Abstract
Tumor-associated macrophages (TAMs) play an important role in the immune response to cancer, but the mechanisms by which the tumor microenvironment controls TAMs and T cell immunity are not completely understood. Here we report that kynurenine produced by glioblastoma cells activates aryl hydrocarbon receptor (AHR) in TAMs to modulate their function and T cell immunity. AHR promotes CCR2 expression, driving TAM recruitment in response to CCL2. AHR also drives the expression of KLF4 and suppresses NF-κB activation in TAMs. Finally, AHR drives the expression of the ectonucleotidase CD39 in TAMs, which promotes CD8+ T cell dysfunction by producing adenosine in cooperation with CD73. In humans, the expression of AHR and CD39 was highest in grade 4 glioma, and high AHR expression was associated with poor prognosis. In summary, AHR and CD39 expressed in TAMs participate in the regulation of the immune response in glioblastoma and constitute potential targets for immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
13 June 2019
In the version of this article initially published, author Alexandre Prat’s surname was misspelled. The error has been corrected in the HTML and PDF versions of the article.
References
De Henau, O. et al. Overcoming resistance to checkpoint blockade therapy by targeting PI3Kγ in myeloid cells. Nature 539, 443–447 (2016).
Kaneda, M. M. et al. PI3Kγ is a molecular switch that controls immune suppression. Nature 539, 437–442 (2016).
Quail, D. F. et al. The tumor microenvironment underlies acquired resistance to CSF-1R inhibition in gliomas. Science 352, aad3018 (2016).
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2014).
Condeelis, J. & Pollard, J. W. Macrophages: obligate partners for tumor cell migration, invasion, and metastasis. Cell 124, 263–266 (2006).
Franklin, R. A. et al. The cellular and molecular origin of tumor-associated macrophages. Science 344, 921–925 (2014).
Junttila, M. R. & de Sauvage, F. J. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 501, 346–354 (2013).
Noy, R. & Pollard, J. W. Tumor-associated macrophages: from mechanisms to therapy. Immunity 41, 49–61 (2014).
Qian, B. Z. et al. CCL2 recruits inflammatory monocytes to facilitate breast-tumour metastasis. Nature 475, 222–225 (2011).
Qian, B. Z. & Pollard, J. W. Macrophage diversity enhances tumor progression and metastasis. Cell 141, 39–51 (2010).
Preusser, M., Lim, M., Hafler, D. A., Reardon, D. A. & Sampson, J. H. Prospects of immune checkpoint modulators in the treatment of glioblastoma. Nat. Rev. Neurol. 11, 504–514 (2015).
Sharma, P. & Allison, J. P. The future of immune checkpoint therapy. Science 348, 56–61 (2015).
Wen, P. Y. & Reardon, D. A. Neuro-oncology in 2015: progress in glioma diagnosis, classification and treatment. Nat. Rev. Neurol. 12, 69–70 (2016).
Hambardzumyan, D., Gutmann, D. H. & Kettenmann, H. The role of microglia and macrophages in glioma maintenance and progression. Nat. Neurosci. 19, 20–27 (2016).
Thomas, A. A. et al. Regulatory T cells are not a strong predictor of survival for patients with glioblastoma. Neuro Oncol. 17, 801–809 (2015).
Pyonteck, S. M. et al. CSF-1R inhibition alters macrophage polarization and blocks glioma progression. Nat. Med. 19, 1264–1272 (2013).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nat. Rev. Immunol. 12, 253–268 (2012).
Gabriely, G., Wheeler, M. A., Takenaka, M. C. & Quintana, F. J. Role of AHR and HIF-1α in glioblastoma metabolism. Trends Endocrinol. Metab. 28, 428–436 (2017).
Opitz, C. A. et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature 478, 197–203 (2011).
Gutiérrez-Vázquez, C. & Quintana, F. J. Regulation of the immune response by the aryl hydrocarbon receptor. Immunity 48, 19–33 (2018).
