Abstract
In humans, brain oscillations support critical features of memory formation. However, understanding the molecular mechanisms underlying this activity remains a major challenge. Here, we measured memory-sensitive oscillations using intracranial electroencephalography recordings from the temporal cortex of patients performing an episodic memory task. When these patients subsequently underwent resection, we employed transcriptomics on the temporal cortex to link gene expression with brain oscillations and identified genes correlated with oscillatory signatures of memory formation across six frequency bands. A co-expression analysis isolated oscillatory signature-specific modules associated with neuropsychiatric disorders and ion channel activity, with highly correlated genes exhibiting strong connectivity within these modules. Using single-nucleus transcriptomics, we further revealed that these modules are enriched for specific classes of both excitatory and inhibitory neurons, and immunohistochemistry confirmed expression of highly correlated genes. This unprecedented dataset of patient-specific brain oscillations coupled to genomics unlocks new insights into the genetic mechanisms that support memory encoding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The RNA-seq dataset used for memory oscillatory signature analysis in this study are available at GEO with accession number GSE139914.
Code availability
Custom R codes for the quality control, MVA, correlative analysis, permutation/bootstraps, WGCNA, snRNA-seq analysis, snATAC-seq analysis, visualizations, functional enrichments, GWAS enrichment and gene set enrichments are available at https://github.com/konopkalab/Within_Subject.
References
Morgan, S. E. et al. Cortical patterning of abnormal morphometric similarity in psychosis is associated with brain expression of schizophrenia-related genes. Proc. Natl Acad. Sci. USA 116, 9604–9609 (2019).
Lombardo, M. V. et al. Large-scale associations between the leukocyte transcriptome and BOLD responses to speech differ in autism early language outcome subtypes. Nat. Neurosci. 21, 1680–1688 (2018).
Romero-Garcia, R., Warrier, V., Bullmore, E. T., Baron-Cohen, S. & Bethlehem, R. A. I. Synaptic and transcriptionally downregulated genes are associated with cortical thickness differences in autism. Mol. Psychiatry 24, 1053–1064 (2019).
Wang, G. Z. et al. Correspondence between resting-state activity and brain gene expression. Neuron 88, 659–666 (2015).
Patania, A. et al. Topological gene expression networks recapitulate brain anatomy and function. Netw. Neurosci. 3, 744–762 (2019).
Le, B. D. & Stein, J. L. Mapping causal pathways from genetics to neuropsychiatric disorders using genome-wide imaging genetics: current status and future directions. Psychiatry Clin. Neurosci. 73, 357–369 (2019).
Konopka, G. Cognitive genomics: linking genes to behavior in the human brain. Netw. Neurosci. 1, 3–13 (2017).
Berto, S., Wang, G. Z., Germi, J., Lega, B. C. & Konopka, G. Human genomic signatures of brain oscillations during memory encoding. Cereb. Cortex 28, 1733–1748 (2018).
Long, N. M., Burke, J. F. & Kahana, M. J. Subsequent memory effect in intracranial and scalp EEG. NeuroImage 84, 488–494 (2014).
Mukamel, R. & Fried, I. Human intracranial recordings and cognitive neuroscience. Annu. Rev. Psychol. 63, 511–537 (2012).
Sederberg, P. B. et al. Hippocampal and neocortical gamma oscillations predict memory formation in humans. Cereb. Cortex 17, 1190–1196 (2007).
Nakamura, K. & Kubota, K. The primate temporal pole: its putative role in object recognition and memory. Behav. Brain Res. 77, 53–77 (1996).
Hill, P. F., King, D. R., Lega, B. C. & Rugg, M. D. Comparison of fMRI correlates of successful episodic memory encoding in temporal lobe epilepsy patients and healthy controls. NeuroImage 207, 116397 (2020).
Sederberg, P. B., Howard, M. W. & Kahana, M. J. A context-based theory of recency and contiguity in free recall. Psychol. Rev. 115, 893–912 (2008).
Arora, A. et al. Comparison of logistic regression, support vector machines, and deep learning classifiers for predicting memory encoding success using human intracranial EEG recordings. J. Neural Eng. 15, 066028 (2018).
Lin, J. J. et al. Theta band power increases in the posterior hippocampus predict successful episodic memory encoding in humans. Hippocampus 27, 1040–1053 (2017).
