Abstract
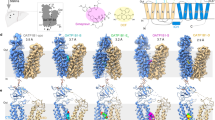
Organic anion transporters (OATs) of the SLC22 family have crucial roles in the transport of organic anions, including metabolites and therapeutic drugs, and in transporter-mediated drug-drug interactions. In the kidneys, OATs facilitate the elimination of metabolic waste products and xenobiotics. However, their transport activities can lead to the accumulation of certain toxic compounds within cells, causing kidney damage. Moreover, OATs are important drug targets, because their inhibition modulates the elimination or retention of substrates linked to diseases. Despite extensive research on OATs, the molecular basis of their substrate and inhibitor binding remains poorly understood. Here we report the cryo-EM structures of rat OAT1 (also known as SLC22A6) and its complexes with para-aminohippuric acid and probenecid at 2.1, 2.8 and 2.9 Å resolution, respectively. Our findings reveal a highly conserved substrate binding mechanism for SLC22 transporters, wherein four aromatic residues form a cage to accommodate the polyspecific binding of diverse compounds.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cryo-EM density maps of apo-rOAT1, the rOAT1–PAH complex and the rOAT1–probenecid complex have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-40352 (refined by CryoSPARC), EMD-40354 and EMD-40355, respectively. The map of apo-rOAT1 refined by RELION4 is available with accession code EMD-40948. The atomic coordinates of the structures of apo-rOAT1, the rOAT1–PAH complex and the rOAT1–probenecid complex have been deposited in the Protein Data Bank (PDB) under accession codes 8SDU, 8SDY and 8SDZ, respectively. The modified version of RELION4 used in this work is available at GitHub https://github.com/jiangjiansen/relion_composite_masks. Source data are provided with this paper.
References
Lopez-Nieto, C. E. et al. Molecular cloning and characterization of NKT, a gene product related to the organic cation transporter family that is almost exclusively expressed in the kidney. J. Biol. Chem. 272, 6471–6478 (1997).
Simonson, G. D., Vincent, A. C., Roberg, K. J., Huang, Y. & Iwanij, V. Molecular cloning and characterization of a novel liver-specific transport protein. J. Cell Sci. 107, 1065–1072 (1994).
Mori, K. et al. Kidney-specific expression of a novel mouse organic cation transporter-like protein. FEBS Lett. 417, 371–374 (1997).
Brady, K. P. et al. A novel putative transporter maps to the osteosclerosis (oc) mutation and is not expressed in the oc mutant mouse. Genomics 56, 254–261 (1999).
Nishimura, M. & Naito, S. Tissue-specific mRNA expression profiles of human ATP-binding cassette and solute carrier transporter superfamilies. Drug Metab. Pharmacokinet. 20, 452–477 (2005).
Hilgendorf, C. et al. Expression of thirty-six drug transporter genes in human intestine, liver, kidney, and organotypic cell lines. Drug Metab. Dispos. 35, 1333–1340 (2007).
Rizwan, A. N. & Burckhardt, G. Organic anion transporters of the SLC22 family: biopharmaceutical, physiological, and pathological roles. Pharm. Res. 24, 450–470 (2007).
Nigam, S. K. et al. The organic anion transporter (OAT) family: a systems biology perspective. Physiol. Rev. 95, 83–123 (2015).
Nigam, S. K. The SLC22 transporter family: a paradigm for the impact of drug transporters on metabolic pathways, signaling, and disease. Annu. Rev. Pharmacol. Toxicol. 58, 663–687 (2018).
Hagos, Y. & Wolff, N. A. Assessment of the role of renal organic anion transporters in drug-induced nephrotoxicity. Toxins (Basel) 2, 2055–2082 (2010).
Sakiyama, M. et al. A common variant of organic anion transporter 4 (OAT4/SLC22A11) gene is associated with renal underexcretion type gout. Drug Metab. Pharmacokinet. 29, 208–210 (2014).
Yee, S. W. & Giacomini, K. M. Emerging roles of the human solute carrier 22 family. Drug Metab. Dispos. 50, 1193–1210 (2021).
Chow, P. N. An improved toluene-triton-based liquid scintillation system for counting 14C-labeled compounds at ambient temperature. Anal. Biochem. 60, 322–328 (1974).
Masuda, S., Saito, H. & Inui, K. I. Interactions of nonsteroidal anti-inflammatory drugs with rat renal organic anion transporter, OAT-K1. J. Pharmacol. Exp. Ther. 283, 1039–1042 (1997).
Cha, S. H. et al. Molecular cloning and characterization of multispecific organic anion transporter 4 expressed in the placenta. J. Biol. Chem. 275, 4507–4512 (2000).
