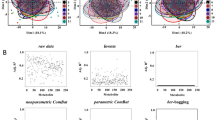
Abstract
Metabolic phenotyping is an important tool in translational biomedical research. The advanced analytical technologies commonly used for phenotyping, including mass spectrometry (MS) and nuclear magnetic resonance (NMR) spectroscopy, generate complex data requiring tailored statistical analysis methods. Detailed protocols have been published for data acquisition by liquid NMR, solid-state NMR, ultra-performance liquid chromatography (LC-)MS and gas chromatography (GC-)MS on biofluids or tissues and their preprocessing. Here we propose an efficient protocol (guidelines and software) for statistical analysis of metabolic data generated by these methods. Code for all steps is provided, and no prior coding skill is necessary. We offer efficient solutions for the different steps required within the complete phenotyping data analytics workflow: scaling, normalization, outlier detection, multivariate analysis to explore and model study-related effects, selection of candidate biomarkers, validation, multiple testing correction and performance evaluation of statistical models. We also provide a statistical power calculation algorithm and safeguards to ensure robust and meaningful experimental designs that deliver reliable results. We exemplify the protocol with a two-group classification study and data from an epidemiological cohort; however, the protocol can be easily modified to cover a wider range of experimental designs or incorporate different modeling approaches. This protocol describes a minimal set of analyses needed to rigorously investigate typical datasets encountered in metabolic phenotyping.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All the data and software reported in the paper are open source and freely available on GitHub and Zenodo repositories: https://github.com/Gscorreia89/chemometrics-tutorials, https://github.com/phenomecentre/metabotyping-dementia-urine and https://doi.org/10.5281/zenodo.4053166.
Code availability
All the data and software reported in the paper are open source and freely available on GitHub and Zenodo repositories: https://github.com/Gscorreia89/chemometrics-tutorials, https://github.com/phenomecentre/metabotyping-dementia-urine and https://doi.org/10.5281/zenodo.4053166.
References
Nicholson, J. K., Lindon, J. C. & Holmes, E. ‘Metabonomics’: understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica 29, 1181–1189 (2008).
Holmes, E., Wilson, I. D. & Nicholson, J. K. Metabolic phenotyping in health and disease. Cell 134, 714–717 (2008).
Nicholson, J. K. et al. Metabolic phenotyping in clinical and surgical environments. Nature 491, 384–392 (2012).
Surowiec, I. et al. Quantification of run order effect on chromatography - mass spectrometry profiling data. J. Chromatogr. A 1568, 229–234 (2018).
Leek, J. T. et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat. Rev. Genet. 11, 733–739 (2010).
Lewis, M. R. et al. Development and application of UPLC-ToF MS for precision large scale urinary metabolic phenotyping. Anal. Chem. https://doi.org/10.1021/acs.analchem.6b01481 (2016).
Fages, A. et al. Batch profiling calibration for robust NMR metabonomic data analysis. Anal. Bioanal. Chem. 405, 8819–8827 (2013).
Dieterle, F., Ross, A., Schlotterbeck, G. & Senn, H. Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in 1H NMR metabonomics. Anal. Chem. 78, 4281–4290 (2006).
Posma, J. M. et al. Optimized phenotypic biomarker discovery and confounder elimination via covariate-adjusted projection to latent structures from metabolic spectroscopy data. J. Proteome Res. https://doi.org/10.1021/acs.jproteome.7b00879 (2018).
Blaise, B. J. et al. Statistical recoupling prior to significance testing in nuclear magnetic resonance based metabonomics. Anal. Chem. 81, 6242–6251 (2009).
Navratil, V., Pontoizeau, C., Billoir, E. & Blaise, B. J. SRV: an open-source toolbox to accelerate the recovery of metabolic biomarkers and correlations from metabolic phenotyping datasets. Bioinformatics 29, 1348–1349 (2013).
Kuhl, C., Tautenhahn, R., Böttcher, C., Larson, T. R. & Neumann, S. CAMERA: an integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass spectrometry data sets. Anal. Chem. 84, 283–289 (2012).
Moseley, H. N. B. Error analysis and propagation in metabolomics data analysis. Comput. Struct. Biotechnol. J. 4, e201301006 (2013).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Forsberg, E. M. et al. Data processing, multi-omic pathway mapping, and metabolite activity analysis using XCMS Online. Nat. Protoc. https://doi.org/10.1038/nprot.2017.151 (2018).