Mascanfroni, I. D. et al. Metabolic control of type 1 regulatory T cell differentiation by AHR and HIF1-α. Nat. Med. 21, 638–646 (2015).
Bessede, A. et al. Aryl hydrocarbon receptor control of a disease tolerance defence pathway. Nature 511, 184–190 (2014).
Rothhammer, V. et al. Microglial control of astrocytes in response to microbial metabolites. Nature 557, 724–728 (2018).
Mukthavaram, R. et al. Effect of the JAK2/STAT3 inhibitor SAR317461 on human glioblastoma tumorspheres. J. Transl Med. 13, 269 (2015).
Thota, B. et al. STAT-1 expression is regulated by IGFBP-3 in malignant glioma cells and is a strong predictor of poor survival in patients with glioblastoma. J. Neurosurg. 121, 374–383 (2014).
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat. Med. 22, 586–597 (2016).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Bowman, R. L. et al. Macrophage ontogeny underlies differences in tumor-specific education in brain malignancies. Cell Rep. 17, 2445–2459 (2016).
Platten, M. et al. Monocyte chemoattractant protein-1 increases microglial infiltration and aggressiveness of gliomas. Ann. Neurol. 54, 388–392 (2003).
Mayo, L. et al. Regulation of astrocyte activation by glycolipids drives chronic CNS inflammation. Nat. Med. 20, 1147–1156 (2014).
Phillips, R. J., Lutz, M. & Premack, B. Differential signaling mechanisms regulate expression of CC chemokine receptor-2 during monocyte maturation. J. Inflamm. (Lond.) 2, 14 (2005).
Sozzani, S. et al. MCP-1 and CCR2 in HIV infection: regulation of agonist and receptor expression. J. Leukoc. Biol. 62, 30–33 (1997).
Platten, M. et al. Treatment of autoimmune neuroinflammation with a synthetic tryptophan metabolite. Science 310, 850–855 (2005).
Liao, X. et al. Krüppel-like factor 4 regulates macrophage polarization. J. Clin. Invest. 121, 2736–2749 (2011).
Yeste, A. et al. Tolerogenic nanoparticles inhibit T cell-mediated autoimmunity through SOCS2. Sci. Signal. 9, ra61 (2016).
Takenaka, M. C., Robson, S. & Quintana, F. J. Regulation of the T cell response by CD39. Trends Immunol. 37, 427–439 (2016).
Deaglio, S. et al. Adenosine generation catalyzed by CD39 and CD73 expressed on regulatory T cells mediates immune suppression. J. Exp. Med. 204, 1257–1265 (2007).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T Cells. Cell 166, 1500–1511.e9 (2016).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Nirschl, C. J. et al. IFNγ-dependent tissue-immune homeostasis is co-opted in the tumor microenvironment. Cell 170, 127–141 (2017).
Wang, Q. et al. Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32, 42–56.e6 (2017).
Wherry, E. J. T cell exhaustion. Nat. Immunol. 12, 492–499 (2011).
Kohanbash, G. et al. Isocitrate dehydrogenase mutations suppress STAT1 and CD8+ T cell accumulation in gliomas. J. Clin. Invest. 127, 1425–1437 (2017).
Thaiss, C. A., Zmora, N., Levy, M. & Elinav, E. The microbiome and innate immunity. Nature 535, 65–74 (2016).
Vétizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Okey, A. B., Vella, L. M. & Harper, P. A. Detection and characterization of a low affinity form of cytosolic Ah receptor in livers of mice nonresponsive to induction of cytochrome P1-450 by 3-methylcholanthrene. Mol. Pharmacol. 35, 823–830 (1989).
Gabriely, G. et al. Targeting latency-associated peptide promotes antitumor immunity. Sci. Immunol. 2, 2 (2017).
Bennett, M. L. et al. New tools for studying microglia in the mouse and human CNS. Proc. Natl Acad. Sci. USA 113, E1738–E1746 (2016).
Durafourt, B. A., Moore, C. S., Blain, M. & Antel, J. P. Isolating, culturing, and polarizing primary human adult and fetal microglia. Methods Mol. Biol. 1041, 199–211 (2013).
