Abstract
The contribution of lineage identity and differentiation state to malignant transformation is controversial. We have previously shown that adult neural stem and early progenitor cells give origin to glioblastoma. Here we systematically assessed the tumor-initiating potential of adult neural populations at various stages of lineage progression. Cell type–specific tamoxifen-inducible Cre recombinase transgenes were used to target glioblastoma-relevant tumor suppressors Nf1, Trp53 and Pten in late-stage neuronal progenitors, neuroblasts and differentiated neurons. Mutant mice showed cellular and molecular defects demonstrating the impact of tumor suppressor loss, with mutant neurons being the most resistant to early changes associated with tumor development. However, we observed no evidence of glioma formation. These studies show that increasing lineage restriction is accompanied by decreasing susceptibility to malignant transformation, indicating a glioblastoma cell-of-origin hierarchy in which stem cells sit at the apex and differentiated cell types are least susceptible to tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data Availability
Sequence data supporting the findings in this paper are publicly accessible in Gene Expression Omnibus as GSE117258. The data supporting the findings of this study are available from the corresponding authors upon request.
References
Sanai, N., Alvarez-Buylla, A. & Berger, M. S. Neural stem cells and the origin of gliomas. N. Engl. J. Med. 353, 811–822 (2005).
Bond, A. M., Ming, G. L. & Song, H. Adult mammalian neural stem cells and neurogenesis: five decades later. Cell Stem Cell 17, 385–395 (2015).
Gage, F. H. & Temple, S. Neural stem cells: generating and regenerating the brain. Neuron 80, 588–601 (2013).
Lim, D. A. & Alvarez-Buylla, A. The adult ventricular-subventricular zone (V-SVZ) and olfactory bulb (OB) neurogenesis. Cold Spring Harb. Perspect. Biol. 8, 8 (2016).
Huse, J. T. & Holland, E. C. Genetically engineered mouse models of brain cancer and the promise of preclinical testing. Brain Pathol. 19, 132–143 (2009).
Alcantara Llaguno, S. et al. Malignant astrocytomas originate from neural stem/progenitor cells in a somatic tumor suppressor mouse model. Cancer Cell 15, 45–56 (2009).
Alcantara Llaguno, S. R. et al. Adult lineage-restricted CNS progenitors specify distinct glioblastoma subtypes. Cancer Cell 28, 429–440 (2015).
Galvao, R. P. et al. Transformation of quiescent adult oligodendrocyte precursor cells into malignant glioma through a multistep reactivation process. Proc. Natl Acad. Sci. USA 111, E4214–E4223 (2014).
Chow, L. M. et al. Cooperativity within and among Pten, p53, and Rb pathways induces high-grade astrocytoma in adult brain. Cancer Cell 19, 305–316 (2011).
Holland, E. C. et al. Combined activation of Ras and Akt in neural progenitors induces glioblastoma formation in mice. Nat. Genet. 25, 55–57 (2000).
Bachoo, R. M. et al. Epidermal growth factor receptor and Ink4a/Arf: convergent mechanisms governing terminal differentiation and transformation along the neural stem cell to astrocyte axis. Cancer Cell 1, 269–277 (2002).
Friedmann-Morvinski, D. et al. Dedifferentiation of neurons and astrocytes by oncogenes can induce gliomas in mice. Science 338, 1080–1084 (2012).
Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Griffith, L. C. Calcium/calmodulin-dependent protein kinase II: an unforgettable kinase. J. Neurosci. 24, 8391–8393 (2004).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Luikart, B. W. et al. TrkB has a cell-autonomous role in the establishment of hippocampal Schaffer collateral synapses. J. Neurosci. 25, 3774–3786 (2005).
Zhu, Y. et al. Ablation of NF1 function in neurons induces abnormal development of cerebral cortex and reactive gliosis in the brain. Genes Dev. 15, 859–876 (2001).
Kwon, C. H. et al. Pten regulates neuronal arborization and social interaction in mice. Neuron 50, 377–388 (2006).