Pluskal, T. T., Castillo, S., Villar-Briones, A., Oresic, M. & Orešič, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinforma. 11, 395 (2010).
Li, B. et al. NOREVA: normalization and evaluation of MS-based metabolomics data. Nucleic Acids Res. https://doi.org/10.1093/nar/gkx449 (2017).
Hughes, G. et al. MSPrep—summarization, normalization and diagnostics for processing of mass spectrometry-based metabolomic data. Bioinformatics https://doi.org/10.1093/bioinformatics/btt589 (2014).
Wehrens, R., Weingart, G. & Mattivi, F. MetaMS: an open-source pipeline for GC-MS-based untargeted metabolomics. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. https://doi.org/10.1016/j.jchromb.2014.02.051 (2014).
Wang, S. & Yang, H. pseudoQC: a regression-based simulation software for correction and normalization of complex metabolomics and proteomics datasets. Proteomics https://doi.org/10.1002/pmic.201900264 (2019).
Biswas, A. et al. Metdat: a modular and workflow-based free online pipeline for mass spectrometry data processing, analysis and interpretation. Bioinformatics https://doi.org/10.1093/bioinformatics/btq436 (2010).
Shen, X., Zhu, Z. J. & Wren, J. MetFlow: an interactive and integrated workflow for metabolomics data cleaning and differential metabolite discovery. Bioinformatics https://doi.org/10.1093/bioinformatics/bty1066 (2019).
Hao, L. et al. Metandem: an online software tool for mass spectrometry-based isobaric labeling metabolomics. Anal. Chim. Acta https://doi.org/10.1016/j.aca.2019.08.046 (2019).
Verhoeven, A., Giera, M. & Mayboroda, O. A. KIMBLE: a versatile visual NMR metabolomics workbench in KNIME. Anal. Chim. Acta https://doi.org/10.1016/j.aca.2018.07.070 (2018).
Hao, J., Astle, W., De Iorio, M. & Ebbels, T. M. D. BATMAN—an R package for the automated quantification of metabolites from nuclear magnetic resonance spectra using a Bayesian model. Bioinformatics 28, 2088–2090 (2012).
Beirnaert, C. et al. speaq 2.0: A complete workflow for high-throughput 1D NMR spectra processing and quantification. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1006018 (2018).
Chawade, A., Alexandersson, E. & Levander, F. Normalyzer: a tool for rapid evaluation of normalization methods for omics data sets. J. Proteome Res. https://doi.org/10.1021/pr401264n (2014).
Wang, S. et al. MetaboGroup S: a group entropy-based web platform for evaluating normalization methods in blood metabolomics data from maintenance hemodialysis patients. Anal. Chem. https://doi.org/10.1021/acs.analchem.8b03065 (2018).
Xia, J., Sinelnikov, I. V., Han, B. & Wishart, D. S. MetaboAnalyst 3.0—making metabolomics more meaningful. Nucleic Acids Res. 43, W251–W257 (2015).
Giacomoni, F. et al. Workflow4Metabolomics: a collaborative research infrastructure for computational metabolomics. Bioinformatics https://doi.org/10.1093/bioinformatics/btu813 (2015).
Wen, B., Mei, Z., Zeng, C. & Liu, S. metaX: a flexible and comprehensive software for processing metabolomics data. BMC Bioinformatics https://doi.org/10.1186/s12859-017-1579-y (2017).
Cardoso, S., Afonso, T., Maraschin, M. & Rocha, M. WebSpecmine: a website for metabolomics data analysis and mining. Metabolites https://doi.org/10.3390/metabo9100237 (2019).
Sud, M. et al. Metabolomics Workbench: an international repository for metabolomics data and metadata, metabolite standards, protocols, tutorials and training, and analysis tools. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv1042 (2016).
Bictash, M. et al. Opening up the ‘Black Box’: metabolic phenotyping and metabolome-wide association studies in epidemiology. J. Clin. Epidemiol. 63, 970–979 (2010).
Cloarec, O. et al. Evaluation of the orthogonal projection on latent structure model limitations caused by chemical shift variability and improved visualization of biomarker changes in 1H NMR spectroscopic metabonomic studies. Anal. Chem. 77, 517–526 (2005).
