Abstract
Measurements of neuronal activity across brain areas are important for understanding the neural correlates of cognitive and motor processes such as attention, decision-making and action selection. However, techniques that allow cellular resolution measurements are expensive and require a high degree of technical expertise, which limits their broad use. Wide-field imaging of genetically encoded indicators is a high-throughput, cost-effective and flexible approach to measure activity of specific cell populations with high temporal resolution and a cortex-wide field of view. Here we outline our protocol for assembling a wide-field macroscope setup, performing surgery to prepare the intact skull and imaging neural activity chronically in behaving, transgenic mice. Further, we highlight a processing pipeline that leverages novel, cloud-based methods to analyze large-scale imaging datasets. The protocol targets laboratories that are seeking to build macroscopes, optimize surgical procedures for long-term chronic imaging and/or analyze cortex-wide neuronal recordings. The entire protocol, including steps for assembly and calibration of the macroscope, surgical preparation, imaging and data analysis, requires a total of 8 h. It is designed to be accessible to laboratories with limited expertise in imaging methods or interest in high-throughput imaging during behavior.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw datasets used to generate the visual sign, stimuli triggered averages and linear regression analysis maps are available in a public repository, maintained by Cold Spring Harbor Laboratory with https://doi.org/10.14224/1.38599. Example datasets to test the analysis pipeline are at http://labshare.cshl.edu/shares/library/repository/38599/2021-01-20-Update/.
Code availability
Source code used in this protocol is available in the online repositories without access restrictions under a general public license at https://github.com/churchlandlab. The code and NeuroCAAS platform will remain available for the foreseeable future.
References
Stringer, C. et al. Spontaneous behaviors drive multidimensional, brainwide activity. Science 364, 255 (2019).
Vanni, M. P., Chan, A. W., Balbi, M., Silasi, G. & Murphy, T. H. Mesoscale mapping of mouse cortex reveals frequency-dependent cycling between distinct macroscale functional modules. J. Neurosci. 37, 7513–7533 (2017).
Clancy, K. B., Orsolic, I. & Mrsic-Flogel, T. D. Locomotion-dependent remapping of distributed cortical networks. Nat. Neurosci. 22, 778–786 (2019).
Musall, S., Kaufman, M. T., Juavinett, A. L., Gluf, S. & Churchland, A. K. Single-trial neural dynamics are dominated by richly varied movements. Nat. Neurosci. 22, 1677–1686 (2019).
Allen, W. E. et al. Global representations of goal-directed behavior in distinct cell types of mouse neocortex. Neuron 94, 891–907.e6 (2017).
Pinto, L. et al. Task-dependent changes in the large-scale dynamics and necessity of cortical regions. Neuron 104, 810–824.e9 (2019).
Zatka-Haas, P., Steinmetz, N. A., Carandini, M. & Harris, K. D. A perceptual decision requires sensory but not action coding in mouse cortex. Preprint at bioRxiv https://doi.org/10.1101/501627 (2020).
Salkoff, D. B., Zagha, E., McCarthy, E. & McCormick, D. A. Movement and performance explain widespread cortical activity in a visual detection task. Cereb. Cortex 30, 421–437 (2020).
Grinvald, A., Lieke, E., Frostig, R. D., Gilbert, C. D. & Wiesel, T. N. Functional architecture of cortex revealed by optical imaging of intrinsic signals. Nature 324, 361–364 (1986).
Frostig, R. D., Lieke, E. E., Ts’o, D. Y. & Grinvald, A. Cortical functional architecture and local coupling between neuronal activity and the microcirculation revealed by in vivo high-resolution optical imaging of intrinsic signals. Proc. Natl Acad. Sci. USA 87, 6082–6086 (1990).
Shmuel, A. & Grinvald, A. Functional organization for direction of motion and its relationship to orientation maps in cat area 18. J. Neurosci. 16, 6945–6964 (1996).
Garrett, M. E., Nauhaus, I., Marshel, J. H. & Callaway, E. M. Topography and areal organization of mouse visual cortex. J. Neurosci. 34, 12587–12600 (2014).
Andermann, M. L., Kerlin, A. M., Roumis, D. K., Glickfeld, L. L. & Reid, R. C. Functional specialization of mouse higher visual cortical areas. Neuron 72, 1025–1039 (2011).
Juavinett, A. L., Nauhaus, I., Garrett, M. E., Zhuang, J. & Callaway, E. M. Automated identification of mouse visual areas with intrinsic signal imaging. Nat. Protoc. 12, 32–43 (2017).
Ferezou, I. et al. Spatiotemporal dynamics of cortical sensorimotor integration in behaving mice. Neuron 56, 907–923 (2007).
Akemann, W., Mutoh, H., Perron, A., Rossier, J. & Knöpfel, T. Imaging brain electric signals with genetically targeted voltage-sensitive fluorescent proteins. Nat. Methods 7, 643–649 (2010).
