Abstract
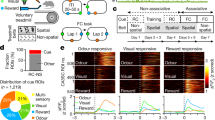
Salient experiences are often relived in the mind. Human neuroimaging studies suggest that such experiences drive activity patterns in visual association cortex that are subsequently reactivated during quiet waking. Nevertheless, the circuit-level consequences of such reactivations remain unclear. Here, we imaged hundreds of neurons in visual association cortex across days as mice learned a visual discrimination task. Distinct patterns of neurons were activated by different visual cues. These same patterns were subsequently reactivated during quiet waking in darkness, with higher reactivation rates during early learning and for food-predicting versus neutral cues. Reactivations involving ensembles of neurons encoding both the food cue and the reward predicted strengthening of next-day functional connectivity of participating neurons, while the converse was observed for reactivations involving ensembles encoding only the food cue. We propose that task-relevant neurons strengthen while task-irrelevant neurons weaken their dialog with the network via participation in distinct flavors of reactivation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request.
Code availability
The code that support the findings of this study is available on Github at https://github.com/asugden/pipe, https://github.com/asugden/flow and https://github.com/asugden/pool. For specific links, see above.
References
Nádasdy, Z., Hirase, H., Czurkó, A., Csicsvari, J. & Buzsáki, G. Replay and time compression of recurring spike sequences in the hippocampus. J. Neurosci. 19, 9497–9507 (1999).
Lee, A. K. & Wilson, M. A. Memory of sequential experience in the hippocampus during slow wave sleep. Neuron 36, 1183–1194 (2002).
Ji, D. & Wilson, M. A. Coordinated memory replay in the visual cortex and hippocampus during sleep. Nat. Neurosci. 10, 100–107 (2007).
Singer, A. C. & Frank, L. M. Rewarded outcomes enhance reactivation of experience in the hippocampus. Neuron 64, 910–921 (2009).
Squire, L. R., Genzel, L., Wixted, J. T. & Morris, R. G. Memory consolidation. Cold Spring Harb. Perspect. Biol. 7, a021766 (2015).
Rothschild, G., Eban, E. & Frank, L. M. A cortical-hippocampal-cortical loop of information processing during memory consolidation. Nat. Neurosci. 20, 251–259 (2016).
Foster, D. J. Replay comes of age. Annu. Rev. Neurosci. 40, 581–602 (2017).
Euston, D. R., Tatsuno, M. & McNaughton, B. L. Fast-forward playback of recent memory sequences in prefrontal cortex during sleep. Science 318, 1147–1150 (2007).
Peyrache, A., Khamassi, M., Benchenane, K., Wiener, S. I. & Battaglia, F. P. Replay of rule-learning related neural patterns in the prefrontal cortex during sleep. Nat. Neurosci. 12, 919–926 (2009).
Xu, S., Jiang, W., Poo, M.-M. & Dan, Y. Activity recall in a visual cortical ensemble. Nat. Neurosci. 15, 449–455 (2012).
Girardeau, G., Inema, I. & Buzsáki, G. Reactivations of emotional memory in the hippocampus–amygdala system during sleep. Nat. Neurosci. 20, 1634–1642 (2017).
Puentes-Mestril, C. & Aton, S. J. Linking network activity to synaptic plasticity during sleep: hypotheses and recent data. Front. Neural Circuits 11, 61 (2017).
Norimoto, H. et al. Hippocampal ripples down-regulate synapses. Science 359, 1524–1527 (2018).
Girardeau, G., Benchenane, K., Wiener, S. I., Buzsáki, G. & Zugaro, M. B. Selective suppression of hippocampal ripples impairs spatial memory. Nat. Neurosci. 12, 1222–1223 (2009).
Jadhav, S. P., Kemere, C., German, P. W. & Frank, L. M. Awake hippocampal sharp-wave ripples support spatial memory. Science 336, 1454–1458 (2012).
Jadhav, S. P., Rothschild, G., Roumis, D. K. & Frank, L. M. Coordinated excitation and inhibition of prefrontal ensembles during awake hippocampal sharp-wave ripple events. Neuron 90, 113–127 (2016).
Xia, F. et al. Parvalbumin-positive interneurons mediate neocortical-hippocampal interactions that are necessary for memory consolidation. eLife 6, 191 (2017).
Maingret, N., Girardeau, G., Todorova, R., Goutierre, M. & Zugaro, M. B. Hippocampo-cortical coupling mediates memory consolidation during sleep. Nat. Neurosci. 19, 959–964 (2016).
Burgess, C. R. et al. Hunger-dependent enhancement of food cue responses in mouse postrhinal cortex and lateral amygdala. Neuron 91, 1154–1169 (2016).
Sacco, T. & Sacchetti, B. Role of secondary sensory cortices in emotional memory storage and retrieval in rats. Science 329, 649–656 (2010).
Grosso, A. et al. The higher order auditory cortex is involved in the assignment of affective value to sensory stimuli. Nat. Commun. 6, 8886 (2015).
Ramesh, R. N., Burgess, C. R., Sugden, A. U., Gyetvan, M. & Andermann, M. L. Intermingled ensembles in visual association cortex encode stimulus identity or predicted outcome. Neuron 100, 900–915 (2018).
Deuker, L. et al. Memory consolidation by replay of stimulus-specific neural activity. J. Neurosci. 33, 19373–19383 (2013).
Murty, V. P., Tompary, A., Adcock, R. A. & Davachi, L. Selectivity in postencoding connectivity with high-level visual cortex is associated with reward-motivated memory. J Neurosci 37, 537–545 (2017).
Logothetis, N. K. et al. Hippocampal-cortical interaction during periods of subcortical silence. Nature 491, 547–553 (2012).
Norman, Y. et al. Hippocampal sharp-wave ripples linked to visual episodic recollection in humans. Science 365, eaax1030 (2019).