David, B. A. et al. Isolation and high-dimensional phenotyping of gastrointestinal immune cells. Immunology 151, 56–70 (2017).
Mascanfroni, I. D. et al. IL-27 acts on DCs to suppress the T cell response and autoimmunity by inducing expression of the immunoregulatory molecule CD39. Nat. Immunol. 14, 1054–1063 (2013).
Shin, S. et al. NRF2 modulates aryl hydrocarbon receptor signaling: influence on adipogenesis. Mol. Cell. Biol. 27, 7188–7197 (2007).
Karpurapu, M. et al. Krüppel like factor 4 promoter undergoes active demethylation during monocyte/macrophage differentiation. PLoS One 9, e93362 (2014).
Acknowledgements
This work was supported by the following grants: NS087867, ES02530, AI126880 and AI093903 from the National Institutes of Health (NIH) and RSG-14-198-01-LIB from the American Cancer Society to F.J.Q.; the Brain Science Foundation grant to G.Gabriely and ES07381 to D.S. M.C.T. received support from an educational grant from Mallinkrodt Pharmaceuticals (A219074). M.A.W. was supported by a training grant from the NIH (1F32NS101790) and the Dana Farber Cancer Institute (T32CA207201). V.R. was supported by a fellowship from the German Research Foundation (DFG RO4866 1/1) and an educational grant from Mallinkrodt Pharmaceuticals (A219074). C.G.-V. is supported by an Alfonso Martín Escudero Foundation postdoctoral fellowship. A.B. was supported by a doctoral fellowship (CBB BDS-00503-16) from FAPEMIG, Brazil. We thank T. Seyfried (Biology Department, Boston College, MA, USA) for providing the CT2A cell line.
Author information
Authors and Affiliations
Contributions
M.C.T., V.R., I.D.M., G.Gabriely, C.-C.C., M.A.W., C.G.-V., K.A.L., J.K., A.B., L.M. and T.V. performed the in vitro and in vivo experiments. S.G., S.Z., A.P. and M.A.W. performed the immunofluorescence studies. E.C.T., H.Z. and M.F.F. performed the bioinformatics analysis, and S.R., D.S., J.A., S.C.R., S.G., L.H., G.Getz, D.A.R and H.L.W. provided unique reagents. M.C.T., G.Gabriely and F.J.Q. discussed and interpreted the findings. M.C.T. and F.J.Q. wrote the manuscript and F.J.Q. designed and supervised the study.
Corresponding author
Ethics declarations
Competing interests
F.J.Q. is a member of the Scientific Advisory Board of Kyn Therapeutics. D.S. is a co-founder of and holds equity in Hercules Pharmaceuticals. The other authors declare no competing interests.
Additional information
Journal peer review information: Nature Neuroscience thanks Ingo Mellinghoff, Judith Varner, and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Effect of TCM on human macrophages.
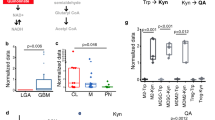
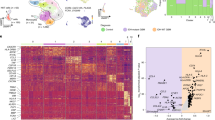
(a) Heatmap of M1- and M2-like genes in GBM-infiltrating CD14+ cells (GBM) and microglia from healthy individuals (Control) using Nanostring (n = 3 biologically independent samples). (b) Expression of Ahr and M2-like genes in BMDMs stimulated with TCM from CT2A cells per 24 hours (n = 3 technical replicates). Data are representative of two independent experiments with similar results. Unpaired two-tailed t test was used for statistical analysis. (c) Relative gene expression of CYP1B, KLF4, MCR1, LLGL1, STAT3, STAT1, CD274, IL10 and ENTPD1 in CD14+ blood cells from healthy donors treated with TCM from different glioma cell lines per 24 hours. DMEM sample was used as reference sample. Each symbol represents an individual (n = 4 biologically independent samples). Ordinary one-way ANOVA was used for statistical analysis. (d) Left panels. Representative immunofluorescence images of human gliomas stained for in the top CD68 (green), TMEM119 (red), AHR (cyan) and nucleus (blue), in the middle AHR (green), CD4 (red), CD3 (cyan) and nucleus (blue) and in the bottom AHR (green), CD8 (red), CD3 (cyan) and nucleus (blue). Right panels. Quantification of AHR+CD68+TMEM119Neg cells in tumor from grade 1 (n = 2), grade 2 (n = 14), grade 3 (n = 8) and grade 4 (n = 9) (top); AHR+CD4+CD3+ cells (middle) and AHR+CD8+CD3+ cells (bottom) in tumor from grade 1 (n = 5), grade 2 (n = 9), grade 3 (n = 13) and grade 4 (n = 8). Each symbol represents one individual and all data are mean ± s.e.m. Kruskal-Wallis test was used for statistical analysis. Scale bars, 20 μm.
