Abstract
Multiple aspects of neural activity, from neuronal firing to neuromodulator release and signaling, underlie brain function and ultimately shape animal behavior. The recently developed and constantly growing toolbox of genetically encoded sensors for neural activity, including calcium, voltage, neurotransmitter and neuromodulator sensors, allows precise measurement of these signaling events with high spatial and temporal resolution. Here, we describe the engineering, characterization and application of our recently developed dLight1, a suite of genetically encoded dopamine (DA) sensors based on human inert DA receptors. dLight1 offers high molecular specificity, requisite affinity and kinetics and great sensitivity for measuring DA release in vivo. The detailed workflow described in this protocol can be used to systematically characterize and validate dLight1 in increasingly intact biological systems, from cultured cells to acute brain slices to behaving mice. For tool developers, we focus on characterizing five distinct properties of dLight1: dynamic range, affinity, molecular specificity, kinetics and interaction with endogenous signaling; for end users, we provide comprehensive step-by-step instructions for how to leverage fiber photometry and two-photon imaging to measure dLight1 transients in vivo. The instructions provided in this protocol are designed to help laboratory personnel with a broad range of experience (at the graduate or post-graduate level) to develop and utilize novel neuromodulator sensors in vivo, by using dLight1 as a benchmark.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All DNA plasmids and viruses mentioned in this protocol can be obtained from either the Tian laboratory at UC Davis or Addgene under a materials transfer agreement. All data present in this article are available from the authors upon request. Implemented and curated computer codes have been deposited in github (https://github.com/GradinaruLab/dLight1/).
References
Marder, E. Neuromodulation of neuronal circuits: back to the future. Neuron 76, 1–11 (2012).
Lee, S. H. & Dan, Y. Neuromodulation of brain states. Neuron 76, 209–222 (2012).
Tritsch, N. X. & Sabatini, B. L. Dopaminergic modulation of synaptic transmission in cortex and striatum. Neuron 76, 33–50 (2012).
Rusakov, D. A., Savtchenko, L. P., Zheng, K. & Henley, J. M. Shaping the synaptic signal: molecular mobility inside and outside the cleft. Trends Neurosci. 34, 359–369 (2011).
Cachope, R. & Cheer, J. F. Local control of striatal dopamine release. Front. Behav. Neurosci. 8, 188 (2014).
Kehr, J. & Yoshitake, T. Monitoring molecules in neuroscience: historical overview and current advancements. Front. Biosci. (Elite Ed.) 5, 947–954 (2013).
Wightman, R. M. Detection technologies. Probing cellular chemistry in biological systems with microelectrodes. Science 311, 1570–1574 (2006).
Darvesh, A. S. et al. In vivo brain microdialysis: advances in neuropsychopharmacology and drug discovery. Expert Opin. Drug Discov. 6, 109–127 (2011).
Park, J., Takmakov, P. & Wightman, R. M. In vivo comparison of norepinephrine and dopamine release in rat brain by simultaneous measurements with fast-scan cyclic voltammetry. J. Neurochem. 119, 932–944 (2011).
Bucher, E. S. & Wightman, R. M. Electrochemical analysis of neurotransmitters. Annu. Rev. Anal. Chem. (Palo Alto Calif.) 8, 239–261 (2015).
Jaquins-Gerstl, A. & Michael, A. C. A review of the effects of FSCV and microdialysis measurements on dopamine release in the surrounding tissue. Analyst 140, 3696–3708 (2015).
Muller, A., Joseph, V., Slesinger, P. A. & Kleinfeld, D. Cell-based reporters reveal in vivo dynamics of dopamine and norepinephrine release in murine cortex. Nat. Methods 11, 1245–1252 (2014).
Lee, D. et al. Temporally precise labeling and control of neuromodulatory circuits in the mammalian brain. Nat. Methods 14, 495–503 (2017).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nat. Methods 6, 875–881 (2009).
Patriarchi, T. et al. Ultrafast neuronal imaging of dopamine dynamics with designed genetically encoded sensors. Science 360, eaat4422 (2018).
