Abstract
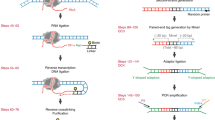
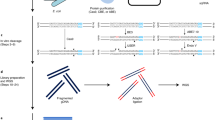
RNA–chromatin interactions represent an important aspect of the transcriptional regulation of genes and transposable elements. However, analyses of chromatin-associated RNAs (caRNAs) are often limited to one caRNA at a time. Here, we describe the iMARGI (in situ mapping of RNA–genome interactome) technique, which is used to discover caRNAs and reveal their respective genomic interaction loci. iMARGI starts with in situ crosslinking and genome fragmentation, followed by converting each proximal RNA–DNA pair into an RNA–linker–DNA chimeric sequence. These chimeric sequences are subsequently converted into a sequencing library suitable for paired-end sequencing. A standardized bioinformatic software package, iMARGI-Docker, is provided to decode the paired-end sequencing data into caRNA–DNA interactions. Compared to its predecessor MARGI (mapping RNA–genome interactions), the number of input cells for iMARGI is 3–5 million (a 100-fold reduction), experimental time is reduced, and clear checkpoints have been established. It takes a few hours a day and a total of 8 d to complete the construction of an iMARGI sequencing library and 1 d to carry out data processing with iMARGI-Docker.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
An iMARGI dataset has been deposited into the NCBI Sequence Read Archive (SRA) under accession no. SRR8206679.
Code availability
The iMARGI-Docker software and its documentation are available at: https://sysbio.ucsd.edu/imargi_pipeline. The software is completely open source, under the BSD 2 license. The source code is available at https://github.com/Zhong-Lab-UCSD/iMARGI-Docker. The pre-built Docker image can be pulled from the Docker Hub. The version used in this paper is v.1.1.1.
References
Nguyen, T. C., Zaleta-Rivera, K., Huang, X., Dai, X. & Zhong, S. RNA, Action through Interactions. Trends Genet. 34, 867–882 (2018).
Penny, G. D., Kay, G. F., Sheardown, S. A., Rastan, S. & Brockdorff, N. Requirement for Xist in X chromosome inactivation. Nature 379, 131–137 (1996).
Yang, F. et al. The lncRNA Firre anchors the inactive X chromosome to the nucleolus by binding CTCF and maintains H3K27me3 methylation. Genome Biol. 16, 52 (2015).
Graf, M. et al. Telomere length determines TERRA and R-loop regulation through the cell cycle. Cell 170, 72–85.e14 (2017).
Yu, R., Wang, X. & Moazed, D. Epigenetic inheritance mediated by coupling of RNAi and histone H3K9 methylation. Nature 558, 615–619 (2018).
Watanabe, T. et al. Role for piRNAs and noncoding RNA in de novo DNA methylation of the imprinted mouse Rasgrf1 locus. Science 332, 848–852 (2011).
Miao, Y. et al. Enhancer-associated long non-coding RNA LEENE regulates endothelial nitric oxide synthase and endothelial function. Nat. Commun. 9, 292 (2018).
Place, R. F., Li, L. C., Pookot, D., Noonan, E. J. & Dahiya, R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc. Natl Acad. Sci. USA 105, 1608–1613 (2008).
Morris, K. V., Chan, S. W., Jacobsen, S. E. & Looney, D. J. Small interfering RNA-induced transcriptional gene silencing in human cells. Science 305, 1289–1292 (2004).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Simon, M. D. et al. The genomic binding sites of a noncoding RNA. Proc. Natl Acad. Sci. USA 108, 20497–20502 (2011).
Chu, C., Qu, K., Zhong, F. L., Artandi, S. E. & Chang, H. Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol. Cell 44, 667–678 (2011).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Sridhar, B. et al. Systematic mapping of RNA-chromatin interactions in vivo. Curr. Biol. 27, 602–609 (2017).
Yan, Z. et al. Genome-wide co-localization of RNA-DNA interactions and fusion RNA pairs. Proc. Natl Acad. Sci. USA 116, 3328–3337 (2019).
Bell, J. C. et al. Chromatin-associated RNA sequencing (ChAR-seq) maps genome-wide RNA-to-DNA contacts. eLife 7, e27024 (2018).
Li, X. et al. GRID-seq reveals the global RNA–chromatin interactome. Nat. Biotechnol. 35, 940–950 (2017).
Chen, W. et al. RNAs as proximity-labeling media for identifying nuclear speckle positions relative to the genome. iScience 4, 204–215 (2018).