Trygg, J., Holmes, E. & Lundstedt, T. Chemometrics in metabonomics. J. Proteome Res. 6, 469–479 (2007).
Tzoulaki, I., Ebbels, T. M. D., Valdes, A., Elliott, P. & Ioannidis, J. P. A. Design and analysis of metabolomics studies in epidemiologic research: a primer on -omic technologies. Am. J. Epidemiol. 180, 129–139 (2014).
Ren, S., Hinzman, A. A., Kang, E. L., Szczesniak, R. D. & Lu, L. J. Computational and statistical analysis of metabolomics data. Metabolomics 11, 1492–1513 (2015).
Xia, J., Psychogios, N., Young, N. & Wishart, D. S. MetaboAnalyst: a web server for metabolomic data analysis and interpretation. Nucleic Acids Res. 37, W652–W660 (2009).
Gromski, P. S. et al. A tutorial review: metabolomics and partial least squares-discriminant analysis—a marriage of convenience or a shotgun wedding. Anal. Chim. Acta 879, 10–23 (2015).
Smilde, A. K. et al. Dynamic metabolomic data analysis: a tutorial review. Metabolomics 6, 3–17 (2010).
Beckonert, O. et al. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat. Protoc. 2, 2692–2703 (2007).
Beckonert, O. et al. High-resolution magic-angle-spinning NMR spectroscopy for metabolic profiling of intact tissues. Nat. Protoc. 5, 1019–1032 (2010).
Southam, A. D., Weber, R. J. M., Engel, J., Jones, M. R. & Viant, M. R. A complete workflow for high-resolution spectral-stitching nanoelectrospray direct-infusion mass-spectrometry-based metabolomics and lipidomics. Nat. Protoc. 12, 255–273 (2017).
Dunn, W. B. et al. Metabolic profiling of serum using ultra performance liquid chromatography and the LTQ-Orbitrap mass spectrometry system. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 871, 288–298 (2008).
Want, E. J. et al. Global metabolic profiling procedures for urine using UPLC-MS. Nat. Protoc. 5, 1005–1018 (2010).
Want, E. J. et al. Global metabolic profiling of animal and human tissues via UPLC-MS. Nat. Protoc. 8, 17–32 (2013).
Dona, A. C. et al. Precision high-throughput proton NMR spectroscopy of human urine, serum, and plasma for large-scale metabolic phenotyping. Anal. Chem. 86, 9887–9894 (2014).
Dunn, W. B. et al. Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nat. Protoc. 6, 1060–1083 (2011).
Jiménez, B. et al. Quantitative lipoprotein subclass and low molecular weight metabolite analysis in human serum and plasma by 1H NMR spectroscopy in a multilaboratory trial. Anal. Chem. https://doi.org/10.1021/acs.analchem.8b02412 (2018).
Broadhurst, D. et al. Guidelines and considerations for the use of system suitability and quality control samples in mass spectrometry assays applied in untargeted clinical metabolomic studies. Metabolomics https://doi.org/10.1007/s11306-018-1367-3 (2018).
Mahieu, N. G. & Patti, G. J. Systems-level annotation of a metabolomics data set reduces 25 000 features to fewer than 1000 unique metabolites. Anal. Chem. 89, 10397–10406 (2017).
Craig, A., Cloarec, O., Holmes, E., Nicholson, J. K. & Lindon, J. C. Scaling and normalization effects in NMR spectroscopic metabonomic data sets. Anal. Chem. 78, 2262–2267 (2006).
Johansson, E., Wold, S. & Sjödin, K. Minimizing effects of closure on analytical data. Anal. Chem. 56, 1685–1688 (1984).
Chayes, F. & Trochimczyk, J. An effect of closure on the structure of principal components. J. Int. Assoc. Math. Geol. 10, 323–333 (1978).
Rietjens, M. Reduction of error propagation due to normalization: {Effect} of error propagation and closure on spurious correlations. Anal. Chim. Acta 316, 205–215 (1995).
Saccenti, E. Correlation patterns in experimental data are affected by normalization procedures: consequences for data analysis and network inference. J. Proteome Res. 16, 619–634 (2017).
Kohl, S. M. et al. State-of-the art data normalization methods improve NMR-based metabolomic analysis. Metabolomics 8, 146–160 (2012).
Wu, Y. & Li, L. Sample normalization methods in quantitative metabolomics. J. Chromatogr. A 1430, 80–95 (2016).