Venkatakrishnan, A. J. et al. Molecular signatures of G-protein-coupled receptors. Nature 494, 185–194 (2013).
Manglik, A. & Kruse, A. C. Structural basis for G protein-coupled receptor activation. Biochemistry 56, 5628–5634 (2017).
Svoboda, K. & Yasuda, R. Principles of two-photon excitation microscopy and its applications to neuroscience. Neuron 50, 823–839 (2006).
Dana, H. et al. High-performance GFP-based calcium indicators for imaging activity in neuronal populations and microcompartments. Nat. Methods 16, 649–657 (2019).
Abdelfattah, A. S. et al. Bright and photostable chemigenetic indicators for extended in vivo voltage imaging. Science 365, 699–704 (2019).
Piatkevich, K. D. et al. Population imaging of neural activity in awake behaving mice in multiple brain regions. Nature 574, 413–417 (2019).
Broussard, G. J. et al. In vivo measurement of afferent activity with axon-specific calcium imaging. Nat. Neurosci. 21, 1272–1280 (2018).
Marvin, J. S. et al. An optimized fluorescent probe for visualizing glutamate neurotransmission. Nat. Methods 10, 162–170 (2013).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Jing, M. et al. A genetically encoded fluorescent acetylcholine indicator for in vitro and in vivo studies. Nat. Biotechnol. 36, 726–737 (2018).
Sun, F. et al. A genetically encoded fluorescent sensor enables rapid and specific detection of dopamine in flies, fish, and mice. Cell 174, 481–496.e19 (2018).
Feng, J. et al. A genetically encoded fluorescent sensor for rapid and specific in vivo detection of norepinephrine. Neuron 102, 745–761.e8 (2019).
Mingote, S. et al. Functional connectome analysis of dopamine neuron glutamatergic connections in forebrain regions. J. Neurosci. 35, 16259–16271 (2015).
de Jong, J. W. et al. A neural circuit mechanism for encoding aversive stimuli in the mesolimbic dopamine system. Neuron 101, 133–151.e7 (2018).
Corre, J. et al. Dopamine neurons projecting to medial shell of the nucleus accumbens drive heroin reinforcement. eLife 7, e39945 (2018).
Dong, H. et al. Dorsal striatum dopamine levels fluctuate across the sleep-wake cycle and respond to salient stimuli in mice. Front. Neurosci. 13, 242 (2019).
Zhang, Y. et al. Capping of the N-terminus of PSD-95 by calmodulin triggers its postsynaptic release. EMBO J. 33, 1341–1353 (2014).
Thomas, P. & Smart, T. G. HEK293 cell line: a vehicle for the expression of recombinant proteins. J. Pharmacol. Toxicol. Methods 51, 187–200 (2005).
Tsvetanova, N. G. & von Zastrow, M. Spatial encoding of cyclic AMP signaling specificity by GPCR endocytosis. Nat. Chem. Biol. 10, 1061–1065 (2014).
Thorne, N., Inglese, J. & Auld, D. S. Illuminating insights into firefly luciferase and other bioluminescent reporters used in chemical biology. Chem. Biol. 17, 646–657 (2010).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. eLife 5, e12727 (2016).
Goodman, O. B. et al. Beta-arrestin acts as a clathrin adaptor in endocytosis of the beta2-adrenergic receptor. Nature 383, 447–450 (1996).
Vickery, R. G. & von Zastrow, M. Distinct dynamin-dependent and -independent mechanisms target structurally homologous dopamine receptors to different endocytic membranes. J. Cell Biol. 144, 31–43 (1999).
Irannejad, R. et al. Conformational biosensors reveal GPCR signalling from endosomes. Nature 495, 534–538 (2013).
Lee, Y. B., Glover, C. P., Cosgrave, A. S., Bienemann, A. & Uney, J. B. Optimizing regulatable gene expression using adenoviral vectors. Exp. Physiol. 90, 33–37 (2005).
Schnütgen, F. et al. A directional strategy for monitoring Cre-mediated recombination at the cellular level in the mouse. Nat. Biotechnol. 21, 562–565 (2003).