Wekselblatt, J. B., Flister, E. D., Piscopo, D. M. & Niell, C. M. Large-scale imaging of cortical dynamics during sensory perception and behavior. J. Neurophysiol. 115, 2852–2866 (2016).
Gilad, A., Gallero-Salas, Y., Groos, D. & Helmchen, F. Behavioral strategy determines frontal or posterior location of short-term memory in neocortex. Neuron 99, 814–828.e7 (2018).
Guo, Z. V. et al. Flow of cortical activity underlying a tactile decision in mice. Neuron 81, 179–194 (2014).
Silasi, G., Xiao, D., Vanni, M. P., Chen, A. C. N. & Murphy, T. H. Intact skull chronic windows for mesoscopic wide-field imaging in awake mice. J Neurosci Methods 267, 141–149 (2016).
Cunningham, J. P. & Yu, B. M. Dimensionality reduction for large-scale neural recordings. Nat. Neurosci. 17, 1500–1509 (2014).
Buchanan, E. K. et al. Penalized matrix decomposition for denoising, compression, and improved demixing of functional imaging data. Preprint at https://arxiv.org/abs/1807.06203 (2018).
Saxena, S. et al. Localized semi-nonnegative matrix factorization (LocaNMF) of widefield calcium imaging data. PLoS Comput. Biol. 16, e1007791 (2020).
Kim, E. J., Juavinett, A. L., Kyubwa, E. M., Jacobs, M. W. & Callaway, E. M. Three types of cortical layer 5 neurons that differ in brain-wide connectivity and function. Neuron 88, 1253–1267 (2015).
Matho, K. S. et al. Genetic dissection of glutamatergic neuron subpopulations and developmental trajectories in the cerebral cortex. Preprint at bioRxiv https://doi.org/10.1101/2020.04.22.054064 (2020).
Murphy, T. H. et al. High-throughput automated home-cage mesoscopic functional imaging of mouse cortex. Nat. Commun. 7, 11611 (2016).
Murphy, T. H. et al. Automated task training and longitudinal monitoring of mouse mesoscale cortical circuits using home cages. eLife 9, e55964 (2020).
Cramer, J. V. et al. In vivo widefield calcium imaging of the mouse cortex for analysis of network connectivity in health and brain disease. NeuroImage 199, 570–584 (2019).
Shimaoka, D., Harris, K. D. & Carandini, M. Effects of arousal on mouse sensory cortex depend on modality. Cell Rep. 25, 3230 (2018).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. eLife 5, e12727 (2016).
Xiao, D. et al. Mapping cortical mesoscopic networks of single spiking cortical or sub-cortical neurons. eLife 6, e19976 (2017).
Deffieux, T., Demene, C., Pernot, M. & Tanter, M. Functional ultrasound neuroimaging: a review of the preclinical and clinical state of the art. Curr. Opin. Neurobiol. 50, 128–135 (2018).
Lake, E. M. R. et al. Simultaneous cortex-wide fluorescence Ca 2+ imaging and whole-brain fMRI. Nat. Methods 17, 1262–1271 (2020).
Shemesh, O. A. et al. Precision calcium imaging of dense neural populations via a cell body-targeted calcium indicator. Preprint at bioRxiv https://doi.org/10.1101/773069 (2019).
Abe, T., Kinsella, I., Saxena, S., Paninski, L. & Cunningham, J. P. Neuroscience cloud analysis as a service. Preprint at bioRxiv https://doi.org/10.1101/2020.06.11.146746 (2020).
Ratzlaff, E. H. & Grinvald, A. A tandem-lens epifluorescence macroscope: hundred-fold brightness advantage for wide-field imaging. J. Neurosci. Methods 36, 127–137 (1991).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Ma, Y. et al. Wide-field optical mapping of neural activity and brain haemodynamics: considerations and novel approaches. Phil. Trans. R. Soc. B 371, 20150360 (2016).
Musall, S., Haiss, F., Weber, B. & von der Behrens, W. Deviant processing in the primary somatosensory cortex. Cereb. Cortex 27, 863–876 (2015).
Lerner, T. N. et al. Intact-brain analyses reveal distinct information carried by SNc dopamine subcircuits. Cell 162, 635–647 (2015).
Valley, M. T. et al. Separation of hemodynamic signals from GCaMP fluorescence measured with wide-field imaging. J. Neurophysiol. 123, 356–366 (2019).
Vanni, M. P. & Murphy, T. H. Mesoscale transcranial spontaneous activity mapping in GCaMP3 transgenic mice reveals extensive reciprocal connections between areas of somatomotor cortex. J. Neurosci. 34, 15931–15946 (2014).
Xiao, D. et al. Mapping cortical mesoscopic networks of single spiking cortical or sub-cortical neurons. eLife. 6, e19976 (2017).
Kim, T. H. et al. Long-term optical access to an estimated one million neurons in the live mouse cortex. Cell Rep. 17, 3385–3394 (2016).
Ghanbari, L. et al. Cortex-wide neural interfacing via transparent polymer skulls. Nat. Commun. 10, 1500 (2019).
