Abstract
THEMIS plays an indispensable role in T cells, but its mechanism of action has remained highly controversial. Using the systematic proximity labeling methodology PEPSI, we identify THEMIS as an uncharacterized substrate for the phosphatase SHP1. Saturated mutagenesis assays and mass spectrometry analysis reveal that phosphorylation of THEMIS at the evolutionally conserved Tyr34 residue is oppositely regulated by SHP1 and the kinase LCK. Similar to THEMIS−/− mice, THEMISY34F/Y34F knock-in mice show a significant decrease in CD4 thymocytes and mature CD4 T cells, but display normal thymic development and peripheral homeostasis of CD8 T cells. Mechanistically, the Tyr34 motif in THEMIS, when phosphorylated upon T cell antigen receptor activation, appears to act as an allosteric regulator, binding and stabilizing SHP1 in its active conformation, thus ensuring appropriate negative regulation of T cell antigen receptor signaling. However, cytokine signaling in CD8 T cells fails to elicit THEMIS Tyr34 phosphorylation, indicating both Tyr34 phosphorylation-dependent and phosphorylation-independent roles of THEMIS in controlling T cell maturation and expansion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The mass spectrometry proteomics data, phospho-site mass spectrometry proteomics data and HDX data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository51 with the dataset identifier PXD028689. Source data is provided with this paper.
References
Godfrey, D. I., Kennedy, J., Suda, T. & Zlotnik, A. A developmental pathway involving four phenotypically and functionally distinct subsets of CD3−CD4−CD8− triple-negative adult mouse thymocytes defined by CD44 and CD25 expression. J. Immunol. 150, 4244–4252 (1993).
Zuniga-Pflucker, J. C. T-cell development made simple. Nat. Rev. Immunol. 4, 67–72 (2004).
Germain, R. N. T-cell development and the CD4–CD8 lineage decision. Nat. Rev. Immunol. 2, 309–322 (2002).
Singer, A., Adoro, S. & Park, J. H. Lineage fate and intense debate: myths, models and mechanisms of CD4- versus CD8-lineage choice. Nat. Rev. Immunol. 8, 788–801 (2008).
Liu, Y. et al. Themis is indispensable for IL-2 and IL-15 signaling in T cells. Sci. Signal 15, eabi9983 (2022).
Fu, G. et al. Themis controls thymocyte selection through regulation of T cell antigen receptor-mediated signaling. Nat. Immunol. 10, 848–856 (2009).
Johnson, A. L. et al. Themis is a member of a new metazoan gene family and is required for the completion of thymocyte positive selection. Nat. Immunol. 10, 831–839 (2009).
Lesourne, R. et al. Themis, a T cell-specific protein important for late thymocyte development. Nat. Immunol. 10, 840–847 (2009).
Patrick, M. S. et al. Gasp, a Grb2-associating protein, is critical for positive selection of thymocytes. Proc. Natl Acad. Sci. USA 106, 16345–16350 (2009).
Allen, P. M. Themis imposes new law and order on positive selection. Nat. Immunol. 10, 805–806 (2009).
Kakugawa, K. et al. A novel gene essential for the development of single positive thymocytes. Mol. Cell. Biol. 29, 5128–5135 (2009).
Fu, G. et al. Themis sets the signal threshold for positive and negative selection in T-cell development. Nature 504, 441–445 (2013).
Choi, S., Cornall, R., Lesourne, R. & Love, P. E. THEMIS: two models, different thresholds. Trends Immunol. 38, 622–632 (2017).
Choi, S. et al. THEMIS enhances TCR signaling and enables positive selection by selective inhibition of the phosphatase SHP-1. Nat. Immunol. 18, 433–441 (2017).
Mehta, M. et al. Themis-associated phosphatase activity controls signaling in T cell development. Proc. Natl Acad. Sci. USA 115, E11331–E11340 (2018).
Gascoigne, N. R. & Acuto, O. THEMIS: a critical TCR signal regulator for ligand discrimination. Curr. Opin. Immunol. 33, 86–92 (2015).
Paster, W. et al. A THEMIS:SHP1 complex promotes T-cell survival. EMBO J. 34, 393–409 (2015).