Supplementary Figure 2 Immunoblot analysis of AHR, NFkBp65 and TRAF6.
(a) Immunoblots of AHR (top) and GAPDH (bottom) in total protein lysates of RAW264.7 macrophages overexpressing or not the miR-29b, corresponding to the immunoblot in the Fig. 2n. Control samples (1, 3, 5), miR-29b transfected cell samples (2, 4, 6). (b) Representative immunoblots of AHR (upper panel) and GAPDH (lower panel) in total protein lysates of BMDMs from WT and AHRLysM mice, corresponding to the immunoblot in the Supplementary Fig. 3b. BMDMs WT (1, 3 and 5) and AHRLysM (2, 4 and 6) samples. (c) Western Blot analysis of NF-κB (p65) (upper panel), GAPDH (middle panel) and histone 3 (lower panel) in cytoplasmatic and nuclear fraction of BMDMs stimulated or not with TCM for 90 minutes from WT and AHRLysM mice, corresponding to the immunoblot in the Fig. 4l. Samples: 1 (control WT), 2 (TCM WT), 3 (control AHRLysM) and 4 (TCM AHRLysM). (d) Immunoblots of TRAF6 (upper panel) and GAPDH (lower panel) in total protein lysates of BMDMs stimulated or not with TCM for 6 hours from WT and AHRLysM mice, corresponding to the immunoblot in the Supplementary Fig. 4k. Samples: 1 (control WT), 2 (TCM WT), 3 (control AHRLysM) and 4 (TCM AHRLysM). L: ladder.
Supplementary Figure 3 AHR expression in WT and AHRLysM mice.
(a) Ahr expression in sorted bone marrow inflammatory monocytes (CD3NegB220NegLy6GNeg NK1.1NegSiglec-FNegCD11b+Ly6CHi) from naive WT and AHRLysM mice (n = 3 independent mice). Representative of two independent experiments with similar results. (b) Western-blot analysis of AHR in BMDMs from WT and AHRLysM mice (n = 3 biologically independent samples). Data are representative of two independent experiments with similar results; images were cropped and the full scans are shown in Supplementary Fig. 2b. (c-e) qPCR analysis of Ahr expression in microglia (c), B cells (CD19+) and T cells (CD3+ CD4+ and CD3+ CD8+) (d), neutrophils (CD115NegCD11b+CD11cNegLy6G+) and dendritic cells (CD115NegCD11bNegCD11c+ MHCIIHi) (e) (n = 3 independent mice). (f) Frequency of neutrophils and dendritic cells in the CD45+CD11b+ gate of glioma-infiltrating cells. CyTOF analysis was performed in WT mice 15 days after GL261 cell implantation. The dot plot graphs are representative of two independent experiments with similar results. (g) Frequency of microglia, splenic B cells and T cells in naive WT and AHRLysM mice (n = 4 independent mice). (h) Total number of macrophages (CD11b+F4/80+) and inflammatory monocytes (CD11b+F4/80+Ly6CHi) in spleen from WT and AHRLysM (n = 4 independent mice). (i) Survival curve analysis from WT and AHRLysM mice implanted intracranially with CT2A cells (n = 10 independent mice). Representative of two independent experiments with similar results. Survival analysis was performed using a Kaplan-Meier plot using a log-rank (Mantel-Cox) test. (j) Flow cytometry analysis of TMEM119 (blue), CX3CR1 (red) and Ly6C (magenta) expression in TAMs, 15 days after GL261 implantation in WT mice. Representative of two independent experiments with similar results. (k) Ly6C expression on TAMs gate (LinNegCD11b+CD45+) in GBM from WT and AHRLysM mice on day 15 (n = 3 independent mice). In the left, representative