Steinmetz, N. A., Zatka-Haas, P., Carandini, M. & Harris, K. D. Distributed coding of choice, action and engagement across the mouse brain. Nature 576, 266–273 (2019).
Steinmetz, N. A. et al. Aberrant cortical activity in multiple GCaMP6-expressing transgenic mouse lines. eNeuro https://doi.org/10.1523/ENEURO.0207-17.2017 (2017).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Gilad, A. & Helmchen, F. Spatiotemporal refinement of signal flow through association cortex during learning. Nat. Commun. 11, 1–14 (2020).
Montgomery, M. K. et al. Glioma-induced alterations in neuronal activity and neurovascular coupling during disease progression. Cell Rep. 31, 107500 (2020).
Acknowledgements
We thank M. Kaufman and K. Odoemene for help with developing early versions of the protocol; P. Gupta, F. Albeanu and J. Wekselblatt for technical advice; N. Steinmetz, M. Pachitariu and K. Harris for help with wide-field analysis; and Z. Josh Huang for providing FezF2 mice. Financial support was received from the Swiss National Science foundation (S.M., grant no. P2ZHP3_161770), the Deutsche Forschungsgemeinschaft (German Research Foundation, DFG - 368482240/GRK2416), the NIH (grant no. EY R01EY022979 and BRAIN initiative 5R01EB026949) and the Army Research Office under contract no. W911NF-16-1-0368 as part of the collaboration between the US DOD, the UK MOD and the UK Engineering and Physical Research Council under the Multidisciplinary University Research Initiative (A.K.C.). X.R.S. was supported by the NINDS BRAIN Initiative of the National Institutes of Health under award number F32MH120888. T.A. was supported by NIH training grant 2T32NS064929-11. S.S. was supported by the Swiss National Science Foundation P400P2 186759 and NIH 5U19NS104649. J.P.C. was supported by Simons 542963 and the McKnight Foundation. L.P. was funded by IARPA MICRONS D16PC00003, NIH 5U01NS103489, 5U19NS104649, 5U19NS107613, 1UF1NS107696, 1UF1NS108213, 1RF1MH120680, DARPA NESD N66001-17-C-4002 and Simons Foundation 543023. L.P. and J.P.C. were supported by NSF Neuronex Award DBI-1707398.
Author information
Authors and Affiliations
Contributions
S.M. and A.K.C. conceptualized early versions of the procedures. S.M. and S.G. implemented early setup versions and acquisition workflow. S.M., X.R.S. and J.C. refined surgical procedures. All authors refined the macroscope building procedures and compiled the required part lists. S.M. and J.C. wrote acquisition software. S.M., X.R.S. and S.G. prepared animals and acquired data in the expected results. J.C., S.M., I.K. and S.S. wrote software for analysis. T.A., I.K., S.S., J.P.C. and L.P conceptualized the general analysis workflow. T.A., I.K. and S.S. deployed software on NeuroCAAS with input from J.C. and S.M.. A.K., J.C. and S.M. prepared figures. J.C., S.M., S.G., A.K. and A.K.C. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Ariel Gilad and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Musall, S. et al. Nat. Neurosci. 22, 1677–1686 (2019): https://doi.org/10.1038/s41593-019-0502-4
Extended data
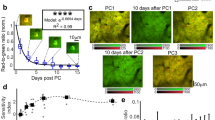
Extended Data Fig. 1 Photobleaching due to repeated or high-power imaging.
a, Photochemical degradation of the fluorophore can occur over days because of repeated imaging. Decrease in fluorescence can be observed after 10 min when imaging at more than 50 mW of blue light (blue trace); signals are more stable at lower intensities (black, cyan traces). b, Top: images from a mouse expressing GCaMP6 in a subpopulation of cortical excitatory projection neurons. Visible decrease in overall fluorescence is evident after daily imaging at day 5 and 10. Bottom: histogram of pixel intensities corresponding to the images above.
Extended Data Fig. 2 Steps for setup calibration.
a, Alignment of the excitation dichroic. b, Alignment of the emission dichroic using Camware. c, Procedure for obtaining uniform illumination with the Camware software and the line profiler. The WidefieldImager also has a calibration mode with similar functionality and supporting cameras from multiple vendors.
Supplementary information
Rights and permissions
About this article
Cite this article
Couto, J., Musall, S., Sun, X.R. et al. Chronic, cortex-wide imaging of specific cell populations during behavior. Nat Protoc 16, 3241–3263 (2021). https://doi.org/10.1038/s41596-021-00527-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00527-z
This article is cited by
-
Efficient coding of natural images in the mouse visual cortex
Nature Communications (2024)
-
Diverse and asymmetric patterns of single-neuron projectome in regulating interhemispheric connectivity
Nature Communications (2024)
-
Distributed context-dependent choice information in mouse posterior cortex
Nature Communications (2023)
-
Pyramidal cell types drive functionally distinct cortical activity patterns during decision-making
Nature Neuroscience (2023)
-
All-optical interrogation of neural circuits in behaving mice
Nature Protocols (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.