Gascoigne, N. R., Brzostek, J., Mehta, M. & Acuto, O. SHP1-ing thymic selection. Eur. J. Immunol. 46, 2091–2094 (2016).
Paster, W. et al. GRB2-mediated recruitment of THEMIS to LAT is essential for thymocyte development. J. Immunol. 190, 3749–3756 (2013).
Flint, A. J., Tiganis, T., Barford, D. & Tonks, N. K. Development of “substrate-trapping” mutants to identify physiological substrates of protein tyrosine phosphatases. Proc. Natl Acad. Sci. USA 94, 1680–1685 (1997).
Liu, Q. et al. A proximity-tagging system to identify membrane protein–protein interactions. Nat. Methods 15, 715–722 (2018).
Mizuno, K., Tagawa, Y., Watanabe, N., Ogimoto, M. & Yakura, H. SLP-76 is recruited to CD22 and dephosphorylated by SHP-1, thereby regulating B cell receptor-induced c-Jun N-terminal kinase activation. Eur. J. Immunol. 35, 644–654 (2005).
Gascoigne, N. R., Rybakin, V., Acuto, O. & Brzostek, J. TCR signal strength and T cell development. Annu. Rev. Cell Dev. Biol. 32, 327–348 (2016).
Brockmeyer, C. et al. T cell receptor (TCR)-induced tyrosine phosphorylation dynamics identifies THEMIS as a new TCR signalosome component. J. Biol. Chem. 286, 7535–7547 (2011).
Brdicka, T., Kadlecek, T. A., Roose, J. P., Pastuszak, A. W. & Weiss, A. Intramolecular regulatory switch in ZAP-70: analogy with receptor tyrosine kinases. Mol. Cell. Biol. 25, 4924–4933 (2005).
Yang, J. et al. Crystal structure of human protein-tyrosine phosphatase SHP-1. J. Biol. Chem. 278, 6516–6520 (2003).
Wang, W. et al. Crystal structure of human protein tyrosine phosphatase SHP-1 in the open conformation. J. Cell. Biochem. 112, 2062–2071 (2011).
Neel, B. G., Gu, H. & Pao, L. The ‘Shp’ing news: SH2 domain-containing tyrosine phosphatases in cell signaling. Trends Biochem. Sci. 28, 284–293 (2003).
O’Reilly, A. M., Pluskey, S., Shoelson, S. E. & Neel, B. G. Activated mutants of SHP-2 preferentially induce elongation of Xenopus animal caps. Mol. Cell. Biol. 20, 299–311 (2000).
Tabula Muris, C. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Daum, G. et al. A general peptide substrate for protein tyrosine phosphatases. Anal. Biochem. 211, 50–54 (1993).
Matalon, O. et al. Dephosphorylation of the adaptor LAT and phospholipase C-gamma by SHP-1 inhibits natural killer cell cytotoxicity. Sci. Signal. 9, ra54 (2016).
Pumphrey, N. J. et al. Differential association of cytoplasmic signalling molecules SHP-1, SHP-2, SHIP and phospholipase C-gamma1 with PECAM-1/CD31. FEBS Lett. 450, 77–83 (1999).
Brzostek, J. et al. T cell receptor and cytokine signal integration in CD8+ T cells is mediated by the protein Themis. Nat. Immunol. 21, 186–198 (2020).
Fry, T. J. et al. IL-7 therapy dramatically alters peripheral T-cell homeostasis in normal and SIV-infected nonhuman primates. Blood 101, 2294–2299 (2003).
Sprent, J. & Surh, C. D. Normal T cell homeostasis: the conversion of naive cells into memory-phenotype cells. Nat. Immunol. 12, 478–484 (2011).
Tamarit, B. et al. Membrane microdomains and cytoskeleton organization shape and regulate the IL-7 receptor signalosome in human CD4 T-cells. J. Biol. Chem. 288, 8691–8701 (2013).
Goplen, N. P. et al. IL-12 signals through the TCR to support CD8 innate immune responses. J. Immunol. 197, 2434–2443 (2016).
