Abstract
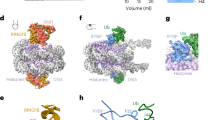
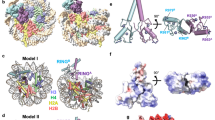
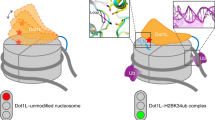
Monoubiquitination of histone H2B-K120/123 plays several roles in regulating transcription, DNA replication and the DNA damage response. The structure of a nucleosome in complex with the dimeric RING E3 ligase Bre1 reveals that one RING domain binds to the nucleosome acidic patch, where it can position the E2 ubiquitin conjugating enzyme Rad6, while the other RING domain contacts the DNA. Comparisons with H2A-specific E3 ligases suggest a general mechanism of tuning histone specificity via the non-E2-binding RING domain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The maps are available in the Electron Microscopy Data Bank (EMDB) database under accession no. EMD-41016 (state 1), EMD-41015 (state 2) and EMD-41011 (state 3). The atomic models are available in the Protein Data Bank database: PDB 8T3Y (state 1), PDB 8T3W(state 2) and PDB 8T3T (state 3). Source data are provided with this paper.
References
Bonnet, J., Devys, D. & Tora, L. Histone H2B ubiquitination: signaling not scrapping. Drug Discov. Today Technol. 12, e19–e27 (2014).
Mattiroli, F. & Penengo, L. Histone ubiquitination: an integrative signaling platform in genome stability. Trends Genet. 37, 566–581 (2021).
Weake, V. M. & Workman, J. L. Histone ubiquitination: triggering gene activity. Mol. Cell 29, 653–663 (2008).
Fleming, A. B., Kao, C. F., Hillyer, C., Pikaart, M. & Osley, M. A. H2B ubiquitylation plays a role in nucleosome dynamics during transcription elongation. Mol. Cell 31, 57–66 (2008).
Pavri, R. et al. Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125, 703–717 (2006).
Hwang, W. W. et al. A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol. Cell 11, 261–266 (2003).
Robzyk, K., Recht, J. & Osley, M. A. Rad6-dependent ubiquitination of histone H2B in yeast. Science 287, 501–504 (2000).
Kim, J., Hake, S. B. & Roeder, R. G. The human homolog of yeast BRE1 functions as a transcriptional coactivator through direct activator interactions. Mol. Cell 20, 759–770 (2005).
Kim, J. et al. RAD6-Mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137, 459–471 (2009).
Sethi, G., Shanmugam, M. K., Arfuso, F. & Kumar, A. P. Role of RNF20 in cancer development and progression—a comprehensive review. Biosci. Rep. 38, BSR20171287 (2018).
Kumar, P. & Wolberger, C. Structure of the yeast Bre1 RING domain. Proteins 83, 1185–1190 (2015).
Turco, E., Gallego, L. D., Schneider, M. & Kohler, A. Monoubiquitination of histone H2B is intrinsic to the Bre1 RING domain-Rad6 interaction and augmented by a second Rad6-binding site on Bre1. J. Biol. Chem. 290, 5298–5310 (2015).
Kim, J. & Roeder, R. G. Direct Bre1–Paf1 complex interactions and RING finger-independent Bre1–Rad6 interactions mediate histone H2B ubiquitylation in yeast. J. Biol. Chem. 284, 20582–20592 (2009).
Gallego, L. D. et al. Structural mechanism for the recognition and ubiquitination of a single nucleosome residue by Rad6–Bre1. Proc. Natl Acad. Sci. USA 113, 10553–10558 (2016).
McGinty, R. K. & Tan, S. Principles of nucleosome recognition by chromatin factors and enzymes. Curr. Opin. Struct. Biol. 71, 16–26 (2021).
Gundogdu, M. & Walden, H. Structural basis of generic versus specific E2-RING E3 interactions in protein ubiquitination. Protein Sci. 28, 1758–1770 (2019).
Pan, M. et al. Structural insights into Ubr1-mediated N-degron polyubiquitination. Nature 600, 334–338 (2021).
Hu, Q. et al. Mechanisms of BRCA1-BARD1 nucleosome recognition and ubiquitylation. Nature 596, 438–443 (2021).
Witus, S. R. et al. BRCA1/BARD1 site-specific ubiquitylation of nucleosomal H2A is directed by BARD1. Nat. Struct. Mol. Biol. 28, 268–277 (2021).
Shukla, P. K. et al. Structure and functional determinants of Rad6-Bre1 subunits in the histone H2B ubiquitin-conjugating complex.Nucleic Acids Res. 51, 2117–2136 (2023).
McGinty, R. K., Henrici, R. C. & Tan, S. Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome. Nature 514, 591–596 (2014).