Sederberg, P. B., Kahana, M. J., Howard, M. W., Donner, E. J. & Madsen, J. R. Theta and gamma oscillations during encoding predict subsequent recall. J. Neurosci. 23, 10809–10814 (2003).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Berkel, S. et al. Mutations in the SHANK2 synaptic scaffolding gene in autism spectrum disorder and mental retardation. Nat. Genet. 42, 489–491 (2010).
Peykov, S. et al. Rare SHANK2 variants in schizophrenia. Mol. Psychiatry 20, 1487–1488 (2015).
Won, H. et al. Autistic-like social behaviour in Shank2-mutant mice improved by restoring NMDA receptor function. Nature 486, 261–265 (2012).
Gandal, M. J. et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science 362, eaat8127 (2018).
Dembrow, N. C., Zemelman, B. V. & Johnston, D. Temporal dynamics of L5 dendrites in medial prefrontal cortex regulate integration versus coincidence detection of afferent inputs. J. Neurosci. 35, 4501–4514 (2015).
Silva, L. R., Amitai, Y. & Connors, B. W. Intrinsic oscillations of neocortex generated by layer 5 pyramidal neurons. Science 251, 432–435 (1991).
Kim, E. J., Juavinett, A. L., Kyubwa, E. M., Jacobs, M. W. & Callaway, E. M. Three types of cortical layer 5 neurons that differ in brain-wide connectivity and function. Neuron 88, 1253–1267 (2015).
Xia, F. et al. Parvalbumin-positive interneurons mediate neocortical-hippocampal interactions that are necessary for memory consolidation. eLife https://doi.org/10.7554/eLife.27868 (2017).
Naka, A. et al. Complementary networks of cortical somatostatin interneurons enforce layer specific control. eLife https://doi.org/10.7554/eLife.43696 (2019).
Veit, J., Hakim, R., Jadi, M. P., Sejnowski, T. J. & Adesnik, H. Cortical gamma band synchronization through somatostatin interneurons. Nat. Neurosci. 20, 951–959 (2017).
Pi, H. J. et al. Cortical interneurons that specialize in disinhibitory control. Nature 503, 521–524 (2013).
Lee, S., Kruglikov, I., Huang, Z. J., Fishell, G. & Rudy, B. A disinhibitory circuit mediates motor integration in the somatosensory cortex. Nat. Neurosci. 16, 1662–1670 (2013).
Kamigaki, T. & Dan, Y. Delay activity of specific prefrontal interneuron subtypes modulates memory-guided behavior. Nat. Neurosci. 20, 854–863 (2017).
Pepper, R. E., Pitman, K. A., Cullen, C. L. & Young, K. M. How do cells of the oligodendrocyte lineage affect neuronal circuits to influence motor function, memory and mood? Front. Cell. Neurosci. 12, 399 (2018).
Um, J. W. & Ko, J. LAR-RPTPs: synaptic adhesion molecules that shape synapse development. Trends Cell Biol. 23, 465–475 (2013).
Kantojarvi, K. et al. Fine mapping of Xq11.1-q21.33 and mutation screening of RPS6KA6, ZNF711, ACSL4, DLG3, and IL1RAPL2 for autism spectrum disorders (ASD). Autism Res. 4, 228–233 (2011).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Azevedo, F. A. et al. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J. Comp. Neurol. 513, 532–541 (2009).
Zhang, Y. et al. Polymorphisms in human dopamine D2 receptor gene affect gene expression, splicing, and neuronal activity during working memory. Proc. Natl Acad. Sci. USA 104, 20552–20557 (2007).
Papassotiropoulos, A. et al. Common Kibra alleles are associated with human memory performance. Science 314, 475–478 (2006).
Jacobs, J. & Kahana, M. J. Direct brain recordings fuel advances in cognitive electrophysiology. Trends Cogn. Sci. 14, 162–171 (2010).
Yamazaki, Y. et al. Oligodendrocytes: facilitating axonal conduction by more than myelination. Neuroscientist 16, 11–18 (2010).
Baker, A. et al. Specialized subpopulations of deep-layer pyramidal neurons in the neocortex: bridging cellular properties to functional consequences. J. Neurosci. 38, 5441–5455 (2018).
Carracedo, L. M. et al. A neocortical delta rhythm facilitates reciprocal interlaminar interactions via nested theta rhythms. J. Neurosci. 33, 10750–10761 (2013).
Jinno, S. et al. Neuronal diversity in GABAergic long-range projections from the hippocampus. J. Neurosci. 27, 8790–8804 (2007).