Doetsch, F., Caillé, I., Lim, D. A., García-Verdugo, J. M. & Alvarez-Buylla, A. Subventricular zone astrocytes are neural stem cells in the adult mammalian brain. Cell 97, 703–716 (1999).
Aprea, J., Nonaka-Kinoshita, M. & Calegari, F. Generation and characterization of Neurod1-CreER(T2) mouse lines for the study of embryonic and adult neurogenesis. Genesis 52, 870–878 (2014).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011).
Mich, J. K. et al. Prospective identification of functionally distinct stem cells and neurosphere-initiating cells in adult mouse forebrain. eLife 3, e02669 (2014).
Oien, K. A. & Dennis, J. L. Diagnostic work-up of carcinoma of unknown primary: from immunohistochemistry to molecular profiling. Ann. Oncol. 23(Suppl 10), x271–x277 (2012).
Kwon, C. H. et al. Pten haploinsufficiency accelerates formation of high-grade astrocytomas. Cancer Res. 68, 3286–3294 (2008).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45(D1), D353–D361 (2017).
Herculano-Houzel, S. & Lent, R. Isotropic fractionator: a simple, rapid method for the quantification of total cell and neuron numbers in the brain. J. Neurosci. 25, 2518–2521 (2005).
Zhu, Y. et al. Early inactivation of p53 tumor suppressor gene cooperating with NF1 loss induces malignant astrocytoma. Cancer Cell 8, 119–130 (2005).
Horton, L. B. et al. Mutagenesis of zinc ligand residue Cys221 reveals plasticity in the IMP-1 metallo-β-lactamase active site. Antimicrob. Agents Chemother. 56, 5667–5677 (2012).
Alcantara Llaguno, S. R. & Parada, L. F. Cell of origin of glioma: biological and clinical implications. Br. J. Cancer 115, 1445–1450 (2016).
Uhrbom, L. et al. Ink4a-Arf loss cooperates with KRas activation in astrocytes and neural progenitors to generate glioblastomas of various morphologies depending on activated Akt. Cancer Res. 62, 5551–5558 (2002).
Kim, J. et al. Reprogramming of postnatal neurons into induced pluripotent stem cells by defined factors. Stem Cells 29, 992–1000 (2011).
Huntly, B. J. et al. MOZ-TIF2, but not BCR-ABL, confers properties of leukemic stem cells to committed murine hematopoietic progenitors. Cancer Cell 6, 587–596 (2004).
Tomasetti, C. & Vogelstein, B. Cancer etiology. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science 347, 78–81 (2015).
Eriksson, P. S. et al. Neurogenesis in the adult human hippocampus. Nat. Med. 4, 1313–1317 (1998).
Sorrells, S. F. et al. Human hippocampal neurogenesis drops sharply in children to undetectable levels in adults. Nature 555, 377–381 (2018).
Boldrini, M. et al. Human hippocampal neurogenesis persists throughout aging. Cell Stem Cell 22, 589–599.e5 (2018).
Gould, E., Reeves, A. J., Graziano, M. S. & Gross, C. G. Neurogenesis in the neocortex of adult primates. Science 286, 548–552 (1999).
Curtis, M. A. et al. Human neuroblasts migrate to the olfactory bulb via a lateral ventricular extension. Science 315, 1243–1249 (2007).
Lee, J. H. et al. Human glioblastoma arises from subventricular zone cells with low-level driver mutations. Nature 560, 243–247 (2018).
Warrell, R. P. Jr, de Thé, H., Wang, Z. Y. & Degos, L. Acute promyelocytic leukemia. N. Engl. J. Med. 329, 177–189 (1993).
Sell, S. Stem cell origin of cancer and differentiation therapy. Crit. Rev. Oncol. Hematol. 51, 1–28 (2004).
Louis, D. N. et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 131, 803–820 (2016).
Lin, S. C. et al. Somatic mutation of p53 leads to estrogen receptor alpha-positive and -negative mouse mammary tumors with high frequency of metastasis. Cancer Res. 64, 3525–3532 (2004).
