Abstract
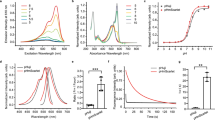
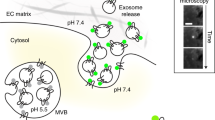
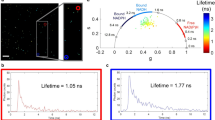
Endocytosis is a fundamental process occurring in all eukaryotic cells. Live cell imaging of endocytosis has helped to decipher many of its mechanisms and regulations. With the pulsed-pH (ppH) protocol, one can detect the formation of individual endocytic vesicles (EVs) with an unmatched temporal resolution of 2 s. The ppH protocol makes use of cargo protein (e.g., the transferrin receptor) coupled to a pH-sensitive fluorescent protein, such as superecliptic pHluorin (SEP), which is brightly fluorescent at pH 7.4 but not fluorescent at pH <6.0. If the SEP moiety is at the surface, its fluorescence will decrease when cells are exposed to a low pH (5.5) buffer. If the SEP moiety has been internalized, SEP will remain fluorescent even during application of the low pH buffer. Fast perfusion enables the complete exchange of low and high pH extracellular solutions every 2 s, defining the temporal resolution of the technique. Unlike other imaging-based endocytosis assays, the ppH protocol detects EVs without a priori hypotheses on the dynamics of vesicle formation. Here, we explain how the ppH protocol quantifies the endocytic activity of living cells and the recruitment of associated proteins in real time. We provide a step-by-step procedure for expression of the reporter proteins with transient transfection, live cell image acquisition with synchronized pH changes and automated analysis. The whole protocol can be performed in 2 d to provide quantitative information on the endocytic process being studied.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
The MATLAB programs that are used to analyze the ppH data are written for MATLAB2018a with the following MATLAB toolboxes: Image Processing, Wavelet, Statistics and Machine Learning; the programs are formatted as a toolbox, scission_analysis, available at MATLAB Central File Exchange as 72744-scission_analysis (https://fr.mathworks.com/matlabcentral/fileexchange/72744-scission_analysis).
References
Sigismund, S. et al. Endocytosis and signaling: cell logistics shape the eukaryotic cell plan. Physiol. Rev. 92, 273–366 (2012).
Thottacherry, J. J., Sathe, M., Prabhakara, C. & Mayor, S. Spoiled for choice: diverse endocytic pathways function at the cell surface. Annu. Rev. Cell Dev. Biol. 35, 55–84 (2019).
Roth, T. F. & Porter, K. R. Yolk protein uptake in the oocyte of the mosquito Aedes aegypti L. J. Cell Biol. 20, 313–332 (1964).
Kaksonen, M. & Roux, A. Mechanisms of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 19, 313–326 (2018).
Johannes, L., Parton, R. G., Bassereau, P. & Mayor, S. Building endocytic pits without clathrin. Nat. Rev. Mol. Cell Biol. 16, 311–321 (2015).
Watanabe, S. & Boucrot, E. Fast and ultrafast endocytosis. Curr. Opin. Cell Biol. 47, 64–71 (2017).
Haucke, V. & Kozlov, M. M. Membrane remodeling in clathrin-mediated endocytosis. J. Cell Sci. 131, jcs216812 (2018).
Rosendale, M. & Perrais, D. Imaging in focus: imaging the dynamics of endocytosis. Int. J. Biochem. Cell Biol. 93, 41–45 (2017).
Robinson, M. S. Forty years of clathrin-coated vesicles. Traffic 16, 1210–1238 (2015).
Anderson, R. G. W., Brown, M. S. & Goldstein, J. L. Role of the coated endocytic vesicle in the uptake of receptor-bound low density lipoprotein in human fibroblasts. Cell 10, 351–364 (1977).
Betz, W., Mao, F. & Bewick, G. Activity-dependent fluorescent staining and destaining of living vertebrate motor nerve terminals. J. Neurosci. 12, 363–375 (1992).
Bitsikas, V., Corrêa, I. R. & Nichols, B. J. Clathrin-independent pathways do not contribute significantly to endocytic flux. eLife 3, e03970 (2014).
Hopkins, C. R. & Trowbridge, I. S. Internalization and processing of transferrin and the transferrin receptor in human carcinoma A431 cells. J. Cell Biol. 97, 508–521 (1983).
