Abstract
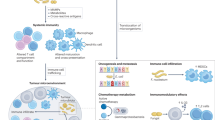
Fecal microbiota transplantation (FMT) represents a potential strategy to overcome resistance to immune checkpoint inhibitors in patients with refractory melanoma; however, the role of FMT in first-line treatment settings has not been evaluated. We conducted a multicenter phase I trial combining healthy donor FMT with the PD-1 inhibitors nivolumab or pembrolizumab in 20 previously untreated patients with advanced melanoma. The primary end point was safety. No grade 3 adverse events were reported from FMT alone. Five patients (25%) experienced grade 3 immune-related adverse events from combination therapy. Key secondary end points were objective response rate, changes in gut microbiome composition and systemic immune and metabolomics analyses. The objective response rate was 65% (13 of 20), including four (20%) complete responses. Longitudinal microbiome profiling revealed that all patients engrafted strains from their respective donors; however, the acquired similarity between donor and patient microbiomes only increased over time in responders. Responders experienced an enrichment of immunogenic and a loss of deleterious bacteria following FMT. Avatar mouse models confirmed the role of healthy donor feces in increasing anti-PD-1 efficacy. Our results show that FMT from healthy donors is safe in the first-line setting and warrants further investigation in combination with immune checkpoint inhibitors. ClinicalTrials.gov identifier NCT03772899.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All clinical metadata have been uploaded to the NCBI Sequence Read Archive, BioProject ID PRJNA928744. Patient baseline clinical data are available in Table 1 and within the text. Study-level clinical data from this study (including the protocol) will be made available upon reasonable request from a qualified medical or scientific professional for the specific purpose laid out in that request and may include de-identified individual participant data. The data for this request will be available after a data access agreement has been signed. Requests should be sent to the corresponding author. Patient-related data not included in the paper were generated as part of a clinical trial and are subject to patient confidentiality.
Change history
03 November 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41591-023-02650-8
References
Robert, C. et al. Five-year outcomes with nivolumab in patients with wild-type BRAF advanced melanoma. JCO 38, 3937–3946 (2020).
Larkin, J. et al. Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 381, 1535–1546 (2019).
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma (KEYNOTE-006): post-hoc 5-year results from an open-label, multicentre, randomised, controlled, phase 3 study. Lancet Oncol. 20, 1239–1251 (2019).
Esfahani, K. et al. Moving towards personalized treatments of immune-related adverse events. Nat. Rev. Clin. Oncol. 17, 504–515 (2020).
Derosa, L. et al. Microbiota-centered interventions: the next breakthrough in immuno-oncology? Cancer Discov. 11, 2396–2412 (2021).
Sepich-Poore, G. D. et al. The microbiome and human cancer. Science 371, eabc4552 (2021).
Routy, B. et al. The gut microbiota influences anticancer immunosurveillance and general health. Nat. Rev. Clin. Oncol. 15, 382–396 (2018).
Aghamajidi, A. & Maleki Vareki, S. The effect of the gut microbiota on systemic and anti-tumor immunity and response to systemic therapy against cancer. Cancers 14, 3563 (2022).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Andrews, L. P., Yano, H. & Vignali, D. A. A. Inhibitory receptors and ligands beyond PD-1, PD-L1 and CTLA-4: breakthroughs or backups. Nat. Immunol. 20, 1425–1434 (2019).
Lee, K. A. et al. Cross-cohort gut microbiome associations with immune checkpoint inhibitor response in advanced melanoma. Nat. Med. 28, 535–544 (2022).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
McCulloch, J. A. et al. Intestinal microbiota signatures of clinical response and immune-related adverse events in melanoma patients treated with anti-PD-1. Nat. Med. 28, 545–556 (2022).
Simpson, R. C. et al. Diet-driven microbial ecology underpins associations between cancer immunotherapy outcomes and the gut microbiome. Nat. Med. 38, 2344–2352 (2022).