Brugnera, E. et al. Coreceptor reversal in the thymus: signaled CD4+8+ thymocytes initially terminate CD8 transcription even when differentiating into CD8+ T cells. Immunity 13, 59–71 (2000).
Singer, A. New perspectives on a developmental dilemma: the kinetic signaling model and the importance of signal duration for the CD4/CD8 lineage decision. Curr. Opin. Immunol. 14, 207–215 (2002).
Fearnley, G. W. et al. The homophilic receptor PTPRK selectively dephosphorylates multiple junctional regulators to promote cell–cell adhesion. eLife https://doi.org/10.7554/eLife.44597 (2019).
Abram, C. L. & Lowell, C. A. Shp1 function in myeloid cells. J. Leukoc. Biol. 102, 657–675 (2017).
Yu, Q., Erman, B., Bhandoola, A., Sharrow, S. O. & Singer, A. In vitro evidence that cytokine receptor signals are required for differentiation of double positive thymocytes into functionally mature CD8+ T cells. J. Exp. Med. 197, 475–487 (2003).
Sun, C. et al. THEMIS-SHP1 recruitment by 4-1BB tunes LCK-mediated priming of chimeric antigen receptor-redirected T cells. Cancer Cell 37, 216–225 e216 (2020).
E-CRISPR. The German Cancer Research Center http://www.e-crisp.org/E-CRISP/ (2014).
Zhang, Y. C. et al. FER-mediated phosphorylation and PIK3R2 recruitment on IRS4 promotes AKT activation and tumorigenesis in ovarian cancer cells. eLife https://doi.org/10.7554/eLife.76183 (2022).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Song, B. et al. Ordered assembly of the cytosolic RNA-sensing MDA5–MAVS signaling complex via binding to unanchored K63-linked poly-ubiquitin chains. Immunity 54, 2218–2230 e2215 (2021).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Acknowledgements
We thank the staff members of the Animal Facility at the National Facility for Protein Science in Shanghai (NFPS), Zhangjiang Lab, China, for supporting mouse housing and care. We thank the Multi-Omics Core Facility (MOCF) at the School of Life Science and Technology, ShanghaiTech University, for technical support with mass spectrometry experiments. We also thank the Molecular and Cell Biology Core Facility (MCBCF) at the School of Life Science and Technology, ShanghaiTech University, for technical support with flow cytometry. This work was supported by the National Natural Science Foundation of China (32070776 and 31872831 to G.F.), the Ministry of Science and Technology of China (SQ2021YFA080048 and 2018YFC1004603 to G.F.), the Shanghai Science and Technology Commission (19JC1413800 to G.F.), the Shanghai Shuguang program (19SG55 to G.F.), the Shanghai Sailing Program (21YF1429900 to Y.W.), the Key Research Project of Zhejiang Lab (2021PE0AC06 to G.F.) and a ShanghaiTech University startup grant. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the paper.
Author information
Authors and Affiliations
Contributions
G.F. and H.W. designed the study. J.L.Z., Z.J., X.Z., Z.Y., J.W., J.C., L.C., M.S., Y.Z., M.H., S.C., X.X. and Y.W. performed the experiments; J.L.Z. was the major contributor; E.Z. generated THEMISY34F/Y34F and THEMIS−/− mice; P.H., T.H., M.Z., L.Z., F.B., J.Z., H.W. and G.F. analyzed and interpreted the data; J.L.Z. and G.F. wrote the paper with help from H.W. G.F. oversaw the whole project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Nicholas Gascoigne and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Dimitris Typas was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PEPSI strategy identifies THEMIS as a substrate of non-receptor tyrosine phosphatase SHP1.
a, Inducible expressing pupE Jurkat T cells were stably transfected with SHP1-WT-PafA-Myc or SHP1-DA-PafA-Myc, indicating that PafA could successfully label SHP1 and SHP1 interactome with pupE which recruits biotin stained by streptavidin antibody. b-c, HEK293T cells were transiently transfected with THEMIS-WT-FLAG and SHP1-WTPafA-Myc or SHP1-DA-PafA-Myc (b) and SHP1-WT or SHP1-DA (c), indicating that THEMIS bound more with SHP1-DA than SHP1-WT. d, Inducible expressing pupE Jurkat T cells were transiently transfected with PafA or SHP1-PafA and showed THEMIS bound to SHP1 but not PafA. e, HEK293T cells were transiently transfected with THEMIS-WT-FLAG and SHP1-WT-PafA-Myc, SHP1-DA-PafA-Myc, PTP1B-WT-PafA-Myc, or PTP1B-DA-PafA-Myc. Results from the co-immunoprecipitation assay indicated that PTP1B, either WT or DA mutant, couldn’t bind to THEMIS.