dot plot graphs of Ly6C expression in TAMs gate, where microglia (CD45LowCD11b+) is shown in blue and peripheral infiltrated macrophages (CD45HiCD11b+) in red. Percentage of Ly6C+ cells in TAMs (right panel). Representative of two independent experiments with similar results. (l, m) Ccl2 and Ccl7 gene expression in naïve brain and GL261 tumor-bearing mice (l) and in CT2A cells (m) (n = 3 tmice per group). Representative of two independent experiments with similar results. (n) Quantification of CCL2 in TCM from human GBM cells by ELISA (n = 5 biologically independent samples). Unpaired two-tailed t test was used to compare two groups (a-d, g, h, k, m and n) and one-way ANOVA was used to compare three or more groups (e and l). All data are presented as mean ± s.e.m.
Supplementary Figure 4 Kyn regulates TAM polarization in vivo via AHR.
(a) Nanostring analysis of peripheral infiltrated macrophages (LinNegCD11b+CD45Hi) in GBM from WT and AHRLysM mice 15 days after GL261 cells implantation (pool of 4 mice per group). Ingenuity pathway analysis of macrophage polarization genes is shown. (b) Tumor size in WT mice 7 days after implantation of GL261-control and GL261-TDO/IDO cells (n = 3 independent mice). Representative images and quantification (left and right, respectively). Data are representative of two independent experiments with similar results. (c,e) Ido1 and Tdo2 expression in tumor tissue from WT mice injected with GL261- control and GL261-TDO/IDO (n = 3 independent mice) (c), and in CT2A cells (n = 3 biologically independent samples) (e). Representative of two independent experiments with similar results. (d) Arg1 expression in sorted TAMs from WT mice injected with GL261-control and GL261-TDO/IDO cells (n = 3 independent mice). Representative of two independent experiments with similar results. (f) Schematic of AHR binding sites (XREs) in the Klf4 promoter. The arrows indicate primers designed to study AHR (sites 1-6) recruitment. (g,h) Klf4 gene was silenced by siRNA in bone marrow derived macrophages. The cells were stimulated with TCM from GL261 cells for 24 hours and gene expression of M1- and M2-like genes was analyzed by qPCR (n = 3 technical replicates). Data are representative of two independent experiments with similar results. (i) Ingenuity Pathway Analysis of NF-κB signaling using data from Nanostring analysis of peripheral infiltrated macrophages (LinNegCD11b+CD45Hi) in GBM from WT and AHRLysM mice 15 days after GL261 cells implantation (pool of 4 mice per group). Colors indicate up- and down-regulation of individual pathway components in red and green, respectively. (j,k) Socs2 expression (j) and TRAF6 protein levels (k) in TCM-stimulated BMDMs from WT and AHRLysM mice. Data in j were repeated two times with similar results, with three technical replicates. The experiment in k was repeated two times with similar results; images were cropped and the full scans are shown in Supplementary Fig. 2d. All data are presented as mean ± s.e.m, Unpaired two-tailed t test was used to compare two groups (b-e, h and j) and one-way ANOVA was used to compare three or more groups (g).
Supplementary Figure 5 CD39 in TAMs drives T cell dysfunction.