Kimura, H. et al. Human organic anion transporters and human organic cation transporters mediate renal transport of prostaglandins. J. Pharmacol. Exp. Ther. 301, 293–298 (2002).
Takeda, M. et al. Characterization of methotrexate transport and its drug interactions with human organic anion transporters. J. Pharmacol. Exp. Ther. 302, 666–671 (2002).
Giacomini, K. M., Galetin, A. & Huang, S. M. The International Transporter Consortium: summarizing advances in the role of transporters in drug development. Clin. Pharmacol. Ther. 104, 766–771 (2018).
Hillgren, K. M. et al. Emerging transporters of clinical importance: an update from the International Transporter Consortium. Clin. Pharmacol. Ther. 94, 52–63 (2013).
FDA. In Vitro Drug Interaction Studies — Cytochrome P450 Enzyme and Transporter-Mediated Drug Interactions Guidance for Industry (2020).
FDA. Clinical Drug Interaction Studies — Cytochrome P450 Enzyme and Transporter-mediated Drug Interactions Guidance for Industry (2020).
EMA. Guideline on the Investigation of Drug Interactions (2012).
Hosoyamada, M., Sekine, T., Kanai, Y. & Endou, H. Molecular cloning and functional expression of a multispecific organic anion transporter from human kidney. Am. J. Physiol. 276, F122–F128 (1999).
Jariyawat, S. et al. The interaction and transport of beta-lactam antibiotics with the cloned rat renal organic anion transporter 1. J. Pharmacol. Exp. Ther. 290, 672–677 (1999).
Maeda, K. et al. Inhibitory effects of p-aminohippurate and probenecid on the renal clearance of adefovir and benzylpenicillin as probe drugs for organic anion transporter (OAT) 1 and OAT3 in humans. Eur. J. Pharm. Sci. 59, 94–103 (2014).
Burckhardt, B. C. et al. Transport of cimetidine by flounder and human renal organic anion transporter 1. Am. J. Physiol. Renal Physiol. 284, F503–F509 (2003).
Yamada, A. et al. Multiple human isoforms of drug transporters contribute to the hepatic and renal transport of olmesartan, a selective antagonist of the angiotensin II AT1-receptor. Drug Metab. Dispos. 35, 2166–2176 (2007).
Zhang, J., Wang, H., Fan, Y., Yu, Z. & You, G. Regulation of organic anion transporters: role in physiology, pathophysiology, and drug elimination. Pharmacol. Ther. 217, 107647 (2021).
Eraly, S. A. et al. Decreased renal organic anion secretion and plasma accumulation of endogenous organic anions in OAT1 knock-out mice. J. Biol. Chem. 281, 5072–5083 (2006).
Fu, Y. et al. Organic anion transporter OAT3 enhances the glucosuric effect of the SGLT2 inhibitor empagliflozin. Am. J. Physiol. Renal Physiol. 315, F386–F394 (2018).
Zou, L. et al. Molecular mechanisms for species differences in organic anion transporter 1, OAT1: implications for renal drug toxicity. Mol. Pharmacol. 94, 689–699 (2018).
Ho, E. S., Lin, D. C., Mendel, D. B. & Cihlar, T. Cytotoxicity of antiviral nucleotides adefovir and cidofovir is induced by the expression of human renal organic anion transporter 1. J. Am. Soc. Nephrol. 11, 383–393 (2000).
Mason, R. M. Studies on the effect of probenecid (benemid) in gout. Ann. Rheum. Dis. 13, 120–130 (1954).
Cunningham, R. F., Israili, Z. H. & Dayton, P. G. Clinical pharmacokinetics of probenecid. Clin. Pharmacokinet. 6, 135–151 (1981).
Granados, J. C., Bhatnagar, V. & Nigam, S. K. Blockade of organic anion transport in humans after treatment with the drug probenecid leads to major metabolic alterations in plasma and urine. Clin. Pharmacol. Ther. 112, 653–664 (2022).
Cundy, K. C. Clinical pharmacokinetics of the antiviral nucleotide analogues cidofovir and adefovir. Clin. Pharmacokinet. 36, 127–143 (1999).
Beyer, K. H., Flippin, H., Verwey, W. F. & Woodward, R. The effect of para-aminohippuric acid on plasma concentration of penicillin in man. JAMA 126, 1007–1009 (1944).
Hong, M. et al. Human organic anion transporter hOAT1 forms homooligomers. J. Biol. Chem. 280, 32285–32290 (2005).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing.eLife 7, e35383 (2018).