Extended Data Fig. 2 Phosphorylation regulation of THEMIS at Tyr34 site.
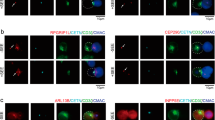
a, Sequence homology comparison of THEMIS Tyr34 site in 9 species. b, THEMIS amino acid sequence showed a 31aa-long peptide after trypsin digestion in the red box. c, As indicated, HEK293T cells were transiently transfected with THEMIS-Myc and GRB2-FLAG or LCK-FLAG. Co-immunoprecipitation assay demonstrated that the interaction between LCK and THEMIS was transient and weak compared to GRB2. d-e, Immunoblotting of truncation variant experiments in the HEK293T cell line exploring the three domains of THEMIS with five different truncation mutants (d), Western blotting results showing that LCK can only phosphorylate the THEMIS Tyr34 site, positioned in the CABIT1 domain (e). f, Sequence alignment between hGARE1, mGARE1, hGARE2, mGARE2, mTHEMIS3, hTHEMIS1, mTHEMIS1, hTHEMIS2, and mTHEMIS2, showing one conserved Tyr residue (Y34 in THEMIS and Y35 in THEMIS2, respectively).
Extended Data Fig. 3 SHP1 dephosphorylates THEMIS at Tyr34 site.
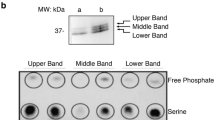
a, In vitro DiFMUP assay, which detects the activity of protein phosphatase SHP1 showed that the constitutively activated SHP1-EA (E74A) mutant has the highest activity; the lowest activity was detected for the phosphatase-dead SHP1-CS (C453S) mutant (n = 5 biologically independent samples). b, Dot blot results indicate that the constitutively activated SHP1-EA mutant can dephosphorylate an artificial THEMIS p-Y34 peptide; the peptide sequence is shown above. c, In vitro dephosphorylation assays which mixed the purified SHP1 protein from E. coli and the THEMIS protein affinity purified from HEK293T cells at 30 °C for 1 hour. Western blotting analysis indicated that SHP1 catalyzed the dephosphorylation of THEMIS Tyr34. d, Src kinase family inhibitor PP2 treatment contributed to the blockade of phosphorylation transduction, including THEMIS Tyr34 site after TCR signaling activation by C305 (anti-TCR) antibody.
Extended Data Fig. 4 THEMIS proteins in THEMIS+/+ and THEMISY34F/Y34F mice are expressed at a comparable level.
a, Distribution of t-SNE of single-cell RNA-seq data from Tabula Muris, colored by different tissue (left) and expression of THEMIS among different tissues (right). b, THEMIS expression in top 10 cell types ordered by the mean of THEMIS log transform expression in individual cell type, data from Tabula Muris. In each box plot, the center line indicates the median, the edges of the box represent the first and third quartiles, and the whiskers extend to span a 1.5 interquartile range from the edges. c, Sanger sequencing of THEMIS-/-, THEMISY34F/Y34F, and THEMISY34F/+ mice tail genotype identification. d, Gray-level analysis of Western bands in Fig. 4A showed the same expression of THEMIS in THEMIS+/+ and THEMISY34F/Y34F mice (n = 3 mice examined over 5 independent blots). e-f, Quantitative fluorescence immunoblotting analysis (e) showed both WT and Y34F THEMIS proteins in total thymocytes were expressed at a comparable level. The analysis data was sourced from 6 mice examined over 3 independent blots of WT and Y34F genotypes (f). g, Flow cytometry intracellular staining of THEMIS protein showed both WT and Y34F THEMIS proteins in DP thymocytes were expressed at a comparable level (n = 5 mice).