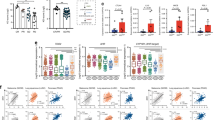
(a) Schematic of generation of LysMCre CD39fl/fl mice. There is insertion of a LoxP site into the exon 5 and the exon 6 in the mouse Entpd1 gene. (b) MFI of CD39 expression determined by flow cytometry in splenic macrophages (F4/80+CD11b+) from naive WT and CD39LysM mice. Data are representative of two independent experiments. (c, d) Percentage of CD39 in CD4+ T cells (c) and in Treg cells (CD4+ FoxP3+) (d) in the blood of WT and CD39LysM mice (n = 5 independent mice). On the left of each figure are the representative dot plots from each group. (e) Frequency of CD3+ T cells, CD3+CD8+ T cells, CD3+CD8+ T cells and Tregs (FoxP3+ CD4+) in the blood from WT and CD39LysM mice (n = 5 independent mice). (f) Effect of CD8+ T-cell depletion in WT and AHRLysM mice performed as previously described 52. Survival curve analysis of GBM mice injected with GL261 cells in WT and AHRLysM (n = 8 independent mice). Log-rank test was used to compare survival among the Isotype WT and Isotype AHRLysM groups, or between Isotype and anti-CD8 treated mice in WT group or in AHRLysM group. Representative of two independent experiments with similar results. (g,h) Flow cytometry analysis of TILs in WT and AHRLysM mice 15 days after GL261 cells implantation. Total number of CD8+ TILs (g) and frequency of PD-1+granzyme B+ cells in CD8+ T cells and of Treg cells (FoxP3+ CD4+) (h). Data are representative of two independent experiments with similar results, using three mice per group. (i,j) Flow cytometry analysis of TILs in WT and AHRLysM mice 28 days after CT2A-luciferase cells implantation. Total number of CD8+ TILs (i) and frequency of IFN-γ+ in CD8+ T cells and of Treg cells (FoxP3+ CD4+) (j). Data are representative of two independent experiments with similar results, using three mice per group. Unpaired two-tailed t test was used to compare two groups (c-e, g-j). All data are shown as mean ± s.e.m.
Supplementary Figure 6 Proposed model of the effects of GBM-produced AHR agonists on TAM function and T cell immunity.
Kyn released by glioma cells activates AHR in TAMs, modulating TAM recruitment into GBM via CCR2, TAM polarization via KLF4/NF-κB, and T-cells via CD39.
Supplementary Figure 7 Gate strategy used to analyze TAMs.
FACS gating strategy for TAM purification from GBM- infiltrated mononuclear cell suspensions.
Supplementary information
Supplementary Figures 1–7.
Supplementary Table 1.
Supplementary Table 2
Data used to construct the heatmap shown in Fig. 6b.
Supplementary Table 3
Data from TCGA used to analyze CCR2 and CCL2 expression in different subtypes of GBM patients shown in Fig. 6c.
Supplementary Table 4
Data used for univariate and multivariate analysis of overall survival of TCGA GBM patients shown in Table 1.
Supplementary Table 5
Data used for univariate and multivariate analysis of overall survival of TCGA GBM patients shown in Supplementary Table 1.
Supplementary Software 1
Code used to generate the results shown in Table 1.
Supplementary Software 2
Code used to generate the results shown in Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Takenaka, M.C., Gabriely, G., Rothhammer, V. et al. Control of tumor-associated macrophages and T cells in glioblastoma via AHR and CD39. Nat Neurosci 22, 729–740 (2019). https://doi.org/10.1038/s41593-019-0370-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0370-y
This article is cited by
-
Metabolic regulation of tumor-associated macrophage heterogeneity: insights into the tumor microenvironment and immunotherapeutic opportunities
Biomarker Research (2024)
-
Lactate dehydrogenase A regulates tumor-macrophage symbiosis to promote glioblastoma progression
Nature Communications (2024)
-
NF-κB in biology and targeted therapy: new insights and translational implications
Signal Transduction and Targeted Therapy (2024)
-
Aryl hydrocarbon receptor suppresses STING-mediated type I IFN expression in triple-negative breast cancer
Scientific Reports (2024)
-
Novel STAT3 oligonucleotide compounds suppress tumor growth and overcome the acquired resistance to sorafenib in hepatocellular carcinoma
Acta Pharmacologica Sinica (2024)