Aschauer, D. F., Kreuz, S. & Rumpel, S. Analysis of transduction efficiency, tropism and axonal transport of AAV serotypes 1, 2, 5, 6, 8 and 9 in the mouse brain. PLoS ONE 8, e76310 (2013).
Grimm, D. et al. In vitro and in vivo gene therapy vector evolution via multispecies interbreeding and retargeting of adeno-associated viruses. J. Virol. 82, 5887–5911 (2008).
Tervo, D. G. et al. A designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92, 372–382 (2016).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Challis, R. C. et al. Systemic AAV vectors for widespread and targeted gene delivery in rodents. Nat. Protoc. 14, 379–414 (2019).
Aurnhammer, C. et al. Universal real-time PCR for the detection and quantification of adeno-associated virus serotype 2-derived inverted terminal repeat sequences. Hum. Gene Ther. Methods 23, 18–28 (2012).
Nassi, J. J., Cepko, C. L., Born, R. T. & Beier, K. T. Neuroanatomy goes viral. Front. Neuroanat. 9, 80 (2015).
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007).
Lo, L. & Anderson, D. J. A Cre-dependent, anterograde transsynaptic viral tracer for mapping output pathways of genetically marked neurons. Neuron 72, 938–950 (2011).
Chatterjee, S. et al. Nontoxic, double-deletion-mutant rabies viral vectors for retrograde targeting of projection neurons. Nat. Neurosci. 21, 638–646 (2018).
Helmchen, F. & Denk, W. Deep tissue two-photon microscopy. Nat. Methods 2, 932–940 (2005).
Palij, P. & Stamford, J. A. Real-time monitoring of endogenous noradrenaline release in rat brain slices using fast cyclic voltammetry: 3. Selective detection of noradrenaline efflux in the locus coeruleus. Brain Res. 634, 275–282 (1994).
John, C. E. & Jones, S. R. in Electrochemical Methods for Neuroscience (eds Michael, A. C. & Borland, L. M.) (CRC Press/Taylor & Francis, Boca Raton, FL, 2007).
Bull, D. R. et al. Application of fast cyclic voltammetry to measurement of electrically evoked dopamine overflow from brain slices in vitro. J. Neurosci. Methods 32, 37–44 (1990).
Courtney, N. A. & Ford, C. P. The timing of dopamine- and noradrenaline-mediated transmission reflects underlying differences in the extent of spillover and pooling. J. Neurosci. 34, 7645–7656 (2014).
Xie, T., McCann, U. D., Kim, S., Yuan, J. & Ricaurte, G. A. Effect of temperature on dopamine transporter function and intracellular accumulation of methamphetamine: implications for methamphetamine-induced dopaminergic neurotoxicity. J. Neurosci. 20, 7838–7845 (2000).
Gunaydin, L. A. et al. Natural neural projection dynamics underlying social behavior. Cell 157, 1535–1551 (2014).
Sparta, D. R. et al. Construction of implantable optical fibers for long-term optogenetic manipulation of neural circuits. Nat. Protoc. 7, 12–23 (2011).
Lerner, T. N. et al. Intact-brain analyses reveal distinct information carried by SNc dopamine subcircuits. Cell 162, 635–647 (2015).
Kim, C. K. et al. Simultaneous fast measurement of circuit dynamics at multiple sites across the mammalian brain. Nat. Methods 13, 325–328 (2016).
Goldey, G. J. et al. Removable cranial windows for long-term imaging in awake mice. Nat. Protoc. 9, 2515–2538 (2014).
Asaad, W. F., Santhanam, N., McClellan, S. & Freedman, D. J. High-performance execution of psychophysical tasks with complex visual stimuli in MATLAB. J. Neurophysiol. 109, 249–260 (2013).
Owen, S. F. & Kreitzer, A. C. An open-source control system for in vivo fluorescence measurements from deep-brain structures. J. Neurosci. Methods 311, 170–177 (2019).
