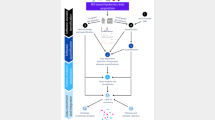
Abstract
Lipidomics studies suffer from analytical and annotation challenges because of the great structural similarity of many of the lipid species. To improve lipid characterization and annotation capabilities beyond those afforded by traditional mass spectrometry (MS)-based methods, multidimensional separation methods such as those integrating liquid chromatography, ion mobility spectrometry, collision-induced dissociation and MS (LC-IMS-CID-MS) may be used. Although LC-IMS-CID-MS and other multidimensional methods offer valuable hydrophobicity, structural and mass information, the files are also complex and difficult to assess. Thus, the development of software tools to rapidly process and facilitate confident lipid annotations is essential. In this Protocol Extension, we use the freely available, vendor-neutral and open-source software Skyline to process and annotate multidimensional lipidomic data. Although Skyline (https://skyline.ms/skyline.url) was established for targeted processing of LC-MS-based proteomics data, it has since been extended such that it can be used to analyze small-molecule data as well as data containing the IMS dimension. This protocol uses Skyline’s recently expanded capabilities, including small-molecule spectral libraries, indexed retention time and ion mobility filtering, and provides a step-by-step description for importing data, predicting retention times, validating lipid annotations, exporting results and editing our manually validated 500+ lipid library. Although the time required to complete the steps outlined here varies on the basis of multiple factors such as dataset size and familiarity with Skyline, this protocol takes ~5.5 h to complete when annotations are rigorously verified for maximum confidence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Data availability
All associated library and data files are publicly available at Panorama Public (https://panoramaweb.org/baker-lipid-ims.url) and Zenodo (https://zenodo.org/record/6374209#.YpzPMxPMJJU)47.
Code availability
Skyline source code is freely available at https://skyline.ms/source.url.
References
Soppert, J., Lehrke, M., Marx, N., Jankowski, J. & Noels, H. Lipoproteins and lipids in cardiovascular disease: from mechanistic insights to therapeutic targeting. Adv. Drug Deliv. Rev. 159, 4–33 (2020).
Maradonna, F. & Carnevali, O. Lipid metabolism alteration by endocrine disruptors in animal models: an overview. Front. Endocrinol. 9, 654 (2018).
Niemann, R. The effects of xenobiotics on hepatic lipid and lipo-protein metabolism. Exp. Pathol. 39, 213–232 (1990).
Miao, H. et al. Lipidomics biomarkers of diet-induced hyperlipidemia and its treatment with Poria cocos. J. Agric. Food Chem. 64, 969–979 (2016).
Perla, F. M., Prelati, M., Lavorato, M., Visicchio, D. & Anania, C. The role of lipid and lipoprotein metabolism in non-alcoholic fatty liver disease. Children 4, 46 (2017).
Wenk, M. R. The emerging field of lipidomics. Nat. Rev. Drug Discov. 4, 594–610 (2005).
Rolim, A. E. H., Henrique-Araújo, R., Ferraz, E. G., de Araújo Alves Dultra, F. K. & Fernandez, L. G. Lipidomics in the study of lipid metabolism: current perspectives in the omic sciences. Gene 554, 131–139 (2015).
Züllig, T., Trötzmüller, M. & Köfeler, H. C. Lipidomics from sample preparation to data analysis: a primer. Anal. Bioanal. Chem. 412, 2191–2209 (2020).
Leaptrot, K. L., May, J. C., Dodds, J. N. & Mclean, J. A. Ion mobility conformational lipid atlas for high confidence lipidomics. Nat. Commun. 10, 985 (2019).
Liebisch, G. et al. Lipidomics needs more standardization. Nat. Metab. 1, 745–747 (2019).
Yang, K., Dilthey, B. G. & Gross, R. W. Identification and quantitation of fatty acid double bond positional isomers: a shotgun lipidomics approach using charge-switch derivatization. Anal. Chem. 85, 9742–9750 (2013).
Ma, X. & Xia, Y. Pinpointing double bonds in lipids by Paternò-Büchi reactions and mass spectrometry. Angew. Chem. Int. Ed. Engl. 53, 2592–2596 (2014).
Brown, S. H. J., Mitchell, T. W. & Blanksby, S. J. Analysis of unsaturated lipids by ozone-induced dissociation. Biochim. Biophys. Acta 1811, 807–817 (2011).
