Abstract
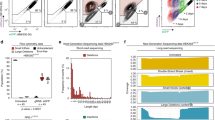
The dynamics of DNA double-strand break (DSB) repairs including homology-directed repair and nonhomologous end joining play an important role in diseases and therapies. However, investigating DSB repair is typically a low-throughput and cross-sectional process, requiring disruption of cells and organisms for subsequent nuclease-, sequencing- or reporter-based assays. In this protocol, we provide instructions for establishing a bioluminescent repair reporter system using engineered Gaussia and Vargula luciferases for noninvasive tracking of homology-directed repair and nonhomologous end joining, respectively, induced by SceI meganuclease, SpCas9 or SpCas9 D10A nickase-mediated editing. We also describe complementation with orthogonal DSB repair assays and omics analyses to validate the reporter readouts. The bioluminescent repair reporter system provides longitudinal and rapid readout (~seconds per sample) to accurately and efficiently measure the efficacy of genome-editing tools and small-molecule modulators on DSB repair. This protocol takes ~2–4 weeks to establish, and as little as 2 h to complete the assay. The entire bioluminescent repair reporter procedure can be performed by one person with standard molecular biology expertise and equipment. However, orthogonal DNA repair assays would require a specialized facility that performs Sanger sequencing or next-generation sequencing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All requests should be addressed to the corresponding authors.
References
Maynard, S., Fang, E. F., Scheibye-Knudsen, M., Croteau, D. L. & Bohr, V. A. DNA damage, DNA repair, aging, and neurodegeneration. Cold Spring Harb. Perspect. Med. https://doi.org/10.1101/cshperspect.a025130 (2015).
Gomez-Herreros, F. DNA Double strand breaks and chromosomal translocations induced by DNA topoisomerase II. Front. Mol. Biosci. 6, 141 (2019).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Chang, H. H. Y., Pannunzio, N. R., Adachi, N. & Lieber, M. R. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat. Rev. Mol. Cell Biol. 18, 495–506 (2017).
Doudna, J. A. The promise and challenge of therapeutic genome editing. Nature 578, 229–236 (2020).
Toulany, M. Targeting DNA double-strand break repair pathways to improve radiotherapy response. Genes (Basel) https://doi.org/10.3390/genes10010025 (2019).
Kesari, S. et al. DNA damage response and repair: insights into strategies for radiation sensitization of gliomas. Future Oncol. 7, 1335–1346 (2011).
Gil Del Alcazar, C. R., Todorova, P. K., Habib, A. A., Mukherjee, B. & Burma, S. Augmented HR repair mediates acquired temozolomide resistance in glioblastoma. Mol. Cancer Res. 14, 928–940 (2016).
Welsh, J. W. et al. Rad51 protein expression and survival in patients with glioblastoma multiforme. Int. J. Radiat. Oncol. Biol. Phys. 74, 1251–1255 (2009).
King, H. O. et al. RAD51 is a selective DNA repair target to radiosensitize glioma stem cells. Stem Cell Rep. 8, 125–139 (2017).
Mashal, R. D., Koontz, J. & Sklar, J. Detection of mutations by cleavage of DNA heteroduplexes with bacteriophage resolvases. Nat. Genet. 9, 177–183 (1995).
Germini, D. et al. A comparison of techniques to evaluate the effectiveness of genome editing. Trends Biotechnol. 36, 147–159 (2018).
Chien, J. C. et al. A multiplexed bioluminescent reporter for sensitive and non-invasive tracking of DNA double strand break repair dynamics in vitro and in vivo. Nucleic Acids Res. 48, e100 (2020).
Haddock, S. H., Moline, M. A. & Case, J. F. Bioluminescence in the sea. Ann. Rev. Mar. Sci. 2, 443–493 (2010).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Bovenberg, M. S. et al. Multiplex blood reporters for simultaneous monitoring of cellular processes. Anal. Chem. 85, 10205–10210 (2013).
Nakajima, Y., Kobayashi, K., Yamagishi, K., Enomoto, T. & Ohmiya, Y. cDNA cloning and characterization of a secreted luciferase from the luminous Japanese ostracod, Cypridina noctiluca. Biosci. Biotechnol. Biochem. 68, 565–570 (2004).