Kim, D. et al. Distinct roles of parvalbumin- and somatostatin-expressing interneurons in working memory. Neuron 92, 902–915 (2016).
Jacobs, J. Hippocampal theta oscillations are slower in humans than in rodents: implications for models of spatial navigation and memory. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130304 (2014).
Yaffe, R. B. et al. Reinstatement of distributed cortical oscillations occurs with precise spatiotemporal dynamics during successful memory retrieval. Proc. Natl Acad. Sci. USA 111, 18727–18732 (2014).
Rutishauser, U., Ross, I. B., Mamelak, A. N. & Schuman, E. M. Human memory strength is predicted by theta-frequency phase-locking of single neurons. Nature 464, 903–907 (2010).
Duzel, E., Penny, W. D. & Burgess, N. Brain oscillations and memory. Curr. Opin. Neurobiol. 20, 143–149 (2010).
Kahana, M. J. The cognitive correlates of human brain oscillations. J. Neurosci. 26, 1669–1672 (2006).
Vaz, A. P., Inati, S. K., Brunel, N. & Zaghloul, K. A. Coupled ripple oscillations between the medial temporal lobe and neocortex retrieve human memory. Science 363, 975–978 (2019).
Lega, B., Burke, J., Jacobs, J. & Kahana, M. J. Slow-theta-to-gamma phase-amplitude coupling in human hippocampus supports the formation of new episodic memories. Cereb. Cortex 26, 268–278 (2016).
Muñoz, M. D., Antolín-Vallespín, M., Tapia-González, S. & Sánchez-Capelo, A. Smad3 deficiency inhibits dentate gyrus LTP by enhancing GABA neurotransmission. J. Neurochem. 137, 190–199 (2016).
Donoghue, T. et al. Parameterizing neural power spectra. Nat. Neurosci. 23, 1655–1665 (2020).
Herweg, N. A., Solomon, E. A. & Kahana, M. J. Theta oscillations in human memory. Trends Cogn. Sci. 24, 208–227 (2020).
Natu, V. S. et al. Stimulation of the posterior cingulate cortex impairs episodic memory encoding. J. Neurosci. 39, 7173–7182 (2019).
Goyal, A. et al. Functionally distinct high and low theta oscillations in the human hippocampus. Nat. Commun. 11, 2469 (2020).
Watrous, A. J., Miller, J., Qasim, S. E., Fried, I. & Jacobs, J. Phase-tuned neuronal firing encodes human contextual representations for navigational goals. eLife https://doi.org/10.7554/eLife.32554 (2018).
Fischl, B. et al. Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33, 341–355 (2002).
Ghose, S., Gleason, K. A., Potts, B. W., Lewis-Amezcua, K. & Tamminga, C. A. Differential expression of metabotropic glutamate receptors 2 and 3 in schizophrenia: a mechanism for antipsychotic drug action? Am. J. Psychiatry 166, 812–820 (2009).
Takahashi, J. S. et al. ChIP-seq and RNA-seq methods to study circadian control of transcription in mammals. Methods Enzymol. 551, 285–321 (2015).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Yeh, Y. H. et al. Transforming growth factor-beta and oxidative stress mediate tachycardia-induced cellular remodelling in cultured atrial-derived myocytes. Cardiovasc. Res. 91, 62–70 (2011).
Del Cid-Pellitero, E., Plavski, A., Mainville, L. & Jones, B. E. Homeostatic changes in GABA and glutamate receptors on excitatory cortical neurons during sleep deprivation and recovery. Front. Syst. Neurosci. 11, 17 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in unique molecular identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311 (2009).
Banerjee-Basu, S. & Packer, A. SFARI Gene: an evolving database for the autism research community. Dis. Models Mech. 3, 133–135 (2010).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
International League Against Epilepsy Consortium on Complex Epilepsies. Genome-wide mega-analysis identifies 16 loci and highlights diverse biological mechanisms in the common epilepsies. Nat. Commun. 9, 5269 (2018).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Savage, J. E. et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat. Genet. 50, 912–919 (2018).
Bipolar Disorder and Schizophrenia Working Group of the Psychiatric Genomics Consortium.Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell 173, 1705–1715.e16 (2018).
Davies, G. et al. Study of 300,486 individuals identifies 148 independent genetic loci influencing general cognitive function. Nat. Commun. 9, 2098 (2018).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Pardinas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Martin, J. et al. A genetic investigation of sex bias in the prevalence of attention-deficit/hyperactivity disorder. Biol. Psychiatry 83, 1044–1053 (2018).
