Abstract
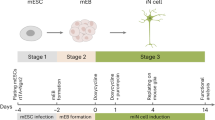
The enteric nervous system (ENS) represents a vast network of neuronal and glial cell types that develops entirely from migratory neural crest (NC) progenitor cells. Considerable improvements in the understanding of the molecular mechanisms underlying NC induction and regional specification have recently led to the development of a robust method to re-create the process in vitro using human pluripotent stem cells (hPSCs). Directing the fate of hPSCs toward the enteric NC (ENC) results in an accessible and scalable in vitro model of ENS development. The application of hPSC-derived enteric neural lineages provides a powerful platform for ENS-related disease modeling and drug discovery. Here we present a detailed protocol for the induction of a regionally specific NC intermediate that occurs over the course of a 15-d interval and is an effective source for the in vitro derivation of functional enteric neurons (ENs) from hPSCs. Additionally, we introduce a new and improved protocol that we have developed to optimize the protocol for future applications in regenerative medicine, in which components of undefined activity have been replaced with fully defined culture conditions. This protocol provides access to a broad range of human ENS lineages within a 30-d period.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data generated or analyzed during this study are included in this published article (and its Supplementary Information files).
References
Shyamala, K., Yanduri, S., Girish, H. C. & Murgod, S. Neural crest: the fourth germ layer. J. Oral Maxillofac. Pathol. 19, 221–229 (2015).
Crane, J. F. & Trainor, P. A. Neural crest stem and progenitor cells. Annu. Rev. Cell Dev. Biol. 22, 267–286 (2006).
Thomas, S. et al. Human neural crest cells display molecular and phenotypic hallmarks of stem cells. Hum. Mol. Genet. 17, 3411–3425 (2008).
Chan, K. K. et al. Hoxb3 vagal neural crest-specific enhancer element for controlling enteric nervous system development. Dev. Dyn. 233, 473–483 (2005).
Fu, M., Chi Hang Lui, V., Har Sham, M., Nga Yin Cheung, A. & Kwong Hang Tam, P. HOXB5 expression is spatially and temporarily regulated in human embryonic gut during neural crest cell colonization and differentiation of enteric neuroblasts. Dev. Dyn. 228, 1–10 (2003).
Heanue, T. A. & Pachnis, V. Enteric nervous system development and Hirschsprung’s disease: advances in genetic and stem cell studies. Nat. Rev. Neurosci. 8, 466–479 (2007).
Cheeseman, B. L., Zhang, D., Binder, B. J., Newgreen, D. F. & Landman, K. A. Cell lineage tracing in the developing enteric nervous system: superstars revealed by experiment and simulation. J. R. Soc. Interface 11, 20130815 (2014).
Wallace, A. S. & Burns, A. J. Development of the enteric nervous system, smooth muscle and interstitial cells of Cajal in the human gastrointestinal tract. Cell Tissue Res. 319, 367–382 (2005).
Kim, J., Lo, L., Dormand, E. & Anderson, D. J. SOX10 maintains multipotency and inhibits neuronal differentiation of neural crest stem cells. Neuron 38, 17–31 (2003).
Lasrado, R. et al. Lineage-dependent spatial and functional organization of the mammalian enteric nervous system. Science 356, 722–726 (2017).
Lee, G., Chambers, S. M., Tomishima, M. J. & Studer, L. Derivation of neural crest cells from human pluripotent stem cells. Nat. Protoc. 5, 688–701 (2010).
Mica, Y., Lee, G., Chambers, S. M., Tomishima, M. J. & Studer, L. Modeling neural crest induction, melanocyte specification, and disease-related pigmentation defects in hESCs and patient-specific iPSCs. Cell Rep. 3, 1140–1152 (2013).
Fattahi, F., Studer, L. & Tomishima, M. J. Neural crest cells from dual SMAD inhibition. Curr. Protoc. Stem Cell Biol. 33, 1H.9.1–1H.9.9 (2015).
Fattahi, F. et al. Deriving human ENS lineages for cell therapy and drug discovery in Hirschsprung disease. Nature 531, 105–109 (2016).
Tchieu, J. et al. A modular platform for differentiation of human PSCs into all major ectodermal lineages. Cell Stem Cell 21, 399–410 (2017).
Hackland, J. O. S. et al. Top-down inhibition of BMP signaling enables robust induction of hPSCs into neural crest in fully defined, xeno-free conditions. Stem Cell Rep. 9, 1043–1052 (2017).
Chalazonitis, A. et al. Neurotrophin-3 induces neural crest-derived cells from fetal rat gut to develop in vitro as neurons or glia. J. Neurosci. 14, 6571–6584 (1994).
Chalazonitis, A., Rothman, T. P., Chen, J. & Gershon, M. D. Age-dependent differences in the effects of GDNF and NT-3 on the development of neurons and glia from neural crest-derived precursors immunoselected from the fetal rat gut: expression of GFRα-1 in vitro and in vivo. Dev. Biol. 204, 385–406 (1998).
Memic, F. et al. Transcription and signaling regulators in developing neuronal subtypes of mouse and human enteric nervous system. Gastroenterology 154, 624–636 (2018).
Lai, F. P.-L. et al. Correction of Hirschsprung-associated mutations in human induced pluripotent stem cells via clustered regularly interspaced short palindromic repeats/Cas9, restores neural crest cell function. Gastroenterology 153, 139–153 (2017).
Burns, A. J. et al. White paper on guidelines concerning enteric nervous system stem cell therapy for enteric neuropathies. Dev. Biol. 417, 229–251 (2016).