Wang, N. et al. Structural basis of human monocarboxylate transporter 1 inhibition by anti-cancer drug candidates. Cell 184, 370–383 e313 (2021).
Drew, D., North, R. A., Nagarathinam, K. & Tanabe, M. Structures and general transport mechanisms by the major facilitator superfamily (MFS).Chem. Rev. 121, 5289–5335 (2021).
Khanppnavar, B. et al. Structural basis of organic cation transporter-3 inhibition. Nat. Commun. 13, 6714 (2022).
Brast, S. et al. The cysteines of the extracellular loop are crucial for trafficking of human organic cation transporter 2 to the plasma membrane and are involved in oligomerization. FASEB J. 26, 976–986 (2012).
Keller, T. et al. The large extracellular loop of organic cation transporter 1 influences substrate affinity and is pivotal for oligomerization. J. Biol. Chem. 286, 37874–37886 (2011).
Miyano, M., Ago, H., Saino, H., Hori, T. & Ida, K. Internally bridging water molecule in transmembrane alpha-helical kink. Curr. Opin. Struct. Biol. 20, 456–463 (2010).
Rizwan, A. N., Krick, W. & Burckhardt, G. The chloride dependence of the human organic anion transporter 1 (hOAT1) is blunted by mutation of a single amino acid. J. Biol. Chem. 282, 13402–13409 (2007).
Perry, J. L., Dembla-Rajpal, N., Hall, L. A. & Pritchard, J. B. A three-dimensional model of human organic anion transporter 1: aromatic amino acids required for substrate transport. J. Biol. Chem. 281, 38071–38079 (2006).
Beyer, K. H. Factors basic to the development of useful inhibitors of renal transport mechanisms. Arch. Int. Pharmacodyn. Ther. 98, 97–117 (1954).
Cihlar, T. & Ho, E. S. Fluorescence-based assay for the interaction of small molecules with the human renal organic anion transporter 1. Anal. Biochem. 283, 49–55 (2000).
Zhao, Y. et al. Conformational preferences of π–π stacking between ligand and protein, analysis derived from crystal structure data geometric preference of π–π interaction. Interdiscip. Sci. 7, 211–220 (2015).
Fraser-Spears, R. et al. Comparative analysis of novel decynium-22 analogs to inhibit transport by the low-affinity, high-capacity monoamine transporters, organic cation transporters 2 and 3, and plasma membrane monoamine transporter. Eur. J. Pharmacol. 842, 351–364 (2019).
Quistgaard, E. M., Low, C., Guettou, F. & Nordlund, P. Understanding transport by the major facilitator superfamily (MFS): structures pave the way. Nat. Rev. Mol. Cell Biol. 17, 123–132 (2016).
Hong, M., Zhou, F., Lee, K. & You, G. The putative transmembrane segment 7 of human organic anion transporter hOAT1 dictates transporter substrate binding and stability. J. Pharmacol. Exp. Ther. 320, 1209–1215 (2007).
Liu, H. C. et al. Molecular properties of drugs interacting with SLC22 transporters OAT1, OAT3, OCT1, and OCT2: a machine-learning approach. J. Pharmacol. Exp. Ther. 359, 215–229 (2016).
Fujita, T. et al. Functional analysis of polymorphisms in the organic anion transporter, SLC22A6 (OAT1). Pharmacogenet. Genomics 15, 201–209 (2005).
Bleasby, K., Hall, L. A., Perry, J. L., Mohrenweiser, H. W. & Pritchard, J. B. Functional consequences of single nucleotidepolymorphisms in the human organic anion transporter hOAT1 (SLC22A6). J. Pharmacol. Exp. Ther. 314, 923–931 (2005).
Zazuli, Z. et al. The impact of genetic polymorphisms in organic cation transporters on renal drug disposition.Int. J. Mol. Sci. 21, 6627 (2020).
Tzvetkov, M. V. et al. The effects of genetic polymorphisms in the organic cation transporters OCT1, OCT2, and OCT3 on the renal clearance of metformin. Clin. Pharmacol. Ther. 86, 299–306 (2009).
Engstrom, K. et al. Polymorphisms in genes encoding potential mercury transporters and urine mercury concentrations in populations exposed to mercury vapor from gold mining. Environ. Health Perspect. 121, 85–91 (2013).
Wolff, N. A. et al. Cationic amino acids involved in dicarboxylate binding of the flounder renal organic anion transporter. J. Am. Soc. Nephrol. 12, 2012–2018 (2001).
Feng, B., Dresser, M. J., Shu, Y., Johns, S. J. & Giacomini, K. M. Arginine 454 and lysine 370 are essential for the anion specificity of the organic anion transporter, rOAT3. Biochemistry 40, 5511–5520 (2001).