Tambini, A., Ketz, N. & Davachi, L. Enhanced brain correlations during rest are related to memory for recent experiences. Neuron 65, 280–290 (2010).
Ko, H. et al. Functional specificity of local synaptic connections in neocortical networks. Nature 473, 87–91 (2011).
Miller, J.-E. K., Ayzenshtat, I., Carrillo-Reid, L. & Yuste, R. Visual stimuli recruit intrinsically generated cortical ensembles. Proc. Natl Acad. Sci. USA 111, E4053–E4061 (2014).
Malvache, A., Reichinnek, S., Villette, V., Haimerl, C. & Cossart, R. Awake hippocampal reactivations project onto orthogonal neuronal assemblies. Science 353, 1280–1283 (2016).
McGinley, M. J., David, S. V. & McCormick, D. A. Cortical membrane potential signature of optimal states for sensory signal detection. Neuron 87, 179–192 (2015).
Kenet, T., Bibitchkov, D., Tsodyks, M., Grinvald, A. & Arieli, A. Spontaneously emerging cortical representations of visual attributes. Nature 425, 954–956 (2003).
Tononi, G. & Cirelli, C. Sleep and the price of plasticity: from synaptic and cellular homeostasis to memory consolidation and integration. Neuron 81, 12–34 (2014).
Paz, R., Bauer, E. P. & Paré, D. Learning-related facilitation of rhinal interactions by medial prefrontal inputs. J. Neurosci. 27, 6542–6551 (2007).
Makino, H. & Komiyama, T. Learning enhances the relative impact of top-down processing in the visual cortex. Nat. Neurosci. 18, 1116–1122 (2015).
Wang, S.-H. & Morris, R. G. M. Hippocampal-neocortical interactions in memory formation, consolidation, and reconsolidation. Annu. Rev. Psychol. 61, 49–79 (2010).
van de Ven, G. M., Trouche, S., McNamara, C. G., Allen, K. & Dupret, D. Hippocampal offline reactivation consolidates recently formed cell assembly patterns during sharp wave-ripples. Neuron 92, 968–974 (2016).
Gomperts, S. N., Kloosterman, F. & Wilson, M. A. VTA neurons coordinate with the hippocampal reactivation of spatial experience. eLife 4, e05360 (2015).
Valdés, J. L., McNaughton, B. L. & Fellous, J.-M. Offline reactivation of experience-dependent neuronal firing patterns in the rat ventral tegmental area. J. Neurophysiol. 114, 1183–1195 (2015).
Timofeev, I. & Chauvette, S. Sleep slow oscillation and plasticity. Curr. Opin. Neurobiol. 44, 116–126 (2017).
Fauth, M. J. & van Rossum, M. C. Self-organized reactivation maintains and reinforces memories despite synaptic turnover. eLife 8, e43717 (2019).
Cossell, L. et al. Functional organization of excitatory synaptic strength in primary visual cortex. Nature 518, 399–403 (2015).
Graf, A. B. A., Kohn, A., Jazayeri, M. & Movshon, J. A. Decoding the activity of neuronal populations in macaque primary visual cortex. Nat. Neurosci. 14, 239–245 (2011).
Leavitt, M. L., Pieper, F., Sachs, A. J. & Martinez-Trujillo, J. C. Correlated variability modifies working memory fidelity in primate prefrontal neuronal ensembles. Proc. Natl Acad. Sci. USA 114, E2494–E2503 (2017).
Atherton, L. A., Dupret, D. & Mellor, J. R. Memory trace replay: the shaping of memory consolidation by neuromodulation. Trends Neurosci. 38, 560–570 (2015).
Ambrose, R. E., Pfeiffer, B. E. & Foster, D. J. Reverse replay of hippocampal place cells is uniquely modulated by changing reward. Neuron 91, 1124–1136 (2016).
Ólafsdóttir, H. F., Carpenter, F. & Barry, C. Task demands predict a dynamic switch in the content of awake hippocampal replay. Neuron 96, 925–934 (2017).
Jung, M. W., Lee, H., Jeong, Y., Lee, J. W. & Lee, I. Remembering rewarding futures: a simulation-selection model of the hippocampus. Hippocampus 28, 913–930 (2018).
Ludvig, E. A., Mirian, M. S., Kehoe, E. J. & Sutton, R. S. Associative learning from replayed experience. Preprint at bioRxiv https://doi.org/10.1101/100800 (2017).
Mattar, M. G. & Daw, N. D. Prioritized memory access explains planning and hippocampal replay. Nat. Neurosci. 21, 1609–1617 (2018).
Madisen, L. et al. Transgenic mice for intersectional targeting of neural sensors and effectors with high specificity and performance. Neuron 85, 942–958 (2015).
Goldey, G. J. et al. Removable cranial windows for long-term imaging in awake mice. Nat. Protoc. 9, 2515–2538 (2014).
Asaad, W. F. & Eskandar, E. N. A flexible software tool for temporally-precise behavioral control in Matlab. J. Neurosci. Methods 174, 245–258 (2008).
Wang, Q. & Burkhalter, A. Area map of mouse visual cortex. J. Comp. Neurol. 502, 339–357 (2007).
Bonin, V., Histed, M. H., Yurgenson, S. & Reid, R. C. Local diversity and fine-scale organization of receptive fields in mouse visual cortex. J. Neurosci. 31, 18506–18521 (2011).
Thevenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Mukamel, E. A., Nimmerjahn, A. & Schnitzer, M. J. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63, 747–760 (2009).
Ziv, Y. et al. Long-term dynamics of CA1 hippocampal place codes. Nat. Neurosci. 16, 264–266 (2013).
Petreanu, L. et al. Activity in motor–sensory projections reveals distributed coding in somatosensation. Nature 489, 299–303 (2012).
Pnevmatikakis, E. A. et al. Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Sheintuch, L. et al. Tracking the same neurons across multiple days in Ca2+ imaging data. Cell Rep. 21, 1102–1115 (2017).