Derosa, L. et al. Gut bacteria composition drives primary resistance to cancer immunotherapy in renal cell carcinoma patients. Eur. Urol. 78, 195–206 (2020).
Derosa, L. et al. Intestinal Akkermansia muciniphila predicts clinical response to PD-1 blockade in patients with advanced non-small-cell lung cancer. Nat. Med. 28, 315–324 (2022).
Messaoudene, M. et al. A natural polyphenol exerts antitumor activity and circumvents anti-PD-1 resistance through effects on the gut microbiota. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-21-0808 (2022).
Baruch, E. N. et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science 371, 602–609 (2021).
Davar, D. et al. Fecal microbiota transplant overcomes resistance to anti-PD-1 therapy in melanoma patients. Science 371, 595–602 (2021).
Craven, L. J., Nair Parvathy, S., Tat-Ko, J., Burton, J. P. & Silverman, M. S. Extended screening costs associated with selecting donors for fecal microbiota transplantation for treatment of metabolic syndrome-associated diseases. Open Forum Infect. Dis. 4, ofx243 (2017).
Parvathy, S. N. et al. Enhanced donor screening for faecal microbial transplantation during COVID-19. Gut 70, 2219–2220 (2021).
Ianiro, G. et al. Variability of strain engraftment and predictability of microbiome composition after fecal microbiota transplantation across different diseases. Nat. Med. 28, 1913–1923 (2022).
Blanco-Míguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01688-w (2023).
Moldoveanu, D. et al. Spatially mapping the immune landscape of melanoma using imaging mass cytometry. Sci. Immunol. 7, eabi5072 (2022).
Kamphorst, A. O. et al. Proliferation of PD-1+ CD8 T cells in peripheral blood after PD-1-targeted therapy in lung cancer patients. Proc. Natl Acad. Sci. USA 114, 4993–4998 (2017).
Kunert, A. et al. CD45RA+CCR7− CD8 T cells lacking co-stimulatory receptors demonstrate enhanced frequency in peripheral blood of NSCLC patients responding to nivolumab. J. Immunother. Cancer 7, 149 (2019).
Ninkov, M. et al. Improved MAIT cell functions following fecal microbiota transplantation for metastatic renal cell carcinoma. Cancer Immunol. Immunother. https://doi.org/10.1007/s00262-022-03329-8 (2022).
Yonekura, S. et al. Cancer induces a stress ileopathy depending on β-adrenergic receptors and promoting dysbiosis that contributes to carcinogenesis. Cancer Discov. 12, 1128–1151 (2022).
Kao, D. et al. Effect of oral capsule- vs colonoscopy-delivered fecal microbiota transplantation on recurrent clostridium difficile infection: a randomized clinical trial. JAMA 318, 1985–1993 (2017).
Saha, S., Mara, K., Pardi, D. S. & Khanna, S. Long-term safety of fecal microbiota transplantation for recurrent clostridioides difficile infection. Gastroenterology 160, 1961–1969 (2021).
Robert, C. et al. Nivolumab in previously untreated melanoma without BRAF mutation. N. Engl. J. Med. 372, 320–330 (2015).
Ribas, A. et al. Association of pembrolizumab with tumor response and survival among patients with advanced melanoma. JAMA 315, 1600–1609 (2016).
Wolchok, J. D. et al. Long-term outcomes with nivolumab plus ipilimumab or nivolumab alone versus ipilimumab in patients with advanced melanoma. JCO 40, 127–137 (2022).
Kuzmanovszki, D. et al. Anti-PD-1 monotherapy in advanced melanoma-real-world data from a 77-month-long retrospective observational study. Biomedicines 10, 1737 (2022).
Ibrahim, T., Mateus, C., Baz, M. & Robert, C. Older melanoma patients aged 75 and above retain responsiveness to anti-PD1 therapy: results of a retrospective single-institution cohort study. Cancer Immunol. Immunother. 67, 1571–1578 (2018).