Thomas, M. C., Mitchell, T. W. & Blanksby, S. J. Ozonolysis of phospholipid double bonds during electrospray ionization: a new tool for structure determination. J. Am. Chem. Soc. 128, 58–59 (2006).
Campbell, J. L. & Baba, T. Near-complete structural characterization of phosphatidylcholines using electron impact excitation of ions from organics. Anal. Chem. 87, 5837–5845 (2015).
Klein, D. R. & Brodbelt, J. S. Structural characterization of phosphatidylcholines using 193 nm ultraviolet photodissociation mass spectrometry. Anal. Chem. 89, 1516–1522 (2017).
Ryan, E., Nguyen, C. Q. N., Shiea, C. & Reid, G. E. Detailed structural characterization of sphingolipids via 193 nm ultraviolet photodissociation and ultra high resolution tandem mass spectrometry. J. Am. Soc. Mass Spectrom. 28, 1406–1419 (2017).
Kyle, J. E. et al. Uncovering biologically significant lipid isomers with liquid chromatography, ion mobility spectrometry and mass spectrometry. Analyst 141, 1649–1659 (2016).
Maclean, B. X. et al. Using Skyline to analyze data-containing liquid chromatography, ion mobility spectrometry, and mass spectrometry dimensions. J. Am. Soc. Mass Spectrom. 29, 2182–2188 (2018).
Hancock, S. E., Poad, B. L. J., Batarseh, A., Abbott, S. K. & Mitchell, T. W. Advances and unresolved challenges in the structural characterization of isomeric lipids. Anal. Biochem. 524, 45–55 (2017).
Paglia, G., Kliman, M., Claude, E., Geromanos, S. & Astarita, G. Applications of ion-mobility mass spectrometry for lipid analysis. Anal. Bioanal. Chem. 407, 4995–5007 (2015).
Groessl, M., Graf, S. & Knochenmuss, R. High resolution ion mobility-mass spectrometry for separation and identification of isomeric lipids. Analyst 140, 6904–6911 (2015).
Egertson, J. D., MacLean, B., Johnson, R., Xuan, Y. & MacCoss, M. J. Multiplexed peptide analysis using data-independent acquisition and Skyline. Nat. Protoc. 10, 887–903 (2015).
Maclean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Pino, L. K. et al. The Skyline ecosystem: informatics for quantitative mass spectrometry proteomics. Mass Spectrom. Rev. 39, 229–244 (2020).
Adams, K. J. et al. Skyline for small molecules: a unifying software package for quantitative metabolomics. J. Proteome Res 19, 1447–1458 (2020).
Kirkwood, K. I. et al. Development and application of multidimensional lipid libraries to investigate lipidomic dysregulation related to smoke inhalation injury severity. J. Proteome Res 21, 232–242 (2022).
Fahy, E. et al. A comprehensive classification system for lipids. J. Lipid Res. 46, 839–862 (2005).
Peng, B. et al. LipidCreator workbench to probe the lipidomic landscape. Nat. Commun. 11, 2057 (2020).
Kind, T. et al. LipidBlast in silico tandem mass spectrometry database for lipid identification. Nat. Methods 10, 755–758 (2013).
Murphy, R. Tandem Mass Spectrometry of Lipids: Molecular Analysis of Complex Lipids (Royal Society of Chemistry, 2015).
Escher, C. et al. Using iRT, a normalized retention time for more targeted measurement of peptides. Proteomics 12, 1111–1121 (2012).
Picache, J. A. et al. Collision cross section compendium to annotate and predict multi-omic compound identities. Chem. Sci. 10, 983–993 (2019).
Kyle, J. E. et al. LIQUID: an-open source software for identifying lipids in LC-MS/MS-based lipidomics data. Bioinformatics 33, 1744–1746 (2017).
Tsugawa, H. et al. MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526 (2015).
Zhou, Z. et al. LipidIMMS Analyzer: integrating multi-dimensional information to support lipid identification in ion mobility—mass spectrometry based lipidomics. Bioinformatics 35, 698–700 (2019).
Koelmel, J. P. et al. LipidMatch: an automated workflow for rule-based lipid identification using untargeted high-resolution tandem mass spectrometry data. BMC Bioinforma. 18, 331 (2017).