Webb, G. I., Boughton, J. R. & Wang, Z. Not so naive Bayes: aggregating one-dependence estimators. Mach. Learn. 58, 5–24 (2005).
Sugden, L. A. et al. Localization of adaptive variants in human genomes using averaged one-dependence estimation. Nat. Commun. 9, 703 (2018).
Aarts, E., Verhage, M., Veenvliet, J. V., Dolan, C. V. & van der Sluis, S. A solution to dependency: using multilevel analysis to accommodate nested data. Nat. Neurosci. 17, 491–496 (2014).
Singmann, H., Bolker, B., Westfall, J., Højsgaard, S. & Fox, J. Package ‘afex’: Analysis of Factorial Experiments, vsn 0.13–145 https://cran.r-project.org/web/packages/afex/ (2015).
Driscoll, L. N., Pettit, N. L., Minderer, M., Chettih, S. N. & Harvey, C. D. Dynamic reorganization of neuronal activity patterns in parietal cortex. Cell 170, 986–999 (2017).
Saramäki, J., Kivelä, M., Onnela, J.-P., Kaski, K. & Kertész, J. Generalizations of the clustering coefficient to weighted complex networks. Phys. Rev. E 75, 027105–4 (2007).
Komiyama, T. et al. Learning-related fine-scale specificity imaged in motor cortex circuits of behaving mice. Nature 464, 1182–1186 (2010).
Kim, M.-H., Znamenskiy, P., Iacaruso, M. F. & Mrsic-Flogel, T. D. Segregated subnetworks of intracortical projection neurons in primary visual cortex. Neuron 100, 1313–1321.e6 (2018).
Hagberg, A. A., Swart, P. J. & Schult, D. A. Exploring network structure, dynamics, and function using NetworkX. In Proc. 7th Python in Science Conference (SciPy2008) (eds Varoquaux, G., Vaught, T. & Millman, J.) 11–15 (SciPy Organizers, 2008).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008–P10013 (2008).
Green, D. M. & Swets, J. A. Signal Detection Theory and Psychophysics (Wiley, 1966).
Roy, N. A., Bak, J. H., Akrami, A., Brody, C. D. & Pillow, J. W. Efficient inference for time-varying behavior during learning. Adv. Neural Inf. Process. Syst. 31, 5695–5705 (2018).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Acknowledgements
We thank G. Goldey for performing initial surgeries, and N. Patel, M. Barbini, G. Niyazov and E. Bamberg for help with mouse training. We thank B. Healy, R. Born, J. Drugowitsch, I. Witten, C. Harvey, Y. Livneh, R. Buckner and members of the Andermann laboratory for useful discussion. We thank V. Jayaraman, R. Kerr, D. Kim, L. Looger, K. Svoboda and the GENIE Project at Janelia Farm Research Campus, Howard Hughes Medical Institute for the use of GCaMP6f, and H. Zeng for sharing GCaMP6f transgenic mice before publication. Support was provided by NIH (no. T32 5T32DK007516 to A.U.S.), NSF (no. GRFP 2016207224 to K.L.M.), NIH (F31 105678 to R.N.R., F32 DK112589-01 to A.L. and R01 DK109930 to M.L.A.), RCN (250259 to K.K.L.), a Davis Family Foundation Postdoctoral Fellowship (to C.R.B.), an NIH Director’s New Innovator Award (DP2 DK105570), a McKnight Scholar Award, a Pew Scholar Award, a Smith Family Foundation Award, a Harvard Mind Brain Behavior Interfaculty Initiative Faculty Research Award and grants from the Klarman Family Foundation, the American Federation for Aging Research and the Harvard Brain Science Initiative Bipolar Disorder Seed Grant, supported by K. and L. Dauten (to M.L.A.).
Author information
Authors and Affiliations
Contributions
A.U.S. and M.L.A. designed experiments and analyses and wrote the manuscript. A.U.S., L.A.S. and M.L.A. designed the classifier. A.U.S. performed the data analysis with statistics designed by J.D.Z. J.D.Z., R.N.R. and C.R.B. contributed additional analyses. A.L. and A.U.S. designed the photometry experiments. O.A. performed the surgeries. O.A. and A.U.S. acquired the data. K.L.M. performed the electrophysiology recordings. K.L.M., A.U.S. and K.K.L. analyzed the electrophysiology data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
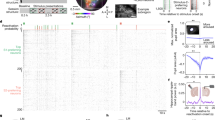
Extended Data Fig. 1 Correlation of behavioral performance and session number and examples of offline cue reactivation events in darkness.
a. Top: Go-NoGo visual discrimination task, and description of the one-time change in cue-outcome contingencies (“reversal”). Bottom: behavioral performance as a function of number of training sessions relative to day of reversal of cue-outcome contingencies (‘day 0’). Data from each of the eight mice are plotted using a unique color. See also Ramesh et al., 2018. b. Left: average cue-evoked response time courses (rows) from each simultaneously recorded cell from a single session. Data are aligned to the onset of the visual cues, each of which was 2 s in duration. ∆F/F0: fractional change in fluorescence. Bottom right: peri-reactivation deconvolved activity (black) of all cells during the post-training and post-satiation ~1 h period of recording in darkness. Data correspond to the 15 food-cue reactivation events with the highest classifier matches (classifier output shown on top), and are sorted in order of decreasing classifier output (top panel; roughly similar to a matching probability, see Methods and Supplementary Information). c. Same plot as in b, but for aversive-cue reactivations during the same example session. d. Portion of an example recording from a different mouse from that shown in Fig. 2a–c, from a sated mouse. Bottom: deconvolved activity of 235 simultaneously recorded cells. Behavioral and other variables are also shown: brain motion (root-mean-squared motion in the imaging plane; red, top), running speed (blue, top), pupil diameter (orange, top), licking (purple ticks, top), output of the classifier for each cue (colored lines, middle; red: aversive cue; blue: neutral cue; green: food cue).