Lamb, J. E., Ray, F., Ward, J. H., Kushner, J. P. & Kaplan, J. Internalization and subcellular localization of transferrin and transferrin receptors in HeLa cells. J. Biol. Chem. 258, 8751–8758 (1983).
Miesenböck, G., De Angelis, D. A. & Rothman, J. E. Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature 394, 192–195 (1998).
Sankaranarayanan, S., De Angelis, D., Rothman, J. E. & Ryan, T. A. The use of pHluorins for optical measurements of presynaptic activity. Biophys. J. 79, 2199–2208 (2000).
Yudowski, G. A. et al. Real-time imaging of discrete exocytic events mediating surface delivery of AMPA receptors. J. Neurosci. 27, 11112–11121 (2007).
Balaji, J. & Ryan, T. A. Single-vesicle imaging reveals that synaptic vesicle exocytosis and endocytosis are coupled by a single stochastic mode. Proc. Natl Acad. Sci. U. S. A. 104, 20576–20581 (2007).
Jullié, D., Choquet, D. & Perrais, D. Recycling endosomes undergo rapid closure of a fusion pore on exocytosis in neuronal dendrites. J. Neurosci. 34, 11106–11118 (2014).
Merrifield, C. J., Perrais, D. & Zenisek, D. Coupling between clathrin-coated-pit invagination, cortactin recruitment, and membrane scission observed in live cells. Cell 121, 593–606 (2005).
Taylor, M. J., Perrais, D. & Merrifield, C. J. A high precision survey of the molecular dynamics of mammalian clathrin-mediated endocytosis. PLoS Biol. 9, e1000604 (2011).
Taylor, M. J., Lampe, M. & Merrifield, C. J. A feedback loop between dynamin and actin recruitment during clathrin-mediated endocytosis. PLoS Biol. 10, e1001302 (2012).
Cauvin, C. et al. Rab35 GTPase triggers switch-like recruitment of the Lowe Syndrome lipid phosphatase OCRL on newborn endosomes. Curr. Biol. 26, 120–128 (2016).
Antonny, B. et al. Membrane fission by dynamin: what we know and what we need to know. EMBO J. 35, 2270–2284 (2016).
Rosendale, M. et al. Functional recruitment of dynamin requires multimeric interactions for efficient endocytosis. Nat. Commun. 10, 4462 (2019).
Jullié, D. et al. A discrete presynaptic vesicle cycle for neuromodulator receptors. Neuron 105, 663–677.e8 (2020).
Kirchhausen, T. Imaging endocytic clathrin structures in living cells. Trends Cell Biol. 19, 596–605 (2009).
Mattheyses, A. L., Simon, S. M. & Rappoport, J. Z. Imaging with total internal reflection fluorescence microscopy for the cell biologist. J. Cell Sci. 123, 3621–3628 (2010).
Rosendale, M., Jullié, D., Choquet, D. & Perrais, D. Spatial and temporal regulation of receptor endocytosis in neuronal dendrites revealed by imaging of single vesicle formation. Cell Rep. 18, 1840–1847 (2017).
Shen, Y., Rosendale, M., Campbell, R. E. & Perrais, D. pHuji, a pH-sensitive red fluorescent protein for imaging of exo- and endocytosis. J. Cell Biol. 207, 419–432 (2014).
Lampe, M., Pierre, F., Al-Sabah, S., Krasel, C. & Merrifield, C. J. Dual single-scission event analysis of constitutive transferrin receptor (TfR) endocytosis and ligand-triggered β2-adrenergic receptor (β2AR) or Mu-opioid receptor (MOR) endocytosis. Mol. Biol. Cell 25, 3070–3080 (2014).
Martineau, M. et al. Semisynthetic fluorescent pH sensors for imaging exocytosis and endocytosis. Nat. Commun. 8, 1412 (2017).
Hua, Y. et al. A readily retrievable pool of synaptic vesicles. Nat. Neurosci. 14, 833–839 (2011).
Chen, M., Van Hook, M. J., Zenisek, D. & Thoreson, W. B. Properties of ribbon and non-ribbon release from rod photoreceptors revealed by visualizing individual synaptic vesicles. J. Neurosci. 33, 2071–2086 (2013).
Fujii, S., Tanaka, H. & Hirano, T. Detection and characterization of individual endocytosis of AMPA-type glutamate receptor around postsynaptic membrane. Genes Cells 22, 583–590 (2017).