Wurdinger, T. et al. A secreted luciferase for ex vivo monitoring of in vivo processes. Nat. Methods 5, 171–173 (2008).
Hendel, A. et al. Quantifying genome-editing outcomes at endogenous loci with SMRT sequencing. Cell Rep. 7, 293–305 (2014).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Certo, M. T. et al. Tracking genome engineering outcome at individual DNA breakpoints. Nat. Methods 8, 671–676 (2011).
Yang, Z. et al. Fast and sensitive detection of indels induced by precise gene targeting. Nucleic Acids Res. 43, e59 (2015).
Ramakrishna, S. et al. Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nuclease-induced mutations. Nat. Commun. 5, 3378 (2014).
Cornu, T. I., Mussolino, C. & Cathomen, T. Refining strategies to translate genome editing to the clinic. Nat. Med. 23, 415–423 (2017).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR-Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015).
Yeh, C. D., Richardson, C. D. & Corn, J. E. Advances in genome editing through control of DNA repair pathways. Nat. Cell Biol. 21, 1468–1478 (2019).
Beckta, J. M., Bindra, R. S. & Chalmers, A. J. Targeting DNA repair in gliomas. Curr. Opin. Neurol. 32, 878–885 (2019).
Huang, R. X. & Zhou, P. K. DNA damage response signaling pathways and targets for radiotherapy sensitization in cancer. Signal Transduct. Target Ther. 5, 60 (2020).
Peng, Y. et al. The deubiquitylating enzyme USP15 regulates homologous recombination repair and cancer cell response to PARP inhibitors. Nat. Commun. 10, 1224 (2019).
Hendel, A., Fine, E. J., Bao, G. & Porteus, M. H. Quantifying on- and off-target genome editing. Trends Biotechnol. 33, 132–140 (2015).
Qiu, P. et al. Mutation detection using Surveyor nuclease. Biotechniques 36, 702–707 (2004).
Brinkman, E. K. et al. Easy quantification of template-directed CRISPR/Cas9 editing. Nucleic Acids Res. 46, e58 (2018).
Pinello, L. et al. Analyzing CRISPR genome-editing experiments with CRISPResso. Nat. Biotechnol. 34, 695–697 (2016).
Porteus, M. H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003).
Szczepek, M. et al. Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat. Biotechnol. 25, 786–793 (2007).
Wang, H. et al. CRISPR-mediated live imaging of genome editing and transcription. Science 365, 1301–1305 (2019).
Liu, Y. et al. Very fast CRISPR on demand. Science 368, 1265–1269 (2020).
Li, A. et al. AAV-CRISPR gene editing is negated by pre-existing immunity to Cas9. Mol. Ther. 28, 1432–1441 (2020).
Charlesworth, C. T. et al. Identification of preexisting adaptive immunity to Cas9 proteins in humans. Nat. Med. 25, 249–254 (2019).
Wagner, D. L. et al. High prevalence of Streptococcus pyogenes Cas9-reactive T cells within the adult human population. Nat. Med. 25, 242–248 (2019).
Bindra, R. S., Goglia, A. G., Jasin, M. & Powell, S. N. Development of an assay to measure mutagenic non-homologous end-joining repair activity in mammalian cells. Nucleic Acids Res. 41, e115–e115 (2013).
Lin, Y., Lukacsovich, T. & Waldman, A. S. Multiple pathways for repair of DNA double-strand breaks in mammalian chromosomes. Mol. Cell. Biol. 19, 8353–8360 (1999).
Gaj, T., Gersbach, C. A. & Barbas, C. F. 3rd ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013).
van Haasteren, J., Li, J., Scheideler, O. J., Murthy, N. & Schaffer, D. V. The delivery challenge: fulfilling the promise of therapeutic genome editing. Nat. Biotechnol. 38, 845–855 (2020).
Yin, H., Kauffman, K. J. & Anderson, D. G. Delivery technologies for genome editing. Nat. Rev. Drug Discov. 16, 387–399 (2017).
