Abstract
The therapeutic response to immune checkpoint blockade (ICB) is highly variable, not only between different cancers but also between patients with the same cancer type. The biological mechanisms underlying these differences in response are incompletely understood. Identifying correlates in patient tumor samples is challenging because of genetic and environmental variability. Murine studies usually compare different tumor models or treatments, introducing potential confounding variables. This protocol describes bilateral murine tumor models, derived from syngeneic cancer cell lines, that display a symmetrical yet dichotomous response to ICB. These models enable detailed analysis of whole tumors in a highly homogeneous background, combined with knowledge of the therapeutic outcome within a few weeks, and could potentially be used for mechanistic studies using other (immuno-)therapies. We discuss key considerations and describe how to use two cell lines as fully optimized models. We discuss experimental details, including proper inoculation technique to achieve symmetry and one-sided surgical tumor removal, which takes only 5 min per mouse. Furthermore, we outline the preparation of bulk tissue or single-cell suspensions for downstream analyses such as bulk RNA-seq, immunohistochemistry, single-cell RNA-seq and flow cytometry.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Data availability
The raw RNA-seq data used to generate Fig. 6 are available from the Gene Expression Omnibus data repository (accession no. GSE117358).
References
Galluzzi, L., Chan, T. A., Kroemer, G., Wolchok, J. D. & López-Soto, A. The hallmarks of successful anticancer immunotherapy. Sci. Transl. Med. 10, eaat7807 (2018).
Lesterhuis, W. J. et al. Dynamic versus static biomarkers in cancer immune checkpoint blockade: unravelling complexity. Nat. Rev. Drug Discov. 16, 264 (2017).
Topalian, S. L. et al. Safety, activity, and immune correlates of anti–PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non–small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
Curran, M. A., Montalvo, W., Yagita, H. & Allison, J. P. PD-1 and CTLA-4 combination blockade expands infiltrating T cells and reduces regulatory T and myeloid cells within B16 melanoma tumors. Proc. Natl Acad. Sci. 107, 4275 (2010).
Wolchok, J. D. et al. Nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 369, 122–133 (2013).
Das, R. et al. Combination therapy with anti–CTLA-4 and anti–PD-1 leads to distinct immunologic changes in vivo. J. Immunol. 194, 950–959 (2015).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568 (2014).
Ayers, M. et al. IFN-γ–related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest 127, 2930–2940 (2017).
Prat, A. et al. Immune-related gene expression profiling after PD-1 blockade in non–small cell lung carcinoma, head and neck squamous cell carcinoma, and melanoma. Cancer Res. 77, 3540–3550 (2017).
Sanmamed, M., Chester, C., Melero, I. & Kohrt, H. Defining the optimal murine models to investigate immune checkpoint blockers and their combination with other immunotherapies. Ann. Oncol. 27, 1190–1198 (2016).
Gustafsson, C. & Vallverdú, J. The best model of a cat is several cats. Trends Biotechnol. 34, 207–213 (2016).
Gubin, M. M. et al. High-dimensional analysis delineates myeloid and lymphoid compartment remodeling during successful immune-checkpoint cancer therapy. Cell 175, 1014–1030.e1019 (2018).
Moynihan, K. D. et al. Eradication of large established tumors in mice by combination immunotherapy that engages innate and adaptive immune responses. Nat. Med. 22, 1402 (2016).
Wei, S. C. et al. Distinct cellular mechanisms underlie anti-CTLA-4 and anti-PD-1 checkpoint blockade. Cell 170, 1120–1133.e1117 (2017).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Mosely, S. I. S. et al. Rational selection of syngeneic preclinical tumor models for immunotherapeutic drug discovery. Cancer Immunol. Res. 5, 29–41 (2017).
Lesterhuis, W. J. et al. Network analysis of immunotherapy-induced regressing tumours identifies novel synergistic drug combinations. Sci. Rep. 5, 12298 (2015).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015).
Sutmuller, R. P. et al. Synergism of cytotoxic T lymphocyte–associated antigen 4 blockade and depletion of CD25+ regulatory T cells in antitumor therapy reveals alternative pathways for suppression of autoreactive cytotoxic T lymphocyte responses. J. Exp. Med. 194, 823–832 (2001).
Zemek, R. M. et al. Sensitization to immune checkpoint blockade through activation of a STAT1/NK axis in the tumor microenvironment. Sci. Transl. Med. 11, eaav7816 (2019).
Ock, C.-Y. et al. Genomic landscape associated with potential response to anti-CTLA-4 treatment in cancers. Nat. Commun. 8, 1050 (2017).
Lee, H.-S. et al. Comprehensive immunoproteogenomic analyses of malignant pleural mesothelioma. JCI Insight 3, e98575 (2018).
Jerby-Arnon, L. et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 175, 984–997 (2018). e924.
van Elsas, A., Hurwitz, A. A. & Allison, J. P. Combination immunotherapy of B16 melanoma using anti–cytotoxic T lymphocyte–associated antigen 4 (Ctla-4) and granulocyte/macrophage colony-stimulating factor (Gm-Csf)-producing vaccines induces rejection of subcutaneous and metastatic tumors accompanied by autoimmune depigmentation. J. Exp. Med. 190, 355–366 (1999).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Ribas, A. et al. Association of response to programmed death receptor 1 (PD-1) blockade with pembrolizumab (MK-3475) with an interferon-inflammatory immune gene signature. J. Clin. Oncol. 33 (Suppl.), 3001 (2015).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Yarchoan, M., Hopkins, A. & Jaffee, E. M. Tumor mutational burden and response rate to PD-1 inhibition. N. Engl. J. Med. 377, 2500–2501 (2017).