Worden, E. J., Hoffmann, N. A., Hicks, C. W. & Wolberger, C. Mechanism of cross-talk between H2B ubiquitination and H3 methylation by Dot1L. Cell 176, 1490–1501 e12 (2019).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA.Methods Enzymol 375, 23–44 (2004).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr D. Biol. Crystallogr 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr D. Struct. Biol. 74, 531–544 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr D. Struct. Biol. 75, 861–877 (2019).
Acknowledgements
We thank Johns Hopkins colleagues C. Liu and E. Twomey for advice on Cryo-EM data collection and processing and X. Zhang for valuable suggestions on constructing plasmids. Supported in part by the National Cancer Institute’s National Cryo-EM Facility at the Frederick National Laboratory for Cancer Research under contract 75N91019D00024. This work was supported by National Institute of General Medical Sciences grant GM130393 (C.W.).
Author information
Authors and Affiliations
Contributions
C.W. and F.Z. conceived and designed the study. F.Z. and C.W.H. performed experiments and processed data. F.Z. carried out modeling and map interpretation with assistance from C.W. F.Z., C.W.H. and C.W. wrote the manuscript. C.W. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Uhn-Soo Cho and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Sara Osman and Carolina Perdigoto were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM processing workflow.
The red dashed line indicates the mask generated for use in 3D classification. Three states of Bre1–nucleosome complex are identified and refined respectively.
Extended Data Fig. 2 Cryo-EM validation and example density.
a–c, Local resolution estimation and gold-standard Fourier shell correlation (GSFSC) curves at a FSC cutoff of 0.143 for each state of Bre1–nucleosome complex. d, Representative cryo-EM density maps for the histones, DNA, and Bre1.
Extended Data Fig. 3 Comparison of enzymatic activity of fused and separated Bre1 and Rad6 enzyme.
In vitro nucleosome H2B ubiquitination assay was performed with fused or separated Bre1 and Rad6 enzyme. Samples were quenched by SDS-loading buffer at indicated timepoints and subjected to immunoblotting. Data is representative of 2 independent experiments.
Extended Data Fig. 4 Electrostatic surface representation view of Bre1 dimer (PDB:4R7E).
Bre1 is shown in a cartoon representation with basic residues highlighted as sticks.
Extended Data Fig. 5 Multiple conformations of the Bre1–nucleosome complex.
a–c, Cryo-EM maps of the Bre1–nucleosome complex with an atomic model fitted to states 1, 2 and 3. d, Superimpose of Bre1–nucleosome complex maps comparing states 1, 2 and 3. White arrows show the changes between each conformational state.
Extended Data Fig. 6 RNF20/40 mutation sites mapped to RING domain and preceded α-helix in cancer genomics.
a, Sequence alignment of coiled coil and RING domains of Bre1, RNF20 and RNF40. Black triangle indicates the mutation sites of RNF20 and RNF40 in cancer genomics. b, Complex structure model of Bre1 RING-Rad6-nucleosome. Bre1 residues corresponding to RNF20/40 mutated sites were shown as sticks with dot sphere.
Extended Data Fig. 7 Canonical interface between E3 RING and E2.
a, Superposition of multiple E2-E3 structures. E2-E3 complex structures of RNF4-UbcH5a (PDB: 4AP4), Ubr1-Rad6 (PDB: 7MEX), Ring1b-UbcH5c (PDB: 4R8P), BRCA1-UbcH5c (PDB: 7LYB) and TRIM25-UbcH5 (PDB: 5FER) are aligned together. b, The modeled structure of Bre1 RING-Rad6. c, Residues located in modeled Bre1A RING-Rad6 interface. The side chains of residues located in the interface are shown as sticks.
Extended Data Fig. 8 Superposition of Bre1–nucleosome complex (state 2 and 1) and Ubr1 RING-Rad6-ubiquitin (PDB: 7MEX).
a, c, Ubr1 RING domain is aligned to Bre1-A RING domain in state 2 (a) and state 1 (c). b, d, Close-up view of Rad6 active site C88, ubiquitin G76, and H2B K120 in state 2 (b) and state 1 (d). The H2B K120 rotamer with the shortest distance from Rad6 C88 and ubiquitin G76 is shown here.
Extended Data Fig. 9 Structural alignment of Bre1 RBD-Rad6 complex and Bre1 RING-Rad6-Ub-nucleosome complex (model of state 3).
a, b, Rad6 in Bre1 RBD-Rad6 complex is aligned to the Rad6 in Bre1 RING-Rad6-Ub-nucleosome complex (model of state 3).
Supplementary information
Source data
Source Data Extended Data Fig. 3 and Table 3
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, F., Hicks, C.W. & Wolberger, C. Mechanism of histone H2B monoubiquitination by Bre1. Nat Struct Mol Biol 30, 1623–1627 (2023). https://doi.org/10.1038/s41594-023-01137-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01137-x
This article is cited by
-
Structural mechanisms of autoinhibition and substrate recognition by the ubiquitin ligase HACE1
Nature Structural & Molecular Biology (2024)