Van Der Kloet, F. M., Bobeldijk, I., Verheij, E. R. & Jellema, R. H. Analytical error reduction using single point calibration for accurate and precise metabolomic phenotyping. J. Proteome Res. 8, 5132–5141 (2009).
Berg, R. A., van den, Hoefsloot, H. C. J., Westerhuis, J. A., Smilde, A. K. & van der Werf, M. J. Centering, scaling, and transformations: improving the biological information content of metabolomics data. BMC Genomics 7, 142 (2006).
Rocke, D. M. & Durbin, B. Approximate variance-stabilizing transformations for gene-expression microarray data. Bioinformatics 19, 966–972 (2003).
Purohit, P. V., Rocke, D. M., Viant, M. R. & Woodruff, D. L. Discrimination models using variance-stabilizing transformation of metabolomic NMR data. OMICS 8, 118–130 (2004).
Bro, R. & Smilde, A. K. Principal component analysis. Anal. Methods 6, 2812–2831 (2014).
Wold, S., Esbensen, K. & Geladi, P. Principal component analysis. Chemom. Intell. Lab. Syst. 2, 37–52 (1987).
Geladi, P. & Kowalski, B. R. Partial least-squares regression: a tutorial. Anal. Chim. Acta 185, 1–17 (1986).
Wold, S. et al. PLS-regression: a basic tool of chemometrics. Chemom. Intell. Lab. Syst. 58, 109–130 (2001).
Barker, M. & Rayens, W. Partial least squares for discrimination. J. Chemom. 17, 166–173 (2003).
Trygg, J. & Wold, S. Orthogonal projections to latent structures (O-PLS). J. Chemom. 16, 119–128 (2002).
Wiklund, S. et al. Visualization of GC/TOF-MS-based metabolomics data for identification of biochemically interesting compounds using OPLS class models. Anal. Chem. 80, 115–122 (2008).
Bylesjo, M. et al. OPLS discriminant analysis: combining the strengths of PLS-DA and SIMCA classification. J. Chemom. 20, 341–351 (2006).
Wold, S., Antti, H., Lindgren, F. & Öhman, J. Orthogonal signal correction of near-infrared spectra. Chemom. Intell. Lab. Syst. 44, 175–185 (1998).
Fearn, T. On orthogonal signal correction. Chemom. Intell. Lab. Syst. 50, 47–52 (2000).
Szymańska, E., Saccenti, E., Smilde, A. K. & Westerhuis, J. A. Double-check: validation of diagnostic statistics for PLS-DA models in metabolomics studies. Metabolomics 8, 3–16 (2012).
Triba, M. N. et al. PLS/OPLS models in metabolomics: the impact of permutation of dataset rows on the K-fold cross-validation quality parameters. Mol. BioSyst. 11, 13–19 (2015).
MacGregor, J. F. & Kourti, T. Statistical process control of multivariate processes. Control Eng. Pract. 3, 403–414 (1995).
Mahalanobis, P. C. On the generalized distance in statistics. Proc. Natl Inst. Sci. India 2, 49–55 (1936).
Eriksson, L., Byrne, T., Johansson, E., Trygg, J. & Vikström, C. Multi- and Megavariate Data Analysis: Basic Principles and Applications (Umetrics Academy, 2013).
Martens, H. & Naes, T. Multivariate Calibration (John Wiley & Sons, 1989).
Hastie, T., Tibshirani, R. & Friedman, J. The elements of statistical learning. Elements 1, 337–387 (2009).
Broadhurst, D. I. D. I. & Kell, D. B. Statistical strategies for avoiding false discoveries in metabolomics and related experiments. Metabolomics 2, 171–196 (2006).
Varma, S. et al. Bias in error estimation when using cross-validation for model selection. BMC Bioinforma. 7, 91 (2006).
Burman, P. A comparative study of ordinary cross-validation, v-fold cross-validation and the repeated learning-testing methods. Biometrika 76, 503–514 (1989).
Lindgren, F., Hansen, B., Karcher, W., Sjöström, M. & Eriksson, L. Model validation by permutation tests: applications to variable selection. J. Chemom. 10, 521–532 (1996).
van der Voet, H. Comparing the predictive accuracy of models using a simple randomization test. Chemom. Intell. Lab. Syst. 25, 313–323 (1994).