Chen, Y. et al. NS21: re-defined and modified supplement B27 for neuronal cultures. J. Neurosci. Methods 171, 239–247 (2008).
Franklin, K. & Paxinos, G. Paxinos and Franklin’s the Mouse Brain in Stereotaxic Coordinates, Compact: The Coronal Plates and Diagrams (Academic Press, 2019).
Menegas, W., Babayan, B. M., Uchida, N. & Watabe-Uchida, M. Opposite initialization to novel cues in dopamine signaling in ventral and posterior striatum in mice. eLife 6, e21886 (2017).
al, J. R. C. E. Dorsal raphe dopamine neurons modulate arousal and promote wakefulness by salient stimuli. Neuron 94, 1205–1219 (2017).
Klapoetke, N. C. et al. Independent optical excitation of distinct neural populations. Nat. Methods 11, 338–346 (2014).
Kibbe, W. A. OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res. 35, W43–W46 (2007).
Acknowledgements
This work was supported by NIH BRAIN Initiative grants U01NS090604, U01NS013522, DP2MH107056 and U01NS103571 (L.T.); grants DP2NS083038, R01NS085938 and P30CA014195 (A.N.); BRAIN Initiative grants U01NS013522 (J.T.W. and M.v.Z.), and NIH grant DP2NS087949 and NIH/NIA grant R01AG047664 (V.G.). K.M. is a DFG research fellow and recipient of a Catharina Foundation postdoctoral scholar award. V.G. is a Heritage Principal Investigator supported by the Heritage Medical Research Institute.
Author information
Authors and Affiliations
Contributions
T.P. and L.T. wrote the manuscript with contributions from J.R.C. and V.G. (fiber photometry and optogenetics), G.J.B. (rAAV preparation and cloning), R.L. (structural modeling), A.M. and M.v.Z. (TIRF microscopy, FACS and cAMP measurements), K.M. and A.N. (in vivo two-photon imaging in behaving mice), and J.W. (ex vivo two-photon imaging).
Corresponding authors
Ethics declarations
Competing interests
T.P. and L.T. are co-inventors on a patent application (WO/2018/098262A1) for the technology described in this paper. L.T. is the co-founder of Seven Biosciences.
Additional information
Peer review information Nature Protocols thanks Thomas Knopfel and other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related link
Key references using this protocol
Patriarchi, T. et al. Science 360, eaat4422 (2018) https://science.sciencemag.org/content/360/6396/eaat4422/
Integrated supplementary information
Supplementary Fig. 1 Design of GPCR-based sensors using the universal cpGFP module.
Shown is a sequence alignment of the contiguous regions of transmembrane helixes 5 and 6 (TM5, TM6) used to develop fluorescent sensors from 10 different GPCRs. The third intracellular loop of the receptors is deleted in the sensors (region shown in orange shading) and replaced with the universal cpGFP module consisting of the indicated residues surrounding cpGFP (indicated in the green box). The complete sequence for the cpGFP module is available from Addgene or in our original publication16). Adapted with permission from16, American Association for the Advancement of Science.
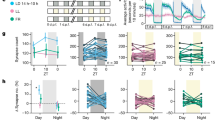
Supplementary Fig. 2 Comparison of locomotion-related dopamine transients using dLight1.1 or dLight1.2 in behaving mice with two-photon microscopy.
a, Schematics of two-photon imaging of head-fixed mouse during treadmill locomotion. b, Mean ΔF/F for all significant positive-going transients in mice expressing either dLight1.1 (n = 2 mice, 131 transients) or dLight1.2 (n = 2 mice, 31 transients). c-d, Mean transient ΔF/F during rest and run for all fields in dLight1.1 (n = 2 mice, 5 fields) and dLight1.2 (n = 2 mice, 8 fields). Adapted with permission from16, American Association for the Advancement of Science.