Sathe, M. et al. Small GTPases and BAR domain proteins regulate branched actin polymerisation for clathrin and dynamin-independent endocytosis. Nat. Commun. 9, 1835 (2018).
Rathje, M. et al. AMPA receptor pHluorin-GluA2 reports NMDA receptor-induced intracellular acidification in hippocampal neurons. Proc. Natl Acad. Sci. U. S. A. 110, 14426–14431 (2013).
Perrais, D., Kleppe, I. C., Taraska, J. W. & Almers, W. Recapture after exocytosis causes differential retention of protein in granules of bovine chromaffin cells. J. Physiol. 560, 413–428 (2004).
Kavalali, E. T. & Jorgensen, E. M. Visualizing presynaptic function. Nat. Neurosci. 17, 10–16 (2013).
Saffarian, S., Cocucci, E. & Kirchhausen, T. Distinct dynamics of endocytic clathrin-coated pits and coated plaques. PLoS Biol. 7, e1000191 (2009).
Mettlen, M., Loerke, D., Yarar, D., Danuser, G. & Schmid, S. L. Cargo- and adaptor-specific mechanisms regulate clathrin-mediated endocytosis. J. Cell Biol. 188, 919–933 (2010).
Caldieri, G. et al. Reticulon 3–dependent ER-PM contact sites control EGFR nonclathrin endocytosis. Science 356, 617–624 (2017).
Boucrot, E. et al. Endophilin marks and controls a clathrin-independent endocytic pathway. Nature 517, 460–465 (2015).
Lakshminarayan, R. et al. Galectin-3 drives glycosphingolipid-dependent biogenesis of clathrin-independent carriers. Nat. Cell Biol. 16, 595–606 (2014).
Renard, H.-F. et al. Endophilin-A2 functions in membrane scission in clathrin-independent endocytosis. Nature 517, 493–496 (2015).
Kononenko, N. L. et al. Clathrin/AP-2 mediate synaptic vesicle reformation from endosome-like vacuoles but are not essential for membrane retrieval at central synapses. Neuron 82, 981–988 (2014).
Watanabe, S. et al. Ultrafast endocytosis at mouse hippocampal synapses. Nature 504, 242–247 (2013).
Lu, J. et al. Postsynaptic positioning of endocytic zones and AMPA receptor cycling by physical coupling of dynamin-3 to Homer. Neuron 55, 874–889 (2007).
Baschieri, F. et al. Frustrated endocytosis controls contractility-independent mechanotransduction at clathrin-coated structures. Nat. Commun. 9, 3825 (2018).
Morel, M., Bartolo, D., Galas, J.-C., Dahan, M. & Studer, V. Microfluidic stickers for cell- and tissue-based assays in microchannels. Lab Chip 9, 1011–1013 (2009).
Courson, D. S. & Rock, R. S. Fast benchtop fabrication of laminar flow chambers for advanced microscopy techniques. PLoS One 4, e6479 (2009).
Acknowledgements
This article is dedicated to Christien James Merrifield (1972–2017), who was instrumental in the development of the ppH protocol. The authors thank Arnaud Rodriguez (Bordeaux Neurocampus) for taking photographs of the imaging and perfusion setup. This work was supported by the Centre National de la Recherche Scientifique (Interface program), the Fondation Recherche Médicale (FRM ING20101221208) and the Agence Nationale pour la Recherche (CaPeBlE ANR-12-BSV5-005 and LocalEndoProbes ANR-17-CE16-0012) to D.P., the FRM, a pre-doctoral fellowship from the University of Bordeaux to M.R. and Labex BRAIN fellowships to M.R. and L.C.
Author information
Authors and Affiliations
Contributions
D.P. designed the experiments. S.S., M.R., L.C., T.N.N.V., D.J. and D.P. performed many experiments that led to this version of the protocol. D.P. wrote most of the analysis software, with contributions from M.R. and D.J. D.P. and S.S. wrote the manuscript, and all authors edited it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Emanuele Cocucci, Derek Toomre and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related Links
Key references using this protocol
Taylor, M. J., Perrais, D. & Merrifield, C. J. PloS Biol. 9, e1000604 (2011): https://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.1000604
Rosendale, M., Jullié, D., Choquet, D. & Perrais, D. Cell Rep. 18, 1840–1847 (2017): https://www.sciencedirect.com/science/article/pii/S2211124717301663
Rosendale, M. et al. Nat. Comm. 10, 4462 (2019): https://www.nature.com/articles/s41467-019-12434-9
Extended data
Extended Data Fig. 1 Description of the perfusion setup for the ppH protocol.