Hartler, J. et al. Deciphering lipid structures based on platform-independent decision rules. Nat. Methods 14, 1171–1174 (2017).
Kochen, M. A. et al. Greazy: open-source software for automated phospholipid tandem mass spectrometry identification. Anal. Chem. 88, 5733–5741 (2016).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 3, 211–221 (2007).
Blaženović, I., Kind, T., Ji, J. & Fiehn, O. Software tools and approaches for compound identification of LC-MS/MS data in metabolomics. Metabolites 8, 1–23 (2018).
Köfeler, H. C. et al. Quality control requirements for the correct annotation of lipidomics data. Nat. Commun. 12, 4771 (2021).
Koelmel, J. P., Ulmer, C. Z., Jones, C. M., Yost, R. A. & Bowden, J. A. Common cases of improper lipid annotation using high-resolution tandem mass spectrometry data and corresponding limitations in biological interpretation. Biochim. Biophys. Acta 1862, 766–770 (2017).
Rampler, E. et al. Recurrent topics in mass spectrometry-based metabolomics and lipidomics—standardization, coverage, and throughput. Anal. Chem. 93, 519–545 (2021).
Frewen, B. E. et al. Analysis of peptide MS/MS spectra from large-scale proteomics experiments using spectrum libraries. Anal. Chem. 78, 5678–5684 (2006).
Odenkirk, M. T. et al. Combining micropunch histology and multidimensional lipidomic measurements for in-depth tissue mapping. ACS Meas. Sci. Au 2, 67–75 (2022).
Kirkwood, K. I. et al. Utilizing Skyline to analyze lipidomics data containing liquid chromatography, ion mobility spectrometry and mass spectrometry dimensions. Zenodo https://zenodo.org/record/6374209#.YpzYhBPMJJU (2022).
Acknowledgements
Portions of this research were supported by grants from the NIH National Institute of Environmental Health Sciences (P30 ES025128, P42 ES027704 and P42 ES031009), NIH National Institute of General Medicine Sciences (R24 GM141156, P41 GM103533 and T32 GM133366), a cooperative agreement with the United States Environmental Protection Agency (STAR RD 84003201) and startup funds from North Carolina State University. In addition, most of the LC-IMS-CID-MS measurements were made in the Molecular Education, Technology, and Research Innovation Center (METRIC) at North Carolina State University.
Author information
Authors and Affiliations
Contributions
K.I.K., B.S.P., N.S., K.T., M.J.M., B.X.M. and E.S.B. developed and optimized the protocol and Skyline tools. K.I.K. drafted the text of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Kirkwood, K. I. et al. J. Proteome Res. 21, 232–242 (2022): https://doi.org/10.1021/acs.jproteome.1c00820
Odenkirk, M. T. et al. ACS Meas. Sci. Au 2, 67–75 (2022): https://doi.org/10.1021/acsmeasuresciau.1c00035
Key data used in this protocol
NCSU Baker Lab—Lipid Libraries: https://panoramaweb.org/baker-lipid-ims.url
Kirkwood, K. I. et al. Zenodo https://doi.org/10.5281/zenodo.6374209
This protocol is an extension to: Nat. Protoc. 10, 887–903 (2015): https://doi.org/10.1038/nprot.2015.055
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Tables 3–5 and Supplementary Figs. 1–17.
Supplementary Table 1
Transition lists
Supplementary Table 2
iRT prediction results
Rights and permissions
About this article
Cite this article
Kirkwood, K.I., Pratt, B.S., Shulman, N. et al. Utilizing Skyline to analyze lipidomics data containing liquid chromatography, ion mobility spectrometry and mass spectrometry dimensions. Nat Protoc 17, 2415–2430 (2022). https://doi.org/10.1038/s41596-022-00714-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00714-6
This article is cited by
-
Ozone-enabled fatty acid discovery reveals unexpected diversity in the human lipidome
Nature Communications (2023)
-
A mass spectrum-oriented computational method for ion mobility-resolved untargeted metabolomics
Nature Communications (2023)
-
Utilizing Skyline to analyze lipidomics data containing liquid chromatography, ion mobility spectrometry and mass spectrometry dimensions
Nature Protocols (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.