Flores, I. et al. The longest telomeres: a general signature of adult stem cell compartments. Genes Dev. 22, 654–667 (2008).
Mi, H. et al. PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 45(D1), D183–D189 (2017).
Acknowledgements
The authors thank D. Laks for processing of the RNA sequencing data and members of the Parada laboratory for helpful suggestions and discussion. We thank N. Socci, E. Feng and V. Boyko for their help in various analyses, and the MSKCC Genomics and Bioinformatics Cores and the Weill Cornell Genomics and Epigenomics Cores for their assistance. This work was supported in part by the Children’s Tumor Foundation Young Investigator Award and National Institutes of Health (NIH) T32 Postdoctoral Training Grant (2T32CA124334-06; PI: Jerry Shay) to S.A.L. L.F.P. is a recipient of NIH R01 grant CA131313-01A1 and National Cancer Institute R35 grant CA210100.
Author information
Authors and Affiliations
Contributions
S.A.L. and L.F.P. conceptualized the study. S.A.L., D.S., A.M.P. and E.V. performed experiments. Z.W. contributed reagents and material. S.A.L., D.S., A.M.P., E.V., D.K.B. and L.F.P. analyzed the data. S.A.L and L.F.P. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following financial competing interests: L.F.P. has advisory or consulting relationships with Bio-Thera Pharmaceuticals (2013–2018), Howard Hughes Medical Institute Scientific Advisory Board (2006–2023) and the National Cancer Institute Board of Scientific Advisors (2013–2018).
Additional information
Journal peer review information Nature Neuroscience thanks Martine Roussel, Mario Suva and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Histologic and molecular analysis of iCK-cre mutants.
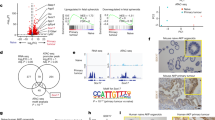
a. Immunofluorescence staining for NeuN in different brain regions. b. Quantification of indicated neuronal lineage markers in iCK-cre mutant (M) vs. control (C) brains. (Left-most panel) Parvalbumin, n=12M,14C, p=0.2930; (Middle-left panel) Calretinin, n=9M,9C, p=0.3138; (Middle-right panel) GABAARα, n=12M,9C, p=0.0173; (Right-most panel) vGlut2, n=7M,7C, p=0.6140. *p<0.05. Two-tailed unpaired Student’s t-test. Data is presented as mean +/- SEM. c. Gene ontology analysis of gene events (n=239 unique genes; listed in Supplementary Table 1) in iCK-cre mutants (n=3) compared to controls (n=3) by PANTHER, showing functional annotations by biologic process (left panel) and molecular function (right panel) using Fisher’s Exact Test with false discovery rate adjustment. d. Quantification of mean telomere length (left panel) and % of short telomeres (right panel) of cortical and dentate gyrus (dg) neurons in iCK-cre mutants (cortex: n=2770; dg: n=45996) vs. controls (cortex: n=3093; dg: n=37722). ****p<0.0001 using two tailed unpaired Students t-test and Chi-square test, respectively. Telomere length is presented as mean+/- SEM, while % short telomeres is presented as a ratio of number of short telomeres (below 25th percentile) over total number of telomeres. e. Telomere FISH images of iCK-cre mutants vs. controls in the cortex and dentate gyrus. All scale bars=100 μm. In a and e, experiments were independently repeated with similar results at least n=3 times using at least n=3 different mouse tissue samples for each group.
Supplementary Figure 2 Expression analysis of iND-cre Tomato reporter and mutant vs. control brains.
a. Tomato staining of iND-cre; R26-stop-tdTomato reporter and control at 4 weeks post-induction. b. Immunostaining of iND-cre reporter with lineage markers at 5 months post-induction c. Immunofluorescence staining of iND-cre mutant vs. control at 10 weeks post-induction. d. Quantification of % BrdU-positive cells in aged iND-cre mutants (n=3) vs. controls (n=3) in each of the indicated brain regions. Data is presented as mean +/- SEM. e. Immunofluorescence staining of aged iND-cre mutant vs. control at >6 months post-induction. All scale bars=100 μm. In a, b, c and e, experiments were independently repeated with similar results at least n=3 times using at least n=3 different mouse tissue samples for each group.