Yin, Y. et al. U1 snRNP regulates chromatin retention of noncoding RNAs. Preprint at bioRxiv, https://www.biorxiv.org/content/10.1101/310433v1 (2018).
Sun, G. et al. A bias-reducing strategy in profiling small RNAs using Solexa. RNA 17, 2256–2262 (2011).
Jayaprakash, A. D., Jabado, O., Brown, B. D. & Sachidanandam, R. Identification and remediation of biases in the activity of RNA ligases in small-RNA deep sequencing. Nucleic Acids Res. 39, e141 (2011).
Sorefan, K. et al. Reducing ligation bias of small RNAs in libraries for next generation sequencing. Silence 3, 4 (2012).
Li, W., Freudenberg, J. & Miramontes, P. Diminishing return for increased Mappability with longer sequencing reads: implications of the k-mer distributions in the human genome. BMC Bioinforma. 15, 2 (2014).
Zhao, J. et al. Genome-wide identification of polycomb-associated RNAs by RIP-seq. Mol. Cell 40, 939–953 (2010).
Hadjiolov, A. A., Tencheva, Z. S. & Bojadjieva-Mikhailova, A. G. Isolation and some characteristics of cell nuclei from brain cortex of adult cat. J. Cell Biol. 26, 383–393 (1965).
Suzuki, K., Bose, P., Leong-Quong, R. Y., Fujita, D. J. & Riabowol, K. REAP: a two minute cell fractionation method. BMC Res. Notes 3, 294 (2010).
Nabbi, A. & Riabowol, K. Isolation of nuclei. Cold Spring Harb. Protoc. 2015, 731–734 (2015).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Dekker, J. et al. The 4D nucleome project. Nature 549, 219–226 (2017).
Quinlan, A. R. & Hall, I. M. J. B. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Abdennur, N. & Mirny, L. Cooler: scalable storage for Hi-C data and other genomically-labeled arrays. Bioinformatics https://doi.org/10.1093/bioinformatics/btz540 (2019).
Kerpedjiev, P. et al. HiGlass: web-based visual exploration and analysis of genome interaction maps. Genome Biol. 19, 125 (2018).
Cao, X., Yan, Z., Wu, Q., Zheng, A. & Zhong, S. GIVE: portable genome browsers for personal websites. Genome Biol. 19, 92 (2018).
Acknowledgements
We thank X. Wen for help with graphics; 4DN DCIC (http://dcic.4dnucleome.org) for discussions on the data processing pipeline; A. Zheng, A. Kaul, K. Faizi, N. Moshiri, R. Zhang, J. Ma, J. Chen, X. Huang and Z. Zhang for testing the iMARGI-Docker software; and K. Sriram and A. Chen for proofreading. This work was funded by DP1HD087990 (to S.Z.), NIH 4D Nucleome U01CA200147 (to S.C. and S.Z.), and NIH R00HL122368 and R01HL145170 (to Z.C.).
Author information
Authors and Affiliations
Contributions
W.W. and S.Z. designed the research; W.W., T.C.N. and S.Z. developed the experimental method and protocol; Z.Y. and S.Z. developed the computational method and data analysis tools; W.W., Z.Y. and S.Z. wrote the manuscript; Z.C. and S.C. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.Z. is a cofounder of Genemo Inc.
Additional information
Peer review information Nature Protocols thanks Michiel de Hoon and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Sridhar, B. et al. Curr. Biol. 27, 602–609 (2017) https://doi.org/10.1016/j.cub.2017.01.011
Yan, Z. et al. Proc. Natl Acad. Sci. USA 116, 3328–3337 (2019) https://doi.org/10.1073/pnas.1819788116
Chen, W. et al. iScience 4, 204–215 (2018) https://doi.org/10.1016/j.isci.2018.06.005
Supplementary information
Rights and permissions
About this article
Cite this article
Wu, W., Yan, Z., Nguyen, T.C. et al. Mapping RNA–chromatin interactions by sequencing with iMARGI. Nat Protoc 14, 3243–3272 (2019). https://doi.org/10.1038/s41596-019-0229-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0229-4
This article is cited by
-
Single-cell multiplex chromatin and RNA interactions in ageing human brain
Nature (2024)
-
Genome-wide analysis of the interplay between chromatin-associated RNA and 3D genome organization in human cells
Nature Communications (2023)
-
The 4D Nucleome Data Portal as a resource for searching and visualizing curated nucleomics data
Nature Communications (2022)
-
Stress-induced RNA–chromatin interactions promote endothelial dysfunction
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.