Oliva, I. G. et al. 607 MCGRAW trial: evaluation of the safety and efficacy of an oral microbiome intervention (SER-401) in combination with nivolumab in first line metastatic melanoma patients. In Regular and Young Investigator Award Abstracts A637–A637 (BMJ Publishing Group, 2022).
Spencer, C. N. et al. Dietary fiber and probiotics influence the gut microbiome and melanoma immunotherapy response. Science 374, 1632–1640 (2021).
Al-Habsi, M. et al. Spermidine activates mitochondrial trifunctional protein and improves antitumor immunity in mice. Science 378, eabj3510 (2022).
Vorwald, V. M. et al. Circulating CD8+ mucosal-associated invariant T cells correlate with improved treatment responses and overall survival in anti-PD-1-treated melanoma patients. Clin. Transl. Immunol. 11, e1367 (2022).
Fan, X., Quezada, S. A., Sepulveda, M. A., Sharma, P. & Allison, J. P. Engagement of the ICOS pathway markedly enhances efficacy of CTLA-4 blockade in cancer immunotherapy. J. Exp. Med. 211, 715–725 (2014).
Xiao, Z., Mayer, A. T., Nobashi, T. W. & Gambhir, S. S. ICOS is an indicator of T-cell-mediated response to cancer immunotherapy. Cancer Res. 80, 3023–3032 (2020).
Filipazzi, P., Huber, V. & Rivoltini, L. Phenotype, function and clinical implications of myeloid-derived suppressor cells in cancer patients. Cancer Immunol. Immunother. 61, 255–263 (2012).
Azuma, K. et al. Clinical significance of plasma-free amino acids and tryptophan metabolites in patients with non-small cell lung cancer receiving PD-1 inhibitor: a pilot cohort study for developing a prognostic multivariate model. J. Immunother. Cancer 10, e004420 (2022).
Mullish, B. H. et al. Microbial bile salt hydrolases mediate the efficacy of faecal microbiota transplant in the treatment of recurrent Clostridioides difficile infection. Gut 68, 1791–1800 (2019).
Walter, J., Armet, A. M., Finlay, B. B. & Shanahan, F. Establishing or exaggerating causality for the gut microbiome: lessons from human microbiota-associated rodents. Cell 180, 221–232 (2020).
Freites-Martinez, A., Santana, N., Arias-Santiago, S. & Viera, A. CTCAE versión 5.0. Evaluación de la gravedad de los eventos adversos dermatológicos de las terapias antineoplásicas. Actas Dermosifiliogr. 112, 90–92 (2021).
Al, K. F., Bisanz, J. E., Gloor, G. B., Reid, G. & Burton, J. P. Evaluation of sampling and storage procedures on preserving the community structure of stool microbiota: a simple at-home toilet-paper collection method. J. Microbiol. Methods 144, 117–121 (2018).
Al, K. F. et al. Fecal microbiota transplantation is safe and tolerable in patients with multiple sclerosis: a pilot randomized controlled trial. Mult. Scler. J. Exp. Transl. Clin. https://doi.org/10.1177/20552173221086662 (2022).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Ndiaye, M. & Mattei, X. Endosymbiotic relationship between a rickettsia-like microorganism and the male germ-cells of Culex tigripes. J. Submicrosc. Cytol. Pathol. 25, 71–77 (1993).
Egermark-Eriksson, I., Carlsson, G. E. & Ingervall, B. Prevalence of mandibular dysfunction and orofacial parafunction in 7-, 11- and 15-year-old Swedish children. Eur. J. Orthod. 3, 163–172 (1981).
Damond, N. et al. A Map of Human Type 1 diabetes progression by imaging mass cytometry. Cell Metab. 29, 755–768 (2019).
Levine, J. H. et al. Data-driven phenotypic dissection of aml reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Kjer-Nielsen, L. et al. MR1 presents microbial vitamin B metabolites to MAIT cells. Nature 491, 717–723 (2012).