Sohail, M. et al. Polygenic adaptation on height is overestimated due to uncorrected stratification in genome-wide association studies. eLife https://doi.org/10.7554/eLife.39702 (2019).
Hoffmann, T. J. et al. A large multiethnic genome-wide association study of adult body mass index identifies novel loci. Genetics 210, 499–515 (2018).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Estrada, K. et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat. Genet. 44, 491–501 (2012).
Schunkert, H. et al. Large-scale association analysis identifies 13 new susceptibility loci for coronary artery disease. Nat. Genet. 43, 333–338 (2011).
Acknowledgements
We thank the patients for participating in the study and the donors and their families for the additional tissue samples. We also thank K. Gleason for assistance with postmortem samples. G.K. is supported by a Jon Heighten Scholarship in Autism Research at UT Southwestern. This work was supported by NIMH (F30MH105158) to M.R.F.; NIDA (5T32DA007290-25) and NHBLI (1T32HL139438-01A1) to F.A.; NINDS (NS106447), a UT BRAIN Initiative Seed Grant (366582), the Chilton Foundation, and the National Center for Advancing Translational Sciences of the NIH under the Center for Translational Medicine’s award number UL1TR001105 to B.C.L. and G.K.; NINDS (NS107357) to B.C.L.; and NIMH (MH103517), The Chan Zuckerberg Initiative, an advised fund of Silicon Valley Community Foundation (HCA-A-1704-01747), and the James S. McDonnell Foundation 21st Century Science Initiative in Understanding Human Cognition—Scholar Award (220020467) to G.K. Postmortem human tissue samples were obtained from the NIH NeuroBioBank (The Harvard Brain Tissue Resource Center, funded through HHSN-271-2013-537 00030C; the Human Brain and Spinal Fluid Resource Center, VA West Los Angeles Healthcare Center; and the University of Miami Brain Endowment Bank) and the UT Neuropsychiatry Research Program (Dallas Brain Collection). We also thank the UT Southwestern Neuroscience Microscopy Facility for providing imaging resources.
Author information
Authors and Affiliations
Contributions
S.B., B.C.L. and G.K. analyzed the data and wrote the paper. M.R.F. and C.D. collected surgical samples, processed RNA and generated bulk RNA-seq libraries. M.R.F. contributed to the design of the project. F.A. generated the snRNA-seq and snATAC-seq data and performed IHC. A.K. preprocessed the snRNA-seq data. E.C. preprocessed the snATAC-seq data. C.A.T. provided postmortem human brain tissue. S.S. analyzed the oscillation data. B.C.L. conducted all surgical procedures and memory testing. B.C.L. and G.K. designed and supervised the study and provided intellectual guidance. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Andrew Jaffe, Ueli Rutishauser, Ziv Williams, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Data quality control.
a, Box plots depicting the probability of recall for items presented at each serial position. Primacy and recency effects are visible, consistent with expectations for performance in the free recall episodic memory paradigm. Whiskers on box plots represent maximum and minimum values. Boxes extend from the 25th to the 75th percentiles, the center lines represent the median. Loess regression with confidence intervals is superimposed to depict the overall distribution. Smooth curves are shown with 95% confidence bands b, Lag conditional response probability curves in our data (lag CRP), indicating expected temporal clustering behavior. Loess regression with confidence intervals depicts the overall distribution. Smooth curves are shown with 95% confidence bands. c, Boxplot showing the comparison of within-subject variance (across all measured electrodes at each band, blue box plot,) with the variance across subjects (at each band, yellow box plot). Across subjects variance is significantly greater than within-subject variance. Reported p-value from Wilcoxon rank sum test (one-sided with alternative greater). Boxplots extend from the 25th to the 75th percentiles, the center lines represent the median. d, Scatter plot showing the fraction of all BA38 electrodes exhibiting a significant subsequent memory effect at each frequency. We observed significant differences predicting recall success across the frequency spectrum, including the delta and gamma bands. Loess regression with confidence intervals depicts the overall distribution. Smooth curves are shown with 95% confidence bands. e, Distribution of SME values for each brain oscillation and cross-correlation based on Spearman’s rank correlation. f, Barplots showing the fraction of electrodes at which oscillations were detected in each frequency band in the recalled and non-recalled conditions. 85% of electrodes exhibited an oscillation in at least one of the delta, theta, or alpha frequency bands. g, Scatter plot showing individual electrode examples of power curves used for oscillation detection via the MODAL algorithm, both before and after subtraction of the best fit line. h, Principal component analysis of the subjects used for the within-subject analysis. Variance explained by each principal component is highlighted in the axis. i, Barplot showing the variance explained by each covariate adjusted across 10 principal components (wVE) for the within-subject data. Technical, biological and sequencing covariates calculated by PICARD (see Methods) are included. l, Principal component analysis of all the subjects used in this study. PMep = post-mortem epileptic subjects, UT = within-subjects, PMctl = post-mortem healthy subjects. m, Variance explained by each covariate adjusted across 10 principal components (wVE). Type corresponds to the three different types of data included in the analysis (PMep, UT, PMctl). Technical, biological and sequencing covariates calculated by PICARD (see Methods) are included. n, Association between the first two components and covariates based on adjusted gene expression. X-axis corresponds to the -log10(P-value) from linear regression modeling between PCs and covariates.