Pinkham, K. et al. Stearoyl CoA desaturase is essential for regulation of endoplasmic reticulum homeostasis and tumor growth in glioblastoma cancer stem cells. Stem Cell Rep. 12, 712–727 (2019).
Huang, F., Mazina, O. M., Zentner, I. J., Cocklin, S. & Mazin, A. V. Inhibition of homologous recombination in human cells by targeting RAD51 recombinase. J. Med. Chem. 55, 3011–3020 (2012).
Leahy, J. J. et al. Identification of a highly potent and selective DNA-dependent protein kinase (DNA-PK) inhibitor (NU7441) by screening of chromenone libraries. Bioorg. Med. Chem. Lett. 14, 6083–6087 (2004).
Verhaegent, M. & Christopoulos, T. K. Recombinant Gaussia luciferase. Overexpression, purification, and analytical application of a bioluminescent reporter for DNA hybridization. Anal. Chem. 74, 4378–4385 (2002).
Thompson, E. M., Nagata, S. & Tsuji, F. I. Vargula hilgendorfii luciferase: a secreted reporter enzyme for monitoring gene expression in mammalian cells. Gene 96, 257–262 (1990).
Vouillot, L., Thelie, A. & Pollet, N. Comparison of T7E1 and surveyor mismatch cleavage assays to detect mutations triggered by engineered nucleases. G3 (Bethesda) 5, 407–415 (2015).
Guell, M., Yang, L. & Church, G. M. Genome editing assessment using CRISPR Genome Analyzer (CRISPR-GA). Bioinformatics 30, 2968–2970 (2014).
Degeling, M. H., Maguire, C. A., Bovenberg, M. S. & Tannous, B. A. Sensitive assay for mycoplasma detection in mammalian cell culture. Anal. Chem. 84, 4227–4232 (2012).
Shimomura, O. & Johnson, F. H. Mechanisms in the quantum yield of Cypridina bioluminescence. Photochem. Photobiol. 12, 291–295 (1970).
Lundholt, B. K., Scudder, K. M. & Pagliaro, L. A simple technique for reducing edge effect in cell-based assays. J. Biomol. Screen. 8, 566–570 (2003).
Szymczak, A. L. et al. Correction of multi-gene deficiency in vivo using a single ‘self-cleaving’ 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Acknowledgements
We thank members of the Lai Lab and the Badr Lab for the very helpful discussions. This work was supported by the Ministry of Science and Technology (MOST) grants (104-2320-B-007-005-MY2, 106-2320-B-007-004-MY3, 109-2628-B-001-017 to C.P.-K.L.), Academia Sinica Innovative Materials and Analysis Technology Exploration (i-MATE) Program (AS-iMATE-107-33 to C.P.-K.L.), Academia Sinica Career Development Award (AS-CDA-109-M04 C.P.-K.L.), the National Institutes of Health (K22CA197053 and R01NS113822 to C.E.B.) and the American Brain Tumor Association (ABTA) Discovery Grant supported by the Uncle Kory Foundation to C.E.B.
Author information
Authors and Affiliations
Contributions
C.P.-K.L. and C.E.B. conceived and designed the study. J.C.C. conducted the in vitro experiments and developed the procedures. C.E.B developed the in vivo procedures. The manuscript was written by J.C.C., C.E.B. and C.P.-K.L. with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A provisional patent application on the BLRR reporter has been submitted.
Additional information
Peer review information Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Chien, J. C. et al. Nucleic Acids Res. 48, e100 (2020): https://doi.org/10.1093/nar/gkaa669
Supplementary information
Rights and permissions
About this article
Cite this article
Chien, J.CY., Badr, C.E. & Lai, C.PK. Multiplexed bioluminescence-mediated tracking of DNA double-strand break repairs in vitro and in vivo. Nat Protoc 16, 3933–3953 (2021). https://doi.org/10.1038/s41596-021-00564-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00564-8
This article is cited by
-
Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
Nature Communications (2024)
-
FSP1 inhibition enhances olaparib sensitivity in BRCA-proficient ovarian cancer patients via a nonferroptosis mechanism
Cell Death & Differentiation (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.