Supplementary Figure 3 Expression analysis of iDlx-cre Tomato reporter.
a. Tomato staining of tamoxifen- and vehicle-treated iDlx-cre;R26-stop-tdTomato reporter at 1 month post-induction. b. Immunostaining of iDlx-cre;R26-stop-tdTomato reporter with lineage markers at 5 months post-induction. All scale bars=100 μm. In a and b, experiments were independently repeated with similar results at least n=3 times using at least n=3 different mouse tissue samples for each group.
Supplementary Figure 4 Molecular and histologic analysis of iDlx-cre mutants.
a. Quantification of % BrdU-positive cells in aged iDlx-cre mutants (n=3) vs. controls (n=3) in each of the indicated brain regions. Data is presented as mean +/- SEM. b. Immunostaining of aged iDlx-cre mutants and controls with lineage markers. c. Western blot analysis of MAPK and PI3K pathway components in iDlx-cre mutant (M) and control (C) brains. Mouse GBM (mGBM) and HeLa cell lysates were used as positive controls. d. Quantification of mean telomere length (left panel) and % of short telomeres (right panel) of dentate gyrus neurons in iDlx-cre mutants (n=50144) vs. controls (n=65974). e. Quantification of mean telomere length (left panel) and % of short telomeres (right panel) of dentate gyrus neurons in iDlx-cre (n=33693) vs. iCK-cre (n=45988) mutants. In d and e, telomere length is presented as mean +/- SEM, while % short telomeres is presented as a ratio of the number of short telomeres (below 25th percentile) over total number of telomeres. ****p<0.0001 using two tailed unpaired Student’s t-test and Chi-square test, respectively. h. Telomere FISH images of iDlx-cre and iCK-cre mutants and controls in the dentate gyrus. All scale bars=100 μm. In b, c, and h, experiments were independently repeated with similar results at least n=3 times using at least n=3 different mouse tissue samples for each group.
Supplementary Figure 5 Molecular profiling of Syn-cre tumors.
RNA sequencing analysis of Syn1-cre, Nestin-creERT2, and NG2-creERTM tumors. a. Dimension reduction analysis of Syn1-cre (n=3), Nestin-creERT2 (n=3), and NG2-creERTM (n=3) mutant tumors. b. Heat map showing expression of NG2-creERTM tumor signature genes (also listed in Supplementary Table 2) in Syn1-cre (n=3), Nestin-creERT2 (n=3), and NG2-creERTM (n=3) tumors.
Supplementary information
Supplementary Figures 1–5
Supplementary Figures 1–5
Supplementary Table 1
Gene events in iCK-cre mutants.
Supplementary Table 2
NG2-creERTM tumor signature genes.
Supplementary Note 1
Antibodies used in the study and related information.
Rights and permissions
About this article
Cite this article
Alcantara Llaguno, S., Sun, D., Pedraza, A.M. et al. Cell-of-origin susceptibility to glioblastoma formation declines with neural lineage restriction. Nat Neurosci 22, 545–555 (2019). https://doi.org/10.1038/s41593-018-0333-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0333-8
This article is cited by
-
Hsa_circITGA4/ miR-1468/EGFR/ PTEN a Master Regulators Axis in Glioblastoma Development and Progression
Molecular Biotechnology (2024)
-
Modeling nervous system tumors with human stem cells and organoids
Cell Regeneration (2023)
-
Stalled oligodendrocyte differentiation in IDH-mutant gliomas
Genome Medicine (2023)
-
Lrig1 expression identifies quiescent stem cells in the ventricular-subventricular zone from postnatal development to adulthood and limits their persistent hyperproliferation
Neural Development (2023)
-
On the origin and development of glioblastoma: multifaceted role of perivascular mesenchymal stromal cells
Acta Neuropathologica Communications (2023)