Extended Data Fig. 2 Further characterization of our classifier, including temporal components and those related to pairwise co-activation of neurons.
We designed a classifier that uses an averaged one-dependence estimator (AODE) – an extension to Naïve Bayes that accounts for pairwise probabilities (Webb et al., 2005; Sugden et al., 2018). a. Mean deconvolved population activity, centered on cue reactivation events found with a version of this classifier that did not include a temporally varying prior. b. Construction of the temporally varying prior. We used three high-pass difference-of-Gaussians kernels to generate the population activity scale factor that modulates the prior probability employed to estimate the probability of a reactivation event. We took the minimum value across the convolutions of each of these three kernels with the mean deconvolved activity across cells, in order to define a temporally varying prior at each time point (see Methods and Supplementary Information for additional details). Each filter is constructed by subtracting from a narrow Gaussian (sigma: 41 imaging frames, 260 ms) one of three broad Gaussians (sigmas: 42, 43, and 44 imaging frames, corresponding to 1 s, purple; 4 s, blue; and 16 s, green respectively), to account for background fluctuations at multiple slower timescales than the timescales expected for cue reactivations (based on both previous studies and on the results of panel a). c. Visualization of the effects of the temporally varying prior – a high-pass temporal filter applied to each cell’s activity time course that biases classification towards brief, synchronous events. High-pass filtering the deconvolved data in Fig. 2c by applying this filter separately to each cell time course shows that the temporally varying prior selectively enhances transient events. Transient events that are not marked as reactivation events by the classifier may be associated with hippocampal sharp-wave ripples, but were not classified as reactivations of one of the three cues, as they were not sufficiently similar to cue responses (right panel; see also Fig. 2a, b). Note that the same temporal filter is applied to all cell time courses, and thus does not modify the relative activity across neurons at the time of cue reactivation. d. Cue reactivation events can occur in bursts with inter-event intervals of ~1 s. These short inter-event intervals in the true distribution of inter-event intervals (black line) are beyond what is expected by chance (95% confidence interval of shuffle is shaded in gray, estimated by assigning each event to a time drawn uniformly at random within the session in which the event was identified). e. Violin plot of rate of cue reactivation events in darkness (width is proportional to relative incidence), for a classifier using Naïve Bayes (left) vs. AODE (right). Black horizontal lines indicate means across sessions, and black vertical lines indicate SEM across sessions. Accounting for pairwise probabilities of co-active neurons using the AODE doubles the number of identified reactivation events for the same classifier output threshold (p < 0.0001, two-tailed Wilcoxon rank-sum test). f. Clusters were defined using the functional connectivity metric of noise correlations, measured across all three cues. Circles represent the activity of neurons (represented as in Fig. 2d; response probabilities of cue-driven neurons are represented by the diameter of circles along outer ring, and joint response probabilities are proportional to the thickness of connecting lines). Cue reactivation events detected by the classifier using AODE but not by the classifier using Naïve Bayes (right column) were often dominated by active cells belonging to a small number of clusters (gray shaded wedges). In contrast, events detected by both the classifier using AODE and the classifier using Naïve Bayes (left column) included active neurons spanning multiple clusters.
Extended Data Fig. 3 The classifier detects few reactivation events following randomization in cell identity or in time.
a. Fraction of cue presentations (colored lines) and inter-trial intervals (gray line) correctly identified by our AODE classifier when trained on two-thirds of the data and tested on the remaining third (mean ± SEM across 109 sessions), as a function of classifier output threshold. Green, blue and red lines correspond to food cue, neutral cue and aversive cue, respectively. b. To demonstrate that the classifier is not identifying random fluctuations in activity during quiet waking periods as cue reactivation events, we plotted the number of reactivation events for each cue type that would still be found following shuffling of cell identities in identified reactivation events compared to real data, as a function of the classifier output threshold (see also Methods and Supplementary Information). There are few events found in randomized data at any classifier output threshold greater than 0.05. c. The fraction of cue reactivation events found in data randomized by identity above our chosen classifier output threshold (same as Fig. 2f, replicated for comparison). Compared to the number of events that exceeded the classifier output threshold of 0.1 in the real data (which was the threshold we used in our analyses), we found a relatively small proportion in the randomized data. Results were similar for a modified classifier using Naïve Bayes (not shown). d. Similarly, we plotted the fraction of these events that would still be found at times of classifier-identified cue reactivation events, but in data randomized across cell identities in which we matched activity levels across cells. Specifically, at time points identified as cue reactivation events, we randomized the cell identities, and determined how many cue reactivation events (across all three cue types) were found above our chosen classifier output threshold of 0.1. e. As in c, the fraction of cue reactivation events found in data randomized in identity with matched activity levels. f. As in b, the fraction of events identified in data randomized in time relative to real data, as a function of classifier output threshold. This was estimated by randomizing data in time via circularly shifting each cell’s time course with random relative delays (by offsetting each time course in time by a random amount for each cell, and wrapping the clipped portion of the time course beyond the end of the recording back to the beginning). g. As in c, the fraction of cue reactivation events identified in data randomized in time. h. Distribution of the number of identified events (across all sessions from all mice, in runs following satiation), for events of each classifier output threshold (from 0.05-1 in bins of 0.05). Dashed gray line represents our chosen classifier output threshold for defining an activity pattern as a cue reactivation event. Note that food-cue reactivation events were more common than neutral-cue reactivation events across a range of classifier output thresholds. All randomizations were performed using 109 sessions across 8 mice, and bars and lines represent the mean ± SEM.