Extended Data Fig. 5 THEMISY34F/Y34F mice show a significant developmental defect in CD4 thymocyte development.
a-e, Mouse T cells (6 ~ 7-week old) were stained with anti-CD4−Qdot605, anti-CD8−APCCy7, anti-TCRb-PerCP-Cy5.5, and anti-CD69-PE-Cy7 in thymocytes or CD4-PerCP-Cy5.5 and CD8-FITC in splenocytes for 1 hour on ice. Quantification of the percentage of CD4SP (a), CD8SP (b), and mature cells (c) in total thymocytes or CD4+ (d) and CD8+ (e) T cells among total splenocytes isolated from male mice (left) and female mice (right) of the three genotypes (male: n = 16, 13, 10; female: n = 8, 6, 12).
Extended Data Fig. 6 The structure of THEMIS predicted by AlphaFold, with Tyr34 highlighted.
a, Recombinant SHP1WT and SHP1EA proteins were incubated with the Tyr34-nonphosphorylated THEMIS peptide, the Y34D THEMIS peptide or the Tyr132-phosphorylated LAT peptide, and then the pTyr peptide was added as substrate, followed by malachite green phosphate assay. Released phosphate from the pTyr peptide was calculated as shown and plotted (n = 3 biologically independent samples). b, The CABIT1, CABIT2, PRS domains, and Tyr34 residue were colored in orange, cyan, pink, and red, respectively (https://www.alphafold.ebi.ac.uk/entry/Q8N1K5). Visualization of the atomic model was processed by UCSF ChimeraX (version 1.2.5).
Extended Data Fig. 7 Fitting curve of SPR assay and comparative HDX profiles.
a, Sensorgrams of binding of the SHP1 N + C protein and the SHP1 PTP protein to the Y34D THEMIS peptide, the Tyr34-phosphorylated THEMIS peptide, and the Tyr686 PECAM peptide. b, Differential HDX view of SHP1 upon binding to the Tyr686-phosphorylated PECAM peptide, the Tyr34-nonphosphorylated THEMIS peptide, and the Tyr34-phosphorylated THEMIS peptide. The values listed under each HDX experiment demonstrated the averaged difference in the percentage of deuterium incorporation of that corresponding peptide derived from two different states across all exchange time points (that is, 0 s, 10 s, 60 s, 300 s, and 900 s). HDX Workbench colored each peptide according to the smooth color gradient HDX perturbation key (D%) shown below. Differences in %D between -5% to +5% were considered nonsignificant and are colored gray.
Extended Data Fig. 8 Gating strategy for flow cytometry analysis.
a, Gating strategy for T cell development analysis in total thymocytes. Cells were first gated by FSC/SSC to exclude debris, followed by gating FCS-A/FCS-W and SSC-A/SSC-W to eliminate non-single cells. Live cells were gated by Indo-1(Violet) negative, followed by total thymocyte development analysis. b, Gating strategy for T cell development analysis in total splenocytes. Cells were first gated by FSC/SSC to exclude debris, followed by gating FCS-A/FCS-W and SSC-A/SSC-W to eliminate non-single cells. Live cells were gated by Pacific Blue negative, followed by total splenocytes development analysis. c, Gating strategy for THEMIS protein expression in DP thymocytes. Cells were first gated by FSC/SSC to exclude debris, followed by gating SSC-A/SSC-W and FCS-A/FCS-W to eliminate non-single cells. Different mice cells were distributed by distinct FITC/BV421 staining, and then DP cells were gated by CD4-BV605 and CD8-APC-Cy7 double positive population. THEMIS protein expression was stained by APC.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Fig. 6
SPR source data.
Source Data Extended Data Fig. 7
SPR source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, J., Jiang, Z., Zhang, X. et al. THEMIS is a substrate and allosteric activator of SHP1, playing dual roles during T cell development. Nat Struct Mol Biol 31, 54–67 (2024). https://doi.org/10.1038/s41594-023-01131-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01131-3