Extended Data Fig. 2 SME gene robustness and overlap with other tasks.
a, Boxplot showing the difference between F-statistics of the SME genes (Multivariate analysis) compared with the other genes. Stars correspond to the Wilcoxon’s rank sum test (N, Sign = 753, NotSign = 14439; one-sided with alternative greater; p < 0.0001 = ****; Benjamini-Hochberg adjusted: Delta, FDR = 2.3x10−249, Theta, FDR = 3.2x10-205, Alpha, FDR = 4.1x10-140, Beta, FDR = 2.1x10-159, Low Gamma, FDR = 7.2x10-207, High Gamma, FDR = 1.3x10-63). Boxes extend from the 25th to the 75th percentiles and the center lines represent the median. b, Violin plots showing the rho^2 of the genes significantly associated with each brain oscillation. Standard errors are calculated based on the rho^2 distribution of the significantly correlated genes. Dots represent the median rho^2 for the specific brain oscillation. c, Violin plots showing the rho^2 of the genes significantly associated with each brain oscillation (Obs = observed) compared with rho^2 derived from the permutation control analyses (Perm = Permutation). Standard errors are calculated based on the rho^2 distribution of the significantly correlated genes. Dots represent the median rho^2 for the specific brain oscillation. 100 random permutations were applied to calculate the Perm values (see Methods). Stars correspond to the Wilcoxon’s rank sum test (unadjusted, one-sided with alternative greater; p < 0.0001 = ****).
Extended Data Fig. 3 WGCNA highlights modules associated with memory oscillations.
a, Representative network dendrogram for the consensus WGCNA. Heatmap shows the correlation between memory oscillatory signatures and genes. Red = positively correlated, Blue = negatively correlated. b, Heatmap showing the module association between memory oscillatory signatures and module eigengenes (Spearman’s rank correlation). Warm colors represent positive correlations and cool colors represent negative correlations. P-values for each correlation together with exact correlation values are contained within each box. c, Bubble-chart showing the enrichment for 300 SME genes decomposed by brain oscillation. Gradient color represents the -log10(FDR) and bubble size represents the odds ratio (OR) from a Fisher’s exact enrichment test of each module with disease-relevant gene lists. Y-axis shows the brain oscillations labels. X-axis indicates the modules of the present study. d, Boxplots showing the differential connectivity (for example number of edges) between SME genes and non-SME genes in the modules associated with memory oscillatory signatures with SME genes enriched. Stars correspond to the results of a Wilcoxon’s rank sum test (one-side test with alternative greater; p < 0.001 = ****, p < 0.01 = **, p < 0.05 = *; Benjamini-Hochberg adjusted: WM4, FDR = 0.016, WM12, FDR = 0.048, WM21, FDR = 4.5x10-4). Boxes extend from the 25th to the 75th percentiles and the center lines represent the median.
Extended Data Fig. 4 Memory-related modules are enriched for gene co-expression modules associated with neuropsychiatric disorders.
a, Bubble-chart showing the enrichment for loci associated with human traits used as negative controls. Gradient color represents the -log10(FDR) from linkage disequilibrium gene set analysis performed by MAGMA. Y-axis shows the acronyms for the GWAS data utilized for this analysis (see Methods). b, Bubble-chart showing the enrichment for modules of co-expressed genes dysregulated in ASD, SCZ or BD. Gradient color represents the -log10(FDR) and bubble size represents the odds ratio (OR) from a Fisher’s exact enrichment test. Y-axis shows the acronyms for the modules associated with neuropsychiatric disorders utilized for this analysis (see Methods). X-axis shows the modules of the present study. Modules significantly correlated with memory-related oscillations are highlighted in bold text.