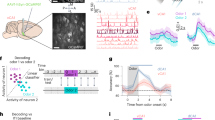
Extended Data Fig. 4 Additional data on combined imaging and hippocampal electrophysiology; reactivation events are not a result of epileptiform events, transient brain motion, or eye motion.
a. Schematic for two-photon calcium imaging in visual association cortex while recording from contralateral hippocampal area CA1 using a silicon multi-electrode probe. b. Same example aversive-cue reactivation event as in Fig. 2g, but with additional data shown. Bottom left: mean activity time courses (fractional change in fluorescence, ∆F/F0) in response to food cues, neutral cues, and aversive cues (columns) for all simultaneously recorded neurons (rows) from the same session, organized by preferred response. Bottom right: deconvolved activity traces of all recorded cortical cells in the period surrounding the aversive-cue reactivation event (red arrow). Top: purple traces: CA1 local field potential trace and zoom-in. When band-pass filtered in the ripple band (150-300 Hz; gray trace), the LFP shows a transient increase just before the detected reactivation event. c-e. The Emx1-Cre;Ai93;CaMK2a-tTA mouse line (Madisen et al., 2015) used in our work has been reported to exhibit epileptiform events in some cases (Steinmetz et al., 2017). Early experiments in our lab using mice obtained directly from the Allen Brain Institute did appear to exhibit visible epileptiform events as detected in the neuropil signal in cortical imaging. While none of these events overlapped with any cue reactivation events, we excluded this mouse from the study. We then acquired mice from Jackson labs (line 024108 rather than 024103 used in (Steinmetz et al., 2017)) which may have been further back-crossed. To further address the possibilities that epileptiform events occurred in the 8 mice used in our study and that these might overlap with the cue reactivations described in our study, we characterized the amplitude and width of all transient events in the neuropil from spontaneous activity recordings in all of our mice (as in (Steinmetz et al., 2017)). As described below, we found that 7/8 mice did not show any evidence of epileptiform events, while 1/8 mice only demonstrated a small number of epileptiform events. Panel c: scatter plot of peak width vs. event peak prominence (height above local background), combined across 7/8 mice included in this study. Typical low-amplitude or high-width peaks (Steinmetz et al., 2017) are shown as blue dots, whereas brief, high-amplitude peaks that may reflect epileptiform events are shown as black dots (defined using a conservative manual threshold as in (Steinmetz et al., 2017)). In these 7 mice, we found almost no epileptiform events total across over a dozen sessions per mouse (0,0,0,0,1,2,3 events total per mouse; events rates ranged between 0 and 0.00003 epileptiform events/second). Panel d: same as c, but for the eighth mouse included in the current study. In this mouse, we found 87 epileptiform events total across all sessions, amounting to 0.0014 epileptiform events/second, or approximately one such ~150 ms event every 12 min. Across all sessions, a total of two epileptiform events were observed within 1 s of any reactivation event, and both of these were neutral-cue reactivation events. Panel e: same as c, but for the mouse excluded from this dataset. Detectable epileptiform events were very rare in this mouse (0.008 events/s). Critically, we found that across all 8 included mice, 0/1444 food cue reactivations, 0/1258 aversive cue reactivations, and 2/1040 neutral cue reactivations occurred within 1 s of an epileptiform event (and the two overlapping neutral cue reactivations were from a mouse with 2 epileptiform events detected in total across all sessions). Thus, epileptiform events are exceptionally rare in the mice used in this study, and those that occurred did not occur near cue reactivation events. To further confirm these observations, we re-analyzed the 7 imaging sessions from two mice in which we imaged from the identical mouse line while recording cortical and hippocampal local field potentials using a 16-channel silicon probe acutely implanted contralateral to the imaging window (see Fig. 2g). The epileptiform events previously described using cortical electrophysiology (Steinmetz et al., 2017) appear as large local field potential (LFP) spikes, characterized by high amplitude (>1000 µV; normal LFP amplitudes are 5-15x smaller) and long duration (>10 ms). Briefly, we used the exact methods as in Steinmetz et al., 2017, and analyzed 20-s LFP traces surrounding 250 cue reactivation events identified using contralateral imaging (5,000 s of data). We found a total of 5 epileptiform events from our LFP data (average rate of 1 events / 1000 s). Critically, all 5 of these events occurred >2 s away from any cue reactivation event. f. Pupil area (normalized to mean area estimated during locomotion in darkness, a state involving dilated pupils) was constricted throughout reactivation events. g. There was no transient increase in brain motion (root-mean-squared motion in the imaging plane) in the moments surrounding a reactivation event (green: food-cue reactivations; blue: neutral-cue reactivations; red: aversive-cue reactivations). h. There was no transient increase in eye motion (root-mean-squared motion) in the moments surrounding a reactivation event. Error bars are ± SEM across sessions (f) and across events (g, h).
Extended Data Fig. 5 Additional evidence supporting a learning-related decrease and a food-cue bias in reactivation rates, and a reactivation-dependent change in future behavioral performance.