Stamp, L. A. Cell therapy for GI motility disorders: comparison of cell sources and proposed steps for treating Hirschsprung disease. Am. J. Physiol. 312, G348–G354 (2017).
Li, W. et al. Characterization and transplantation of enteric neural crest cells from human induced pluripotent stem cells. Mol. Psychiatry 23, 499–508 (2018).
Workman, M. J. et al. Engineered human pluripotent-stem-cell-derived intestinal tissues with a functional enteric nervous system. Nat. Med. 23, 49–59 (2017).
Lee, G. et al. Isolation and directed differentiation of neural crest stem cells derived from human embryonic stem cells. Nat. Biotechnol. 25, 1468–1475 (2007).
Gershon, M. D. & Ratcliffe, E. M. Developmental biology of the enteric nervous system: pathogenesis of Hirschsprung’s disease and other congenital dysmotilities. Semin. Pediatr. Surg. 13, 224–235 (2004).
Chen, G. et al. Chemically defined conditions for human iPSC derivation and culture. Nat. Methods 8, 424–429 (2011).
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat. Biotechnol. 25, 681–686 (2007).
Maldonado, M., Luu, R. J., Ramos, M. E. P. & Nam, J. ROCK inhibitor primes human induced pluripotent stem cells to selectively differentiate towards mesendodermal lineage via epithelial-mesenchymal transition-like modulation. Stem Cell Res. 17, 222–227 (2016).
Sasselli, V., Pachnis, V. & Burns, A. J. The enteric nervous system. Dev. Biol. 366, 64–73 (2012).
Acknowledgements
We thank C. Bispo and A. Carlos (UCSF Flow Cytometry core facility) for excellent technical assistance. The work was supported by the UCSF Program for Breakthrough Biomedical Research and Sandler Foundation, March of Dimes grant no. 1-FY18-394 and 2018 AGA-Rome Foundation Functional GI and MotilityDisorders Pilot Research Award to F.F., the New York State Stem Cell Science (NYSTEM) contract C32599GG and a grant from the National Institutes of Neurological Disorders and Stroke (NINDS) 5R01NS099270 to L.S.
Author information
Authors and Affiliations
Contributions
K.B. contributed to the design, execution and troubleshooting of the experiments, and to writing of the manuscript. L.S. and F.F. contributed to the design and conception of the study, data interpretation, and writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The Memorial Sloan-Kettering Cancer Center has filed a patent application (CA3009509A1) on hPSC-derived ENC lineages for use in Hirschsprung’s disease, with L.S. and F.F. as inventors. L.S. is a co-founder and consultant of BlueRock Therapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference(s) using this protocol
Fattahi, F. et al. Nature 531, 105–109 (2016): https://doi.org/10.1038/nature16951
Tchieu, J. et al. Cell Stem Cell 21, 399–410 (2017): https://doi.org/10.1016/j.stem.2017.08.015
Integrated supplementary information
Supplementary Figure 1 Protocol (days 0–12) for ENC induction using option A.
KSR, knockout serum replacement differentiation medium; LDN, LDN-193189, SB, SB431542, CHIR, CHIR 99021; RA, Retinoic Acid; SB, SB431542.
Supplementary Figure 2 Representative phase contrast image of WA09 embryonic stem cells cultured in E8 medium.
Scale bar = 100 μm.
Supplementary Figure 3 Representative phase contrast images of differentiating cells at different time points of EN induction.
Scale bar = 200 μm.
Supplementary Figure 4 Distinct populations of NOS1+ and CHAT+ cells in hESC–derived EN cultures.
a) Immunofluorescence staining of NOS1 and CHAT on day 75 of EN induction. b) Flow cytometry analysis of NOS1 and CHAT expression on day 75 on EN induction. AF647, Alexa Fluor™ 647; AF488, Alexa Fluor™ 488. Scale bar = 20 μm.
Supplementary Figure 5 Characterization of contaminating cells in hESC–derived EN cultures.
a) Phase contrast image of low density regions of culture plates on day 75 of differentiation. Arrows point to flat non-neuronal contaminating cells. b) Immunofluorescence staining of EN cultures with SMA and TUJ1 on day 75 of differentiation. Scale bar = 100 μm in a and 200 μm in b.
Supplementary Figure 6 Example of FACS gating strategy for purification of CD49D+ ENCs on day 12 of differentiation.
a) Unstained control sample. b) Sample stained with CD49D.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Tables 1 and 2, and Supplementary Methods
Rights and permissions
About this article
Cite this article
Barber, K., Studer, L. & Fattahi, F. Derivation of enteric neuron lineages from human pluripotent stem cells. Nat Protoc 14, 1261–1279 (2019). https://doi.org/10.1038/s41596-019-0141-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0141-y
This article is cited by
-
Safety and efficacy of human ESC-derived corneal endothelial cells for corneal endothelial dysfunction
Cell & Bioscience (2023)
-
Transcriptomics of Hirschsprung disease patient-derived enteric neural crest cells reveals a role for oxidative phosphorylation
Nature Communications (2023)
-
Stem cell-based therapy for hirschsprung disease, do we have the guts to treat?
Gene Therapy (2022)
-
Effects of single bouts of different endurance exercises with different intensities on microRNA biomarkers with and without blood flow restriction: a three-arm, randomized crossover trial
European Journal of Applied Physiology (2021)
-
Innervation: the missing link for biofabricated tissues and organs
npj Regenerative Medicine (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.