Supplementary Fig. 3 Comparison between dLight1.3b and GRAB-DA1m in neurons.
a, Representative images of primary cultured neurons (DIV15) expressing either dLight1.3b or GRAB-DA1m. Cells were transduced by directly applying ~1010 viral particles either AAV9.Synapsin.dLight1.3b or AAV9.Synapsin.GRAB-DA1m onto the medium 14 d prior to imaging (AAVs produced by the UC Davis Viral Vector Core Facility). Cells were imaged under identical conditions (laser intensity, pinhole size, etc.) with a 40x oil-based objective on a Zeiss 710 confocal microscope. Intensity profile plots representative of the lines drawn across the neuronal dendrites are show as insets. Scale bars, 20 µm. b, Basal fluorescence and signal to noise calculations were performed on Fiji (Image J) from regions of interest manually drawn on the neuronal membrane. At least n=3 neurons from 2 separate experiments were included in the analysis. Data are shown as individual ROI values ± SEM. ****p<0.0001, n.s.= not significant, unpaired student’s t-test. c, Time-lapse images were acquired at ~1 second intervals. Drugs (dopamine, DA; haloperidol, Halo; SCH23390, SCH) were directly applied on the cells at the time points and concentrations indicated in the graphs. n≥2 neurons for each sensor. Data are shown as mean ± SEM.
Supplementary Fig. 4 Equipment setup for photometry recordings.
a, Photograph of optical components in the fiber photometry setup described in 51 and 18. b, Photograph of data processor with inputs from photodetector and external TTL and with outputs for LED modulation. Optical components in (a) are located inside the rackbox below. c, Photograph of a mouse expressing dLight1.1 during freely moving behavior inside an operant box. This water-deprived mouse is obtaining sucrose water reward from lickometer. All procedures were performed in accordance with the guidelines of the National Institutes of Health and were approved by the Institutional Animal Care and Use Committee (IACUC) and by the Office of Laboratory Animal Resources at the California Institute of Technology.
Supplementary Fig. 5 Example raw photometry traces.
a, Raw photometry signals demodulated from 490-nm (blue) and 405-nm (violet) channel. Notice photobleaching in both channels. b, Photometry signals after 405-nm signal is fitted to 490-nm signal by applying a least-squares fit. c, ΔF/F trace is calculated for each recording session as. (490-nm signal – fitted 405-nm signal)/fitted 405-nm signal. Notice this can effectively remove the contribution of photobleaching and potential contamination from motion artifacts. All procedures were performed in accordance with the guidelines of the National Institutes of Health and were approved by the Institutional Animal Care and Use Committee (IACUC) and by the Office of Laboratory Animal Resources at the California Institute of Technology.
Supplementary Fig. 6 Equipment setup of perfusion system for titrations on cells.
a, The perfusion system setup for performing neuromodulator titrations during one-photon imaging of sensor fluorescence is based on an inverted confocal microscope (7). A display (1) connected with a digital controller (2) for the eight-valve perfusion system (3), an eight-channel perfusion inlet (4) and a single channel outlet (5) connected to a peristaltic pump (6) for removal of excess buffer from the dish. b, Close up view of the perfusion inlet and outlet setup on the stage adaptor containing the imaged dish.
Supplementary information
Supplementary Information
Supplementary Figures 1–6
Rights and permissions
About this article
Cite this article
Patriarchi, T., Cho, J.R., Merten, K. et al. Imaging neuromodulators with high spatiotemporal resolution using genetically encoded indicators. Nat Protoc 14, 3471–3505 (2019). https://doi.org/10.1038/s41596-019-0239-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0239-2
This article is cited by
-
Genetically encoded sensors enable micro- and nano-scopic decoding of transmission in healthy and diseased brains
Molecular Psychiatry (2021)
-
A versatile GPCR toolkit to track in vivo neuromodulation: not a one-size-fits-all sensor
Neuropsychopharmacology (2021)
-
Spatial decoding of endosomal cAMP signals by a metastable cytoplasmic PKA network
Nature Chemical Biology (2021)
-
A complete pupillometry toolbox for real-time monitoring of locus coeruleus activity in rodents
Nature Protocols (2020)
-
An expanded palette of dopamine sensors for multiplex imaging in vivo
Nature Methods (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.