a. Reactivation rates were significantly higher in sessions following task engagement (black dots: reactivation event counts per session for all 3 cues across 109 training sessions) than in sessions in which naïve animals viewed the same visual stimuli but in the absence of salient outcomes (that is, sessions during initial habituation to the stimuli but prior to any conditioning; gray dots: reactivation event counts per session for all 3 cues across 11 sessions). Data were fit using a GLMM accounting for shared variance within mice (see Methods and Supplementary Information). b. The number of recorded cells did not change across learning (r and p: Pearson correlation; error bars: 95% confidence intervals; N = 109 sessions). c. The overall population activity across all neurons (measured as the mean deconvolved activity across quiet waking periods) did not decrease across learning (fit using the same 109 sessions as in b). d, e. Example plots from a single animal using PsyTrack (Roy et al., 2018) to dynamically determine behavioral variables. d. Individual behavioral variables fit across time in this animal. The vertical black bar represents the time of change in cue-outcome associations (that is, reversal), and thin gray vertical bars separate different days. Colored lines for each angle (0°, 135°, 270°) reflect behavioral sensitivity to a grating drifting at that angle (colors of gratings in legend refer to the cue contingencies associated with that grating prior to and following reversal, as shown using the vertical black line; green: food cue; red: aversive cue; blue: neutral cue). Variables labeled ‘Prev choice’, ‘Prev punish’, and ‘Prev reward’ refer to effects on current performance related to choice (Go or NoGo) or outcome from the previous trial, and ‘Offset’ reflects the baseline licking rate. Error bars: 95% posterior credibility interval (the Bayesian equivalent of a confidence interval). e. Example of behavioral performance calculated per day (orange) and estimated dynamically (blue) across time in the same animal (Roy et al., 2018). Green dots indicate performance at the beginning of each day and red dots indicate performance at the end of each day. f. The change in behavioral performance from the beginning to the end of a training session is not associated with cue reactivation rate during the subsequent quiet waking period (for any of the three cues; same 109 sessions as a–c). r: Spearman correlation. p-values were determined using a GLMM accounting for shared variance within mice (see Methods and Supplementary Information). g. Cumulative distribution of single-session cue reactivation rates (same data as in Fig. 3a). h. Similar to Fig. 3c, d, the food-cue is preferentially reactivated when using the Naïve Bayes classifier instead of the AODE classifier (*** p < 0.001, ** p < 0.01; using a GLMM comparing the effect of cue on the rates of reactivation accounting for shared variance within mice and days), both for sessions prior to reversal (65 sessions) and following reversal (44 sessions). Inset at right shows relative reactivation rates for each session and cue type (normalized to the neutral rate for that session) using Naïve Bayes, colored by cue type. i. Fraction of responsive cells driven by a particular cue (same data as in Fig. 3e but with data points for each session shown). For each session, the fraction of responsive cells driven by a particular cue was normalized to the overall number of cue-responsive cells. Data reflect mean ± SEM across sessions. Thus, the observed enhancement in food cue reactivation rates was not due to a difference in the number of neurons responsive to any given cue (no significant difference, two-tailed Wilcoxon rank-sum test), as we reversed cue contingencies or stopped recording at the point in training where a food-cue bias typically emerges, after the mouse becomes very well trained (Burgess et al., 2016; data not shown). j. Unlike Fig. 3f, reactivation rates of neutral and aversive cues do not significantly predict changes in behavioral performance across sessions (though weak trends are evident). We used a GLMM that accounts for shared variance within mice and for number of days elapsed between the two training sessions. Line: GLMM fit across mice when considering 1 day of elapsed time between sessions. r: Spearman correlation p-values indicate the effect of reactivation rate from the GLMM. Error bars represent mean ± SEM across sessions in panels b-c (109 sessions) and h-i (65, 44, and 109 sessions).
Extended Data Fig. 6 Estimation and simulation of total functional connectivity.
a. A description of the steps taken to compute the change in total connectivity across sequential pairs of days. Step i. For a given day, we generate a matrix containing the mean response, from 0-2 s after the food cue onset, for each trial and each cell driven by the food cue on that day. Step ii. We then compute the matrix of pairwise noise correlations for these cells (described in Methods and Supplementary Information). Step iii. Correlation matrices can be represented as weighted graphs with edge weights defined by the correlation coefficients. As such, we create a weighted graph with food-cue-driven cells as nodes, and using positive noise correlations as edge weights. Step iv. We compute the total connectivity per cell, defined as the clustering coefficient for each node (that is, the geometric mean across connected edges; Hagberg et al., 2008). The measure of total connectivity thus reflects the notion that pairs of cells having stronger noise correlations are more strongly connected and/or receive stronger common input. Step v. For each cell identified on both of two consecutive days (that is, cells lacking an ‘x’ symbol on both days), we calculate the difference in the cell’s total connectivity (clustering coefficient) between the two days. Further analysis (as in Fig. 4d and Fig. 5e) compares this difference in total connectivity between two different groups of cells–cells that either did or did not participate in inter-session reactivation events. b. We considered whether the computation of the total connectivity metric might introduce correlations between estimates of connectivity of simultaneously recorded cells (as each cell’s estimate involved averaging across the pairwise correlations with the other cells), and thereby invalidate the assumption of conditional independence of data points from simultaneously recorded cells in the GLMM analyses used in Figs. 4d and 5e. Therefore, we simulated noise correlation matrices from two distributions (for the two groups of reactivated vs. non-reactivated cells, see above and further described in Methods) and computed changes in total connectivity (steps iii-v) between sequential pairs of these matrices. We found that this metric does introduce a random shift in the change in total connectivity that is common to all cells recorded within a given day-pair (top panel), but that these shifts are accounted for by using a generalized linear mixed regression model that includes fixed offsets for each day-pair. We demonstrated this by measuring, for each pair of simulated cells, the Pearson correlation of changes in total connectivity across multiple day pairs (after subtracting off the mean shift across cells for each day-pair; bottom panel). The resulting correlation was found to be significant in only approximately one out of every 20 simulations (that is, p = 0.05), as expected if there was no underlying correlation introduced by using the total connectivity metric. Given this result, we concluded that there was no correlation introduced by the total connectivity metric above and beyond a common mean effect across cells recorded on any given day-pair, further justifying the use of the GLMM in these analyses. See Data analysis and statistics section of Methods and Supplementary Information for additional details.