Extended Data Fig. 5 snRNA-seq quality control metrices and module enrichment for cell-types dysregulated in cognitive disorders.
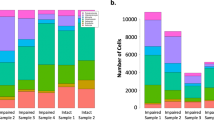
a, Barplot showing the total number of nuclei identified per subject. Colors correspond to the two different batches. b, Quality control boxplots for snRNA-seq with number of genes detected, number of UMIs and percentage of mitochondrial genes. Colors correspond to the two different batches. Boxes extend from the 25th to the 75th percentiles and the center lines represent the median. Dots represent outliers. c, Scatter plot showing the relationship between number of UMIs (X-axis) and detected genes (Y-axis). Each sample is indicated in a different color. d, UMAP plots showing the distribution of nuclei in each subject. Colors correspond to the two different batches. e, Proportion of nuclei representing the identified clusters. Colors correspond to the six different subjects analyzed. f, UMAP plots showing the distribution of the three major cell-classes: GABAergic (blue), Glutamatergic (red), and non-neuronal (green). g, Pie chart showing the proportion of the three major cell-type classes (GABAergic, Glutamatergic, and non-neuronal cells). h, Bubble-chart showing the enrichment of the SME modules for cell-type markers dysregulated in ASD. Color gradient represents the -log10(FDR) and bubble size represent the odds ratio (OR) from a Fisher’s exact enrichment test. Y-axis shows the acronyms for the cell-types defined in the ASD study. i, Bubble-chart showing the enrichment of the SME modules for cell-type markers dysregulated in Alzheimer disease (AD). Color gradient represents the -log10(FDR) and bubble size represent the odds ratio (OR) from a Fisher’s exact enrichment test. Y-axis shows the acronyms for the cell-types defined in the AD study.
Extended Data Fig. 6 snATAC-seq quality control metrices.
a, Barplot showing the total number of nuclei identified per subject. b, Quality control boxplots for each snATAC-seq sample demonstrating the total number of peaks, the number of reads in the peaks and the percentage of reads in peaks. Boxes extend from the 25th to the 75th percentiles and the center lines represent the median. Dots represent outliers. c, Scatter plot showing the relationship between total number of reads (X-axis) and percentage of reads in the peaks (Y-axis). Each sample is indicated in a different color. d, Heatmap of the pairwise similarity between cluster identities. Y-axis shows the snRNA-seq clusters. X-axis shows the snATAC-seq clusters. Gradient corresponds to the percentage of cells for the corresponding prediction label. e, UMAP plots showing the distribution of nuclei in each subject. f, UMAP plots showing the distribution of the three major cell-classes: GABAergic (blue), Glutamatergic (red), and non-neuronal (green). g, Pie chart showing the proportion of the three major cell-type classes.
Supplementary information
Supplementary Table 1
Demographic data. Demographic data with participants, technical covariates, biological covariates, sequencing covariates, SME data, math data, MRI data and behavioral performance metrics.
Supplementary Table 2
Memory biomarkers: gene expression statistics and gene association. All statistics for genes correlated with each brain oscillation and database for association with previous studies.
Supplementary Table 3
Consensus WGCNA modules and gene set association. All statistics for module detection, overlap with memory-correlated genes and database for association with previous studies.
Supplementary Table 4
MAGMA summary statistics for consensus WGCNA modules. All statistics for GWAS enrichment in memory-associated modules.
Supplementary Table 5
snRNA-seq cell-type markers. Genes differentially expressed in each cell type with relative statistics.
Supplementary Table 6
snATAC-seq motif enrichment for peaks associated with module in specific cell types. Motif enrichment statistics for memory modules associated with specific cell types.
Rights and permissions
About this article
Cite this article
Berto, S., Fontenot, M.R., Seger, S. et al. Gene-expression correlates of the oscillatory signatures supporting human episodic memory encoding. Nat Neurosci 24, 554–564 (2021). https://doi.org/10.1038/s41593-021-00803-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-021-00803-x
This article is cited by
-
Functional genomics and systems biology in human neuroscience
Nature (2023)
-
Molecular features driving cellular complexity of human brain evolution
Nature (2023)
-
Genes that give our brains their rhythms
Nature Neuroscience (2021)