Minuesa, G. et al. Transport of lamivudine[(-)-beta-l-2′,3′-dideoxy-3′-thiacytidine] and high-affinity interaction ofnucleoside reverse transcriptase inhibitors with human organic cationtransporters 1, 2, and 3.J. Pharmacol. Exp. Ther. 329, 252–261 (2009).
Mulato, A. S., Ho, E. S. & Cihlar, T. Nonsteroidal anti-inflammatory drugs efficiently reduce the transport and cytotoxicity of adefovir mediated by the human renal organic anion transporter 1. J. Pharmacol. Exp. Ther. 295, 10–15 (2000).
Taniguchi, T. et al. Hypouricemic agents reduce indoxyl sulfate excretion by inhibiting the renal transporters OAT1/3 and ABCG2. Sci. Rep. 11, 7232 (2021).
Kaufhold, M. et al. Differential interaction of dicarboxylates with human sodium-dicarboxylate cotransporter 3 and organic anion transporters 1 and 3. Am. J. Physiol. Renal Physiol. 301, F1026–F1034 (2011).
Tsigelny, I. F. et al. Conformational changes of the multispecific transporter organic anion transporter 1 (OAT1/SLC22A6) suggests a molecular mechanism for initial stages of drug and metabolite transport. Cell Biochem. Biophys. 61, 251–259 (2011).
Janaszkiewicz, A. et al. Insights into the structure and function of the human organic anion transporter 1 in lipid bilayer membranes. Sci. Rep. 12, 7057 (2022).
Shin, H. J. et al. Interactions of urate transporter URAT1 in human kidney with uricosuric drugs. Nephrology (Carlton) 16, 156–162 (2011).
Enomoto, A. et al. Interaction of human organic anion transporters 2 and 4 with organic anion transport inhibitors. J. Pharmacol. Exp. Ther. 301, 797–802 (2002).
Dalbeth, N. et al. Lesinurad, a selective uric acid reabsorption inhibitor, in combination with febuxostat in patients with tophaceous gout: findings of a phase III clinical trial. Arthritis Rheumatol. 69, 1903–1913 (2017).
Taniguchi, T. et al. Pharmacological evaluation of dotinurad, a selective urate reabsorption inhibitor. J. Pharmacol. Exp. Ther. 371, 162–170 (2019).
Pao, S. S., Paulsen, I. T. & Saier, M. H. Jr. Major facilitator superfamily. Microbiol. Mol. Biol. Rev. 62, 1–34 (1998).
Yan, N. Structural biology of the major facilitator superfamily transporters. Annu. Rev. Biophys. 44, 257–283 (2015).
Li, F. et al. Ion transport and regulation in a synaptic vesicle glutamate transporter. Science 368, 893–897 (2020).
Parker, J. L. et al. Cryo-EM structure of PepT2 reveals structural basis for proton-coupled peptide and prodrug transport in mammals.Sci. Adv. 7, eabh3355 (2021).
Killer, M., Wald, J., Pieprzyk, J., Marlovits, T. C. & Low, C. Structural snapshots of human PepT1 and PepT2 reveal mechanistic insights into substrate and drug transport across epithelial membranes. Sci. Adv. 7, eabk3259 (2021).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Acknowledgements
This work was supported by the Intramural Research Program at the National Institutes of Health (NIH), National Heart, Lung, and Blood Institute (NHLBI). This work utilized the NIH Multi-Institute Cryo-EM Facility (MICEF), the computational resources of the NIH High Performing Computation (HPC) Biowulf cluster (http://hpc.nih.gov) and the instruments maintained by the NHLBI Biochemistry Core. We thank H. Wang, Y. Cui and H. He for technical support on the electron microscopes and R. Saracuza for technical support on installation and maintenance of the in-house GPU computers.
Author information
Authors and Affiliations
Contributions
J.J. conceived the project. T.D., T.L., S.S. and Y.H. conducted cell culture and protein expression. T.D. and T.L. performed protein purification and cryo-EM sample preparation. T.D., T.L. and J.J. performed cryo-EM data collection and processing. T.D. and J.J. built the atomic models. J.J. and T.D. wrote the manuscript with input from all of the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Protein purification of rOAT1.
a, Size exclusion chromatography (SEC) elution profile of rOAT1. b, SDS-PAGE image of purified rOAT1. The purification of rOAT1 was repeated more than three times with similar results.
Extended Data Fig. 2 Cryo-EM data of rOAT1.
a, A representative motion-corrected cryo-EM micrograph of rOAT1. A total of over 20,000 micrographs with a quality similar to this one were collected for this work. b, Representative 2D class averages of rOAT1 particles. The side length of each image box is 212 Å.