Efron, B. Bootstrap methods: another look at the jackknife. Ann. Stat. 7, 1–26 (1979).
Zweig, M. H. & Campbell, G. Receiver-operating characteristics (ROC) plots: a fundamental evaluation tool in clinical medicine. Clin. Chem. 39, 561–577 (1993).
de Jong, S. SIMPLS: an alternative approach to partial least squares regression. Chemom. Intell. Lab. Syst. 18, 251–263 (1993).
Galindo-Prieto, B., Eriksson, L. & Trygg, J. Variable influence on projection (VIP) for orthogonal projections to latent structures (OPLS). J. Chemom. 28, 623–632 (2014).
Chong, I.-G. & Jun, C.-H. Performance of some variable selection methods when multicollinearity is present. Chemom. Intell. Lab. Syst. 78, 103–112 (2005).
Frank, I. E. & Friedman, J. H. A statistical view of some chemometrics regression tools. Technometrics 35, 109–135 (1993).
Krämer, N. An overview on the shrinkage properties of partial least squares regression. Comput. Stat. 22, 249–273 (2007).
Abdi, H. H. The Bonferonni and Šidák corrections for multiple comparisons. Encycl. Meas. Stat. 1, 1–9 (2007).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing on JSTOR. J. R. Stat. Soc. B 57, 289–300 (1995).
Benjamini, Y. & Yekutieli, D. The control of the false discovery rate in multiple testing under depencency. Ann. Stat. 29, 1165–1188 (2001).
Ferreira, J. A. & Zwinderman, A. Approximate power and sample size calculations with the Benjamini–Hochberg method. Int. J. Biostat. https://doi.org/10.2202/1557-4679.1018 (2006).
Nyamundanda, G., Gormley, I. C., Fan, Y., Gallagher, W. M. & Brennan, L. MetSizeR: selecting the optimal sample size for metabolic studies using an analysis based approach. BMC Bioinforma. 14, 338–345 (2013).
Jung, S.-H. & Young, S. S. Power and sample size calculation for microarray studies. J. Biopharm. Stat. 22, 30–42 (2012).
Ferreira, J. A. & Zwinderman, A. Approximate sample size calculations with microarray data: an illustration. Stat. Appl. Genet. Mol. Biol. 5, Article25 (2006).
Jung, S.-H., Bang, H. & Young, S. Sample size calculation for multiple testing in microarray data analysis. Biostatistics 6, 157–169 (2005).
Blaise, B. J. et al. Power analysis and sample size determination in metabolic phenotyping. Anal. Chem. 88, 5179–5188 (2016).
Billoir, E., Navratil, V. & Blaise, B. J. Sample size calculation in metabolic phenotyping studies. Brief. Bioinform. 16, 813–819 (2014).
Blaise, B. J. Data-driven sample size determination for metabolic phenotyping studies. Anal. Chem. 85, 8943–8950 (2013).
Continuum Analytics. Anaconda Software Distribution. Computer software. Vers. 2-2.4.0. Continuum Analytics, Nov. 2016. https://continuum.io (2016).
R Core Team & Team, R. C. R: A Language and Environment for Statistical Computing (2017).
Pedregosa, F., Grisel, O., Weiss, R., Passos, A. & Brucher, M. Scikit-learn: Machine Learning in Python. 12, 2825–2830 (2011).
Kluyver, T. et al. Jupyter Notebooks—a publishing format for reproducible computational workflows. Positioning and Power in Academic Publishing: Players, Agents and Agendas https://doi.org/10.3233/978-1-61499-649-1-87 (2016).
Blaise, B. J. et al. Metabolic profiling strategy of Caenorhabditis elegans by whole-organism nuclear magnetic resonance. J. Proteome Res. 8, 2542–2550 (2009).
Blaise, B. J. et al. Metabotyping of Caenorhabditis elegans reveals latent phenotypes. Proc. Natl Acad. Sci. USA. 104, 19808–19812 (2007).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. Ser. B Stat. Methodol. https://doi.org/10.1111/j.1467-9868.2005.00503.x (2005).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Ser. B https://doi.org/10.1111/j.2517-6161.1996.tb02080.x (1996).
Hoerl, A. E. & Kennard, R. W. Ridge regression: biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970).
Breiman, L. Random forests. Mach. Learn. https://doi.org/10.1023/A:1010933404324 (2001).