Corbett, A. J. et al. T-cell activation by transitory neo-antigens derived from distinct microbial pathways. Nature 509, 361–365 (2014).
Dona, A. C. et al. Precision high-throughput proton NMR spectroscopy of human urine, serum, and plasma for large-scale metabolic phenotyping. Anal. Chem. 86, 9887–9894 (2014).
Sands, C. J. et al. The nPYc-Toolbox, a Python module for the pre-processing, quality-control and analysis of metabolic profiling datasets. Bioinformatics 35, 5359–5360 (2019).
Takis, P. G. et al. A computationally lightweight algorithm for deriving reliable metabolite panel measurements from 1D 1H NMR. Anal. Chem. 93, 4995–5000 (2021).
Akoka, S., Barantin, L. & Trierweiler, M. Concentration measurement by proton NMR using the ERETIC method. Anal. Chem. 71, 2554–2557 (1999).
Sarafian, M. H. et al. Bile acid profiling and quantification in biofluids using ultra-performance liquid chromatography tandem mass spectrometry. Anal. Chem. 87, 9662–9670 (2015).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Wolfer, A. M. et al. peakPantheR, an R package for large-scale targeted extraction and integration of annotated metabolic features in LC–MS profiling datasets. Bioinformatics 37, 4886–4888 (2021).
Tautenhahn, R., Böttcher, C. & Neumann, S. Highly sensitive feature detection for high resolution LC/MS. BMC Bioinform. 9, 504 (2008).
Whiley, L. et al. Ultrahigh-performance liquid chromatography tandem mass spectrometry with electrospray ionization quantification of tryptophan metabolites and markers of gut health in serum and plasma—application to clinical and epidemiology cohorts. Anal. Chem. 91, 5207–5216 (2019).
Barr, D. J., Levy, R., Scheepers, C. & Tily, H. J. Random effects structure for confirmatory hypothesis testing: keep it maximal. J. Mem. Lang. 68, 255–278 (2013).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Acknowledgements
We thank the patients and their families. We thank the Clinical Research Unit (CRU) at the London Regional Cancer Program (LRCP) for their support in running the trial. We thank the staff at the CRCHUM animal facility and the Immunomonitoring core facility at the CRCHUM for their help with the experiments. We acknowledge the support of the Rosalind and Morris Goodman Cancer Institute Research Support, the Single Cell Imaging and Mass Cytometry Analysis Platform and the Histology Core facilities at McGill University. The clinical trial was funded by a grant from the Lotte and John Hecht Memorial Foundation awarded to S.M.V. and J.P.B., a grant from the Division of Medical Oncology at Western University awarded to J.G.L. and S.M.V. and a Canadian Cancer Society Impact grant supported by the Lotte and John Hecht Memorial Foundation awarded to B.R., S.M.V. and A.E. The laboratory of B.R. for ancillary analyses and biobanking was funded by Institute du Cancer de Montréal, Terry Fox Marathon of Hope clinician-scientist award. The laboratory of S.M.V. for ancillary analyses and biobanking was funded by a project grant from the Canadian Institute of Health Research (CIHR) (MOP no. 389137) and a LRCP Catalyst Grant Program, Keith Samitt Translational Research grant. Metagenomics sequencing was funded by ONCOBIOME, project number 825410 (Gut OncoMicrobiome Signatures associated with cancer incidence, prognosis and prediction of treatment response). B.R. received salary support from Fonds de Recherche du Quebec Santé. S.M.V. received salary support from the Ontario Institute of Cancer Research (OICR) and The London Health Sciences Foundation Helen and Andy Spriet funds. M.M. reports salary support from Seerave foundation. A.E. reports support from CIHR, Royal College of Physicians and Surgeons of Canada. Metabolomics studies were performed at the Medical Research Council (MRC)-National Institute of Health Research (NIHR) National Phenome Centre at Imperial College London; this center receives financial support from the MRC and NIHR (grant number MC_PC_12025). B.H.M. is the recipient of an NIHR