Extended Data Fig. 7 Generalized linear model for categorizing groups of lateral visual association cortex neurons that encode one or multiple task-related variables; further analyses of cells that also encode reward.
a. Mean deconvolved activity of cell groups identified by a generalized linear model (see Identification of cell responses using a GLM section of Methods) surrounding times of task-related events (for example, cue presentations, motor behavior, reward delivery). Rows, top to bottom: cell types that only respond to the food cue, to the neutral cue, and to the aversive cue, those that only respond in a time-locked manner at moments surrounding the reward presentation or surrounding the punishment presentation, and those with activity time-locked to both the food cue and to the reward presentation. Time zero for each column in a corresponds, from left to right, to the time of food cue onset, neutral cue onset, aversive cue onset, Ensure reward delivery onset, quinine delivery onset, and onset of a lick bout (that is, onset of a lick sequence after ≥ 2 s without any licks). N’s at left indicate numbers of neurons. b. Mean deconvolved activity of cell groups identified by a generalized linear model (see Methods) surrounding reactivation events. The peak cue amplitude for those cells that respond only to the food cue, neutral cue, or aversive cue (top 3 rows) has been scaled to match the amplitude during reactivation events (gray). The other rows (that is, other cell categories) have had their cue-evoked responses scaled by an equal amount. As such, the level of activity during reactivations can be compared with the expected level given the response magnitude of cells of a given category during the cue period used by our classifier. Error bars are ± SEM across the same sets of cells as in a. c. Cells encoding both the reward and the food cue (that is, Food-cue-reward cells) are more common than cells encoding both the reward and other cues (two-tailed Wilcoxon rank-sum test, Bonferroni corrected for 3 comparisons, *** p < 0.0001; mean ± SEM across sessions). d–f. Heatmaps of individual trials (row) from example reward-coding cells that increase their activity at different times relative to reward delivery. Display of data is identical to Fig. 4f.
Extended Data Fig. 8 Additional data demonstrating distinct clusters of cells with strong intra-cluster correlations in trial-by-trial cue responses; comparison of cells belonging to reward-related vs. non-reward-related clusters.
a. The clustering algorithm used in Fig. 2d and Fig. 5 (see Methods) clusters cells into groups that share high within-group, cue-driven, trial-by-trial noise correlations. This algorithm automatically chooses the optimal number of clusters for each session. The number of clusters per session did not differ between stimulus types (two-tailed Wilcoxon rank-sum test). b. Cluster sizes (number of cells per cluster) also did not differ between stimulus types (two-tailed Wilcoxon rank-sum test). c. Within-cluster noise correlations were substantially higher than between-cluster noise correlations (see Methods; two-tailed Wilcoxon rank-sum test; *** p < 0.0001). d. Same violin plots as Fig. 5a, but using the Naïve Bayes classifier. These data confirms that the finding in Fig. 5a is not due to the fact that the AODE classifier used in Fig. 5a includes pairwise activity as part of the information used for identification of reactivation events. e. Similar plots as in d, but separately examining subsets of pairs of cells considered in d that belong to reward-related clusters (pink) or to non-reward-related clusters (orange; two-tailed Wilcoxon rank-sum test; *** p < 0.0001). f. Bottom: CDFs and KDEs of the distributions of cross-day changes in total functional connectivity. Reactivated food-cue-driven cells from reward clusters (purple, 194 cells) increased their next-day total connectivity compared to all non-reactivated food-cue-driven cells (gray, 238 cells). Conversely, reactivated food-cue-driven cells from non-reward clusters (orange, 245 cells) decreased their next-day total functional connectivity. P-values were computed using a linear GLMM involving a categorical comparison involving 3 categories: non-reactivated cells, cells from reward clusters, and cells from non-reward clusters. Permutation tests within days demonstrated that cells from non-reward clusters had significantly decreased next-day connectivity when reactivated (* p < 0.05; see Methods). Top: excess frequency of reactivated cells with a given change in next-day connectivity above that observed for non-reactivated cells (that is, rectified difference of colored vs. gray distributions). g. Data from Fig. 5c, presented as a correlation matrix between all pairs of cells and a hierarchical clustering dendrogram. The clusters identified on the left are labeled on the right. Note the distinct reward-related clusters and non-reward-related clusters. h. Cluster size (that is the number of cells per cluster), with 1 dot per cluster for all clusters from all recordings, separated by reward-related clusters (purple) or non-reward-related clusters (orange). i. Number of clusters of a given type per session. Approximately 44% of sessions contained at least one cluster of each type. j. Within both Day 1 and Day 2 of each pair of days (more specifically, on Day i relative to a pair of days [i, i + 1]), the total functional connectivity was not different between cells belonging to reward-related clusters vs. those belonging to non-reward-related clusters. The overall responsivity of each cell group was also unchanged across days and did not differ between groups (two-tailed Wilcoxon rank-sum test). k. Total connectivity on Day 1 was indistinguishable between (i) all food-cue-driven cells in reward-related clusters, (ii) Food-cue cells in reward-related clusters, defined as cells that are responsive to food cues but not to rewards, and (iii) Food-cue cells in non-reward-related clusters (p = 0.81, Kruskal-Wallis test). This was true both when comparing the subsets of cells that did not participate in any food-cue reactivations on Day 1 (left bars), and when comparing the subsets of cells that participated in at least one food-cue reactivation on Day 1 (right bars, p = 0.96, Kruskal-Wallis test). All error bars are mean ± SEM across sessions, including in violin plots (some bars are too small to show).
Extended Data Fig. 9 Different flavors of food-cue reactivation events from an example session; additional analyses of cross-day changes in total connectivity.
