y.C., Z.H. and L.C. declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Christopher Jewell, Evan Scott and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Huang, X. et al. Nat. Nanotechnol. 16, 214–223 (2021): https://doi.org/10.1038/s41565-020-00813-z
Supplementary information
Supplementary Table
Supplementary Table 1. Example reaction template for DNA−PLGA conjugation.
Source data
Source Data Fig. 2
Uncropped urea–PAGE gels for Fig. 2b–d.
Source Data Fig. 3
Uncropped SDS–PAGE gel for Fig. 3b, urea–PAGE gel used for quantification in Fig. 3c and urea–PAGE gel for Fig. 3e.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hadley, P., Chen, Y., Cline, L. et al. Precise surface functionalization of PLGA particles for human T cell modulation. Nat Protoc 18, 3289–3321 (2023). https://doi.org/10.1038/s41596-023-00887-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00887-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.