Academic Clinical Lectureship (CL-2019-21-002). The Division of Digestive Diseases and MRC-NIHR National Phenome Centre at Imperial College London receive financial and infrastructure support from the NIHR Imperial Biomedical Research Centre based at Imperial College Healthcare NHS Trust and Imperial College London. B.D. acknowledges support from an NSERC Postdoctoral Fellowship Award (PDF-558010-2021). K.F.A. acknowledges support from an American Urological Association Research Scholar Award. Cytofluorimetric analyses in the laboratory of S.M.M.H. were funded through a project grant (PJT – 156295) from the CIHR. I.R.W. reports support from the Terry Fox Research Institute (grant 1084), CIHR (PJT-178341 grant), Canada Research Chairs Program, donations from K. J. Baggs and donations from the Rachel and Jason Schwartz Family Foundation. Financial support was also obtained from the Quebec Cancer Consortium and the Ministère de l’Économie et de l’Innovation du Québec through the Fonds d’accélération des collaborations en santé and the Victor Liu McGill Interdisciplinary Initiative in Infection and Immunity (MI4) initiative. This research was enabled, in part, by support provided by Calcul Québec (www.calculquebec.ca), SHARCNET (www.sharcnet.ca) and Compute Canada (www.computecanada.ca). L.D. was supported by RHU5 ANR-21-RHUS-0017 IMMUNOLIFE; EU-H2020, project no. 825410, project ONCOBIOME, Gut OncoMicrobiome Signatures associated with cancer incidence, prognosis and prediction of treatment response; and the SIGN’IT ARC Foundation.
Author information
Authors and Affiliations
Contributions
S.M.V. conceived the study. S.M.V. and J.G.L. designed the trial and co-wrote the trial protocol. S.M.V. and B.R. designed and supervised translational studies. J.G.L., W.H.M., R.J., S.E., D.L., K.B. and K.E. recruited and/or treated patients. M.M., B.A.D., C.H., K.F.A., L.M.G., M.P., C.R., M.N., G.P., F.A., F.P., M.K., R.F., P. Thebault, P. Takis, J.M., L.R., L.D., J.R.M., A.E., I.R.W., R.L., N.S., S.M.M.H., B.H.M. and J.P.B. collected and/or analyzed the data. S.N.P. and M.S.S. prepared and conducted FMT. B.R., J.G.L. and S.M.V. co-wrote the first draft. All authors provided comments and approved the paper.
Corresponding author
Ethics declarations
Competing interests
B.R. reports grants from Kaleido, Vedanta, Surface and Da Volterra outside the submitted work, as well as consulting fees from BMS, AstraZeneca, Merck and Da Volterra. He is also the co-founder of Science Curebiota. B.H.M. has received consultancy fees from Finch Therapeutics Group and Ferring Pharmaceuticals, outside of the submitted work. R.J. reports grants from BMS, Merck and Iovance Biotherapeutics as well as consulting fees from BMS and Merck. M.K. is an employee at Pfizer. L.D. had consulting and advisory roles for BMS, Sanofi and EverImmune and was supported by the Philantropia Fondation Gustave Roussy. J.R.M. has received consultancies from Enterobiotix and Cultech outside of the submitted work. S.M.V. is a director of the board of IMV. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Christian Jobin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Saheli Sadanand, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
a. Flow diagram indicating the number of patients screened, enrolled in the study, and who received the combination of FMT and anti-PD-1. One patient completed 24 months of pembrolizumab. Seven patients discontinued anti-PD-1 therapy due to progression and five patients discontinued treatment due to grade 3 immune-related adverse events. One patient discontinued anti-PD-1due to undiagnosed cardiac amyloidosis. Six patients remained on anti-PD-1 therapy at data cut off. All 20 patients were evaluable for safety and clinical outcome. b. Baseline characteristics information of the three healthy donors. c. Kaplan Meier curve representation of the progression-free survival, and d. overall survival at data cut-off with a median follow-up of 20.7 months.