a. General view of a setup equipped with local perfusion for the ppH protocol. One can see (1) the peristaltic pump for perfusion of the chamber with a controlled flow, (2) the syringes containing HBS (pH 7.4) and MBS (pH 5.5) solutions (two syringes in the front). The 5 µm pore size filters and stopcocks are visible under the syringes. The two syringes in the back are optional: they may contain a compound to apply to cells. (3) motorized micromanipulator for positioning the application pipette. (4) XY stage of the microscope (Olympus IX71). b. Close up view on the application pipette and open chamber. (1) three way electrovalves (2) pipette holder (3) glass pipette dipped in the solution (open chamber) (4) heated holder with two blue heating elements and (5) in line solution heater for warming up the solution, and (6) suction needle for evacuation of excess solution. c. Diagram of the perfusion setup. For application of HBS/MBS, the stopcocks of the first two syringes are open while the ones of the other two (HBS/MBS+compound, blue) are closed. To apply the compound, the first two stopcocks are closed and the other two are open.
Extended Data Fig. 2 Effects of ill positioned application pipette on the imaging at pH 7.4 and 5.5 during the ppH protocol.
Images of a COS7 cell transfected with TfR-SEP during the ppH protocol under the perfusion of HBS at pH 7.4 (top images) or MBS at pH 5.5 (bottom images). All images are shown with the same scale. In (a), the positioning is optimal. Note the even fluorescence at pH 7.4 which reflects equilibrium (compare with images in transition from one solution to the other, Figure 2b). At pH 5.5, no homogenous fluorescence is visible, only some dots (corresponding to vesicles) are visible. In (b-f), the application pipette was moved in the directions indicated by the bottom diagrams on the three axes. Values are indicated in µm. For these positions, the exchange is not correct. Either the fluorescence of the ‘pH 7.4’ image is too low (d,e) or the fluorescence of the ‘pH 5.5’ image is too high (b,c,f). Scale bar 10 µm.
Supplementary information
Supplementary Information
User Manual for scission_analysis, the MATLAB toolbox described for analysis of ppH experiments and Supplementary Table 1, listing the files generated by scission_analysis and included in the Supplementary Dataset.
Supplementary Video 1
Fast imaging (10 Hz) of a HeLa cell transfected with TfR-SEP with exchange of solutions at pH 7.4 and 5.5, played at real time. Scale bar: 10 µm.
Supplementary Video 2
TetraSpeck beads imaged at 50 Hz with TIRF illumination (473-nm laser with a predicted penetration depth of 80 nm with a 150×, 1.45 NA Olympus objective). Note that the immobile beads attached to the coverslip are brightly fluorescent. They appear saturated to better see the moving beads in solution. The moving beads appear very transiently as they enter the evanescent field and disappear as they leave it.
Supplementary Video 3
092-1_TfR5.stk generated as described in Step 27, accelerated 40 times. Scale bar: 10 µm.
Supplementary Video 4
092-1_TfR7.stk generated as described in Step 27, accelerated 40 times. Scale bar: 10 µm.
Supplementary Video 5
092-1_dyn5.stk generated as described in Step 27, accelerated 40 times. Scale bar: 10 µm.
Supplementary Video 6
092-1_dyn7.stk generated as described in Step 27, accelerated 40 times. Scale bar: 10 µm.
Supplementary Video 7
Merged 092-1_TfR5.stk (green) and 092-1_TfR7.stk (magenta) showing the sites of scission that are located at CCS, and therefore appear white.
Supplementary Data
Raw imaging files and analysis files generated by scission_analysis. The list of files generated and a quick description are provided in Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Sposini, S., Rosendale, M., Claverie, L. et al. Imaging endocytic vesicle formation at high spatial and temporal resolutions with the pulsed-pH protocol. Nat Protoc 15, 3088–3104 (2020). https://doi.org/10.1038/s41596-020-0371-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0371-z
This article is cited by
-
Endocytosis in cancer and cancer therapy
Nature Reviews Cancer (2023)
-
Endocytosis in the axon initial segment maintains neuronal polarity
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.