a. Plot similar to that shown in Fig. 5c, but for a different example session. Response probabilities of cue-driven neurons are represented by the diameters of circles along the outer ring, and joint response probabilities are proportional to the thickness of connecting lines. Food-cue-driven neurons were clustered into groups based on their functional connectivity (food-cue-evoked noise correlations; c.f. Figure 5c). The two ‘reward-related clusters’ (purple wedges) contained at least one Food-cue-reward cell, while the ‘non-reward-related cluster’ (orange wedge) did not. b. Left: as also described in Fig. 5d, food-cue reactivation events could be composed primarily of active food-cue-driven cells from non-reward-related clusters (“#1: reward-poor reactivations”, top), from reward-related clusters (“#3: reward-rich reactivations”, bottom), or of similar numbers of active food-cue-driven cells of both types (“#2: mixed reactivations”, middle). Right: Across 1444 reactivations from 109 sessions in 8 mice, we found a substantial number of food-cue reactivation events of each flavor. Plotted is the kernel density estimate (KDE, or smoothed distribution of frequencies), across cue reactivation events, of the fraction of reactivated cells from reward-related clusters (a value of 1 indicates that all food-cue-driven cells active during the reactivation are from reward-related clusters; a value of 0 indicates that all food-cue-driven cells active during the reactivation are from non-reward-related clusters). Also plotted is the CDF. The large incidence of reactivation events with proportions equaling 0 or 1 is due to sessions in which food-cue-driven cells belonged either entirely to non-reward-related clusters, or to reward-related clusters, respectively. Vertical dashed lines indicate the thresholds that were chosen to categorize food-cue reactivations into one of the three groups defined above (compare circled numbers in left and right panels). c. Left: mean activity time courses (fractional change in fluorescence, ∆F/F0) in response to food cues, neutral cues, and aversive cues (columns) for all simultaneously recorded neurons (rows) from the same example session as in a and organized by preferred response. Right: deconvolved activity time courses (1 row/cell) in the 0.5 s surrounding each food-cue reactivation event (plotted in a similar manner as in Extended Data Fig. 1b-c). Reactivations are sorted, in descending order, by strength of classifier output (posterior estimates; green lines at top). Thick horizontal colored bars below the classifier output estimates are shaded from orange to purple and indicate the proportion of participating food-cue-driven cells that are from reward-related clusters (see color bar at bottom of panel b). Circular plots below each reactivation event indicate which cells participated in the event, and their membership in reward-related or in non-reward-related clusters. Note that the first 8 reactivations are mixed reactivations, while the ninth reactivation (at right) is a reward-rich reactivation, as most participating cells belong to reward-related clusters. d. Same as c but for the next nine food-cue reactivations from the same session (again in descending order of estimates of strength of classifier output). e. Left: changes in total connectivity for non-reactivated cells in Fig. 4d, plotted on the same axis. Right: changes in total connectivity for reactivated Food-cue cells in non-reward-related clusters, Food-cue cells in reward-related clusters, and Food-cue-reward cells. Note that while reactivated cells increased total connectivity more when they belonged to reward-related clusters, non-reactivated cells decreased connectivity more when they belonged to reward-related clusters.
Extended Data Fig. 10 Additional analyses and control experiments regarding changes in VTANAc axon activity during cue reactivations.
a. Setup for simultaneous two-photon calcium imaging in cortex and contralateral fiber photometry recording in NAc of putative VTANAc axons. b. Left: average time courses of bulk GCaMP6 fiber photometry responses in nucleus accumbens (mainly from dopaminergic VTANAc axons; Rodriguez-Lopez et al., 2017) to visual food-cue presentations during training. Responses are averages of ‘hit’ trials (green) in which the mice made an operant response in the response window (2-4 s after cue onset) and received a reward, or ‘miss’ trials in which the mice did not respond in the response window. Percent change in GCaMP6 fluorescence, ∆F/F0. Mean ± SEM across trials, pooled across 33 sessions from 3 mice (mouse #1: 16 sessions; mouse #2: 8 sessions; mouse #3: 9 sessions). Middle and right: same analysis but for neutral cue and aversive cue trials, separated by ‘correct-reject’ trials (no response during response window) and ‘false-alarm’ trials (lick response during response window). Gray bars represent the time of cue presentation and black bars represent the first second of the response window that follows offset of cue presentation. Licking in this response window results in reward delivery, no outcome, or quinine delivery during food-cue trials, neutral-cue trials, or aversive-cue trials, respectively. While some GCaMP6f-labeled axons from EMX1-Cre-expressing neurons do innervate the NAc in this mouse line (as we used this transgenic mouse to track cortical cue reactivations), and while the AAV-GCAMP6s infection of VTA likely led to expression in both dopaminergic and non-dopaminergic NAc-projecting neurons, the observed cue-evoked responses in these NAc recordings show a strong response bias to the rewarded food cue and a decrease in activity in the response window following aversive-cue trials (particularly for false-alarm trials), which closely match the activity profile expected for putative VTA dopaminergic neurons projecting to NAc (Roitman et al., 2008; Cohen et al., 2012; Parker et al., 2016; Lutas et al., 2019). c. Left: reward-rich food-cue reactivations during quiet waking evoked increases in activity in putative VTANAc axons, while reward-poor food-cue reactivations during quiet waking evoked decreases. Mixed reactivations had little effect on the photometry signal. The fraction of reward-rich cells active during a food-cue reactivation predicted the direction and amplitude of the VTANAc fluorescence response between 0.5 and 1 s after reactivation event onset (* p < 0.05, categorical GLMM; see Methods and Supplementary Information). Middle and right: average change in putative VTANAc axon activity following offline neutral-cue reactivations and aversive-cue reactivations, respectively. Mean ± SEM across cue reactivations. Two-tailed Wilcoxon sign-rank tests mean evoked response in the 0.5-1 s after the reactivation event vs. 0 (p-values indicated in panels). d. Heatmaps showing all individual cue reactivation-triggered fractional changes in VTANAc fluorescence for the data plotted in the middle and right panels of c. Black horizontal lines separate groups of trials from each of the three mice. Note the consistent drop in activity (blue) following most reactivation events. e. Similar to d, but for activity during reward-rich, mixed, and reward-poor reactivations (cf. panel c, left).
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and Supplementary Information.
Rights and permissions
About this article
Cite this article
Sugden, A.U., Zaremba, J.D., Sugden, L.A. et al. Cortical reactivations of recent sensory experiences predict bidirectional network changes during learning. Nat Neurosci 23, 981–991 (2020). https://doi.org/10.1038/s41593-020-0651-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0651-5