Sneddon, S. et al. Whole exome sequencing of an asbestos-induced wild-type murine model of malignant mesothelioma. BMC Cancer 17, 396–396 (2017).
Ben-David, U. et al. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560, 325–330 (2018).
Gibney, G. T., Weiner, L. M. & Atkins, M. B. Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 17, e542–e551 (2016).
Hegde, P. S., Karanikas, V. & Evers, S. The where, the when, and the how of immune monitoring for cancer immunotherapies in the era of checkpoint inhibition. Clin. Cancer Res. 22, 1865–1874 (2016).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Devaud, C., John, L. B., Westwood, J. A., Darcy, P. K. & Kershaw, M. H. Immune modulation of the tumor microenvironment for enhancing cancer immunotherapy. OncoImmunology 2, e25961 (2013).
Santegoets, S. J. et al. The anatomical location shapes the immune infiltrate in tumors of same etiology and affects survival. Clin. Cancer Res. 25, 240–252 (2019).
Klein, S. L. & Flanagan, K. L. Sex differences in immune responses. Nat. Rev. Immunol. 16, 626–638 (2016).
Kovacs, E. J. et al. Aging and innate immunity in the mouse: impact of intrinsic and extrinsic factors. Trends Immunol. 30, 319–324 (2009).
Macpherson, A. J. & Harris, N. L. Interactions between commensal intestinal bacteria and the immune system. Nat. Rev. Immunol. 4, 478–485 (2004).
Tsukamoto, A., Serizawa, K., Sato, R., Yamazaki, J. & Inomata, T. Vital signs monitoring during injectable and inhalant anesthesia in mice. Exp. Anim. 64, 57–64 (2015).
Workman, P. et al. Guidelines for the welfare and use of animals in cancer research. Br. J. Cancer 102, 1555–1577 (2010).
Acknowledgements
We acknowledge the facilities, as well as the scientific and technical assistance of the Australian Microscopy & Microanalysis Research Facility at the Centre for Microscopy, Characterisation & Analysis, The University of Western Australia, a facility funded by the University and State and Commonwealth governments. We thank P. Deng and C. Rinaldi for technical assistance. This work was funded by grant 1103980 from the National Health and Medical Research Council (NHMRC). W.J.L. was funded by an NHMRC RD Wright Fellowship and a Fellowship from the Cancer Council of Western Australia.
Author information
Authors and Affiliations
Contributions
R.M.Z. optimized the mouse models, performed and interpreted all mouse experiments and flow cytometry experiments, prepared samples for RNA-seq analysis, and wrote the paper. V.S.F., C.F., and T.H.C. provided technical assistance with surgery and in vivo treatment experiments. E.d.J., T.L. and A.B. interpreted the bulk RNA-seq analyses. L.B. provided critical reagents. R.A.L., M.J.M., and A.K.N. were involved in experimental design and interpretation of all in vivo studies. R.A.L. and V.S.F provided critical revision of the manuscript. W.J.L. designed and developed the models, interpreted the data and supervised the project. R.M.Z and W.J.L. wrote the manuscript with input from all authors. All authors gave final approval of the version to be published.
Corresponding authors
Ethics declarations
Competing interests
A patent application pertaining to aspects of this work has been filed by R.M.Z., W.J.L., R.A.L., and A.B. (PCT/AU2019/050259, “Method for immunotherapy drug treatment”). A.K.N., M.J.M., A.B., R.A.L., and W.J.L. have received research funding and consultancy fees from Douglas Pharmaceuticals. W.J.L. has received research funding from AstraZeneca and consultancy fees from Merck Sharp & Dohme. A.K.N. declares consultancy or advisory board membership for Boehringer Ingelheim, Bayer Pharmaceuticals, Roche Pharmaceutics, Merck Sharp Dohme, Pharmabcine, Atara Biotherapeutics and Trizell, and research funding from AstraZeneca. M.J.M. has served on advisory boards for Merck Sharp & Dohme, Bristol-Myers Squibb, Roche, and AstraZeneca.
Additional information
Peer review information Nature Protocols thanks Gottfried Baier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Zemek, R. M. et al. Sci. Transl. Med. 11, eaav7816 (2019): https://doi.org/10.1126/scitranslmed.aav7816
Lesterhuis, W. J. et al. Sci. Rep. 5, 12298 (2015): https://doi.org/10.1038/srep12298
Integrated supplementary information
Supplementary Figure 1 Response to ICB in the B16-F10 tumor model.
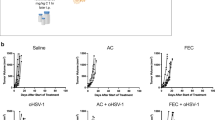
a) Growth curves of C57BL/6 mice bearing subcutaneous B16-F10 tumours, treated with i.p. PBS or (b) with ICB (anti-CTLA4 day 5 and anti-PD-L1 on days 5,7 and 9 post-inoculation) when tumours were at least 9mm2 (n=15 per arm, combined data from 3 experiments).
Supplementary Figure 2 Flow cytometry gating strategy.
Gating strategy used to delineate immune cell subsets in mouse tumour and lymph nodes.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2 and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Zemek, R.M., Fear, V.S., Forbes, C. et al. Bilateral murine tumor models for characterizing the response to immune checkpoint blockade. Nat Protoc 15, 1628–1648 (2020). https://doi.org/10.1038/s41596-020-0299-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0299-3
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.