Extended Data Fig. 3 Cryo-EM data processing workflow for apo-rOAT1.
A set of composite masks used in 3D classification and 3D auto-refinement is shown at the top.
Extended Data Fig. 4 Cryo-EM data processing workflow for the rOAT1-PAH complex.
The 3D reconstruction of apo-rOAT1 was used as the initial model. Some top-view particles were randomly removed before the final 3D auto-refinement to improve the particle orientation distribution.
Extended Data Fig. 5 Cryo-EM data processing workflow for the rOAT1-probenecid complex.
The 3D reconstruction of apo-rOAT1 was used as the initial model.
Extended Data Fig. 6 Resolution estimation of cryo-EM 3D reconstructions of apo-rOAT1, the rOAT1-PAH complex, and the rOAT1-probenecid complex.
a-c, FSC curves (left), particle orientation plots (middle), and local resolution maps (right) of the 3D reconstructions of apo-rOAT1 (a), the rOAT1-PATH complex (b), and the rOAT1-probenecid complex (c). d, FSC curves of non-uniform 3D refinement of apo-rOAT1 using cryoSPARC.
Extended Data Fig. 7 High-resolution details in the cryo-EM density map of apo-rOAT1.
a-c, The cryo-EM densities (grey surfaces) of the transmembrane helices (a), the ECD (b), and the ICD (c) superimposed with the atomic model. d,e, Close-up views of the disulfide bonds in the ECD, showing the cryo-EM density map (grey surfaces) superimposed with the atomic model.
Extended Data Fig. 8 Comparison of the cryo-EM density maps of apo-rOAT1, the rOAT1-PAH complex, and the rOAT1-probenecid complex near the region of the substrate binding site.
a, apo-rOAT1. b, The rOAT1-PAH complex. c, The rOAT1-probenecid complex. The cryo-EM density maps are shown as grey surfaces. The residues of rOAT1 are colored in medium purple (NTD) or cyan (CTD). Water molecules are depicted by red spheres.
Extended Data Fig. 9 Cryo-EM density map of the PAH binding site in the rOAT1-PAH complex.
Three possible models of PAH (white stick models) are superimposed with the cryo-EM density map (grey surfaces) separately. The residues of rOAT1 are colored in medium purple (NTD) or cyan (CTD).
Extended Data Fig. 10 Cryo-EM density map of the probenecid binding site in the rOAT1-probenecid complex.
a, The cryo-EM density map (grey surfaces) of the rOAT1-probenecid complex superimposed with the atomic model. Probenecid is shown as a grey stick model. b, The same view of the cryo-EM density map of apo-rOAT1.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Video 1
Overall structure of apo-rOAT1. The sharpened cryo-EM map of apo-rOAT1 superimposed with the atomic model is first shown, followed by the atomic model with the transmembrane helices rainbow colored.
Supplementary Video 2
Water molecules in the structure of apo-rOAT1. The cryo-EM densities of bound/ordered water molecules are shown as gray surfaces. The hydrogen bonds involving water molecules are depicted by orange dashed lines.
Supplementary Video 3
The binding of PAH in rOAT1. The cryo-EM density map of the rOAT1–PAH complex (gray surface) is superimposed with the atomic model. Three possible poses of PAH (white stick model) in the binding site are shown sequentially.
Supplementary Video 4
The binding of probenecid in rOAT1. The cryo-EM density map of the rOAT1–probenecid complex (gray surface) is superimposed with the atomic model. Probenecid is shown as a gray stick model in the center.
Supplementary Video 5
A putative conformational conversion between the inward-facing and outward-facing states of rOAT1. The animation shows the morph between the inward-facing structure and the outward-facing model of rOAT1. The NTD and CTD rotate as rigid bodies without considering conformational changes within each of the two domains. The side chains of the charged residues (Asp, Glu, Lys, and Arg) are shown as ball-and-stick models.
Source data
Source Data Extended Data Fig. 1b
Unprocessed scan of gel.
Rights and permissions
About this article
Cite this article
Dou, T., Lian, T., Shu, S. et al. The substrate and inhibitor binding mechanism of polyspecific transporter OAT1 revealed by high-resolution cryo-EM. Nat Struct Mol Biol 30, 1794–1805 (2023). https://doi.org/10.1038/s41594-023-01123-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01123-3
This article is cited by
-
Structural insights into human organic cation transporter 1 transport and inhibition
Cell Discovery (2024)
-
OAT1 structures reveal insights into drug transport in the kidney
Nature Structural & Molecular Biology (2023)