Sangster, T., Major, H., Plumb, R., Wilson, A. J. & Wilson, I. D. A pragmatic and readily implemented quality control strategy for HPLC-MS and GC-MS-based metabonomic analysis. Analyst 131, 1075–1078 (2006).
Sands, C. J. et al. The nPYc-Toolbox, a Python module for the pre-processing, quality-control and analysis of metabolic profiling datasets. Bioinformatics https://doi.org/10.1093/bioinformatics/btz566 (2019).
Kamleh, M. A., Ebbels, T. M. D., Spagou, K., Masson, P. & Want, E. J. Optimizing the use of quality control samples for signal drift correction in large-scale urine metabolic profiling studies. Anal. Chem. 84, 2670–2677 (2012).
Wehrens, R. et al. Improved batch correction in untargeted MS-based metabolomics. Metabolomics 12, 88 (2016).
Mehmood, T., Liland, K. H., Snipen, L. & Sæbø, S. A review of variable selection methods in partial least squares regression. Chemom. Intell. Lab. Syst. 118, 62–69 (2012).
Lê Cao, K.-A., Boitard, S. & Besse, P. Sparse PLS discriminant analysis: biologically relevant feature selection and graphical displays for multiclass problems. BMC Bioinforma. 12, 253 (2011).
Cloarec, O. et al. Statistical total correlation spectroscopy: an exploratory approach for latent biomarker identification from metabolic 1H NMR data sets. Anal. Chem. 77, 1282–1289 (2005).
Acknowledgements
B.J.B. is supported by the Analytical Chemistry Trust Fund (Tom West Analytical Fellowship) and the Fondation Bettencourt Schueller. G.C. is supported by the National Institute for Health Research (NIHR) Imperial Biomedical Research Centre (BRC). T.E. acknowledges support from the EU COSMOS project (grant agreement 312941), the EU PhenoMeNal project (Project reference: 654241), UK BBSRC grant BB/T007974/1 and NIH grant R01 HL133932-01. E.H. is supported by the Department of Jobs, Tourism, Science and Innovation, Government of Western Australian Government through the Premier’s Science Fellowship Program. This work was supported by the Medical Research Council (MRC) and National Institute for Health Research (NIHR) (grant number MC_PC_12025) and the MRC UK Consortium for MetAbolic Phenotyping (MAP/UK) (grant number MR/S010483/1). The Division of Systems Medicine is funded by grants from the MRC, BBSRC and NIHR, an Integrative Mammalian Biology (IMB) Capacity Building Award and an FP7- HEALTH- 2009- 241592 EuroCHIP grant and is supported by the NIHR Biomedical Research Centre Funding Scheme. The views expressed are those of the authors and not necessarily those of the (name of funder), the NHS, the NIHR or the Department of Health. AddNeuroMed was supported by the Innovative Medicines Initiative (IMI) Joint Undertaking under EMIF grant agreement, resources of which are composed of financial contribution from the European Union’s Seventh Framework Programme (FP7/2007-2013) and EFPIA companies’ in kind contribution. We thank the clinical leads for the consortium S. Lovestone (PI), H. Soininen, P. Mecocci, M. Tsolaki, B. Vellas and I. Kłoszewska for kind access to the data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Holmes, E. et al. Nature 453, 396–400 (2008): https://doi.org/10.1038/nature06882
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2 and Supplementary Procedures 1.1 and 1.2.
Rights and permissions
About this article
Cite this article
Blaise, B.J., Correia, G.D.S., Haggart, G.A. et al. Statistical analysis in metabolic phenotyping. Nat Protoc 16, 4299–4326 (2021). https://doi.org/10.1038/s41596-021-00579-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00579-1
This article is cited by
-
Quartet metabolite reference materials for inter-laboratory proficiency test and data integration of metabolomics profiling
Genome Biology (2024)
-
Presenting metabolomics analyses: what’s in a number?
The EMBO Journal (2024)
-
Multi-Attribute Subset Selection enables prediction of representative phenotypes across microbial populations
Communications Biology (2024)
-
Gut microbiota-dependent phenylacetylglutamine in cardiovascular disease: current knowledge and new insights
Frontiers of Medicine (2024)
-
Dysregulation of acyl carnitines, pentose phosphate pathway and arginine and ornithine metabolism are associated with decline in intrinsic capacity in Chinese older adults
Aging Clinical and Experimental Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.