Extended Data Fig. 2
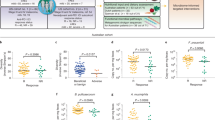
a. Beta-diversity ordination plot of 16S rRNA gene sequencing data showing baseline Aitchison distances between the 19 patients and healthy donors at S1. b. All metagenomics results were obtained from 61 samples from baseline n = 12 R patients and n = 6 NR patients. Longitudinal samples available for n = 10 R and n = 6 NR. Metagenomic sequencing with representation of the alpha-diversity measured by observed species over time from S1 to S4 including n = 18 patients.* p < 0.05.
Extended Data Fig. 3
a. Jaccard dissimilarity index between matching donor and patient measured with metagenomics data separated by outcomes 6 NR vs 10R from S1 to S4. b. 16S rRNA results from 19 patients, Aitchison distance between the patients and their marching donor over time. c. Variability of strain engraftment rate from the donor over time and d. Bray-Curtis dissimilarity rate representation for patients based on alpha-diversity n = 16 patients. e, f. Similarly, strain engraftment rate and Bray-Curtis dissimilarity rates when segregating for body mass index (BMI) below or above the median, respectively n = 16 patients. g. Spearman correlation between baseline alpha-diversity (Shannon index) and BMI. * p < 0.05; ** p < 0.01; *** p < 0.001. S1: Baseline, S2: one week S3: one month, S4 three months after FMT; R: responder, NR: nonresponder; BMI: body mass index.
Extended Data Fig. 4
a. Bray-Curtis comparison and b. strain engraftment rate from metagenomic sequences obtained from the MIMic trial (16 patients; 3 donors), Baruch et al.19 (10 patients; 2 donors) and Davar et al.,20 (15 patients; 7 donors) FMT trials. c. Spearman representation between baseline alpha-diversity (Shannon index) and strain engraftment rate for the 3 FMT trials. d. Correlation between baseline alpha-diversity (Shannon index) for each FMT trials and outcome. e. Pooled baseline alpha-diversity (Shannon index) between R and NR combining the n = 43 patients from the 3 FMT trials. * p < 0.05; ** p < 0.01; *** p < 0.001. S1: Baseline, S2: one week S3: one month, S4 three months after FMT; R: responder, NR: nonresponder.
Extended Data Fig. 5
Metagenomic fecal sample analysis a. LefSe representation of bacterial differential abundance between R and NR at S1 n = 18 patients. b. Heatmap of significant bacteria evolution between S1 vs S3 in NR patients (n = 6). Only bacteria with a p < 0.05 difference between both time points are represented. c. Heatmap of differentially abundant taxa (we.ep < 0.05, ALDEx2) in the 16S rRNA gene dataset for R (n = 13), and d. Taxa evolution between S1 and S3 n = 6 NR patients measured by 16S rRNA gene sequencing. e. Metagenomics strain that preferentially engrafted from donors in R and NR at S2 and S3 n = 16 patients. S1: Baseline, S2: one week S3: one month, S4 three months after FMT; R: responder, NR: nonresponder.
Extended Data Fig. 6
a. Volcano plot from METACyc pathways representation between S3 and S1 in NR (left panel) and R (right panel). b. Regression coefficient of serum bile acids measured by UHPLC–MS between S2 and S1 on 19 patients. c. Serum succinate and lactic acid levels measured by 1H-NMR on 13R and 6 NR over time. Thick lines represent mean-group average levels while thin lines represent individual trajectories. * p < 0.05; ** p < 0.01.
Extended Data Fig. 7
a. Representative Delaunay graphs of 3 CyTOF images showing NR with low immune infiltration (patient 623-1), R with high immune infiltration (patient 803-1), and R with low immune infiltration (patient 623-4). b. Stacked bar plots showing the proportion of each cell type in each patient. Melanoma and other cell types are excluded for visualization purposes. c. Boxplot comparing immune cell proportions in 3 NR and 9 R. P-values calculated from two-sided Wilcoxon rank-sum test. d Boxplot comparing contact enrichment scores between melanoma and immune cells. Higher scores indicate cell contacts occur more frequently than expected by chance. P values calculated from two-sided Wilcoxon rank-sum test. Antigen-Experienced cytotoxic T cells (Tc.ae), Naive cytotoxic T cells (Tc.naive), Antigen-experienced T helper cells (Th.ae), Naive T helper cells (Th.naive).
Extended Data Fig. 8
a. Flow cytometry representation of unsupervised t-SNE panels of circulating immune cells present from n = 13 R and n = 6 NR over time from S1–S4. Each population is color-coded. b. Supervised t-SNE panel on CD8+ T cells representation from S1–S4 between R and NR. ICOS+ populations are represented in red. c. Flow cytometric analysis of peripheral blood MAIT cell frequency over time between R and NR. d. Frequency of PD-1+ MAIT cells at S3 between 11 R and 5 NR. MAIT cells were identified as CD3+MR1-5-OP-RU-loaded tetramer positive population. MR1-6-FP-loaded tetramers were used to set control gates. * p < 0.05.
Extended Data Fig. 9
a. Pooled analysis of tumor growth size (left panel) and tumor size at sacrifice (right panel) of antibiotic-treated mice bearing MCA-205 treated with sequential anti-PD-1 or control IsoPD-1 after receiving FMT from one donor. Experiment was performed for the 3 donors. Each circle represents one animal, n = 15. b. Similar experimental setting was performed in B16-OVA cells in antibiotic-treated mice with FMT from 1 NR S1 and 1 NR S3. Each square represents one animal at the time of sacrifice n = 5. c. Tumor size at sacrifice in MCA-205 model from Fig. 4a,b, where one circle represents the mean value of one FMT performed on 5 mice at the time of sacrifice. Left panel FMT from R S1 or NR S1 + NR S3 samples and right panel from R S3 The color code represents one patient, orange: 802-1, green: 802-2 and purple: 802-4 and pink: 802-05. d. Tumor size at sacrifice in B16 model from Fig. 4c, where one square represents the mean value of one FMT performed on 5 mice at the time of sacrifice. Left panel FMT from R S1 or NR S1 + NR S3 samples and right panel from R S3. The color code represents one patient, orange: 802-1, green: 802-2 and purple: 802-4 and pink: 802-05 e. Flow cytometry analysis of the frequency of TIM3+CD8+ T cells in B16-OVA tumor infiltrating cells from mice that received FMT from 1R patients at S1 or S3 (Fig. 4c). f. 16S rRNA gene analysis of fecal samples from pooled mice associated with anti-PD-1 resistance following FMT from R S1 + NR S1 + NR S3 (anti-PD-1-resistant) n = 74 compared to anti-PD-1 sensitive mice following FMT from R S3 n = 45. Representation of alpha-diversity between both groups and, g. Beta-diversity between both groups. All groups reared in separated cages then received isotype control (IsoPD-1) or anti-PD-1 treatment. Means ± SEM are represented. * p < 0.05; ** p < 0.01; *** p < 0.001. S1: Baseline, S3: one month after FMT; R: responder, NR: nonresponder.
Extended Data Fig. 10
a. Representation of tumor size at sacrifice of germ-free mice recolonized with 2 donors in both isotype control (IsoPD-1) or anti-PD-1. b. Kinetic representation of three different conditions in antibiotic-treated mice. Mice were recolonized at D-15 with R S1 (blue), D2 marching donor (green) and, R S1 followed by two FMT from the matching donor (D2) at day +3 and day +7 (green). Means ± SEM are represented. * p < 0.05; ** p < 0.01. Each circle represents one mouse. S1: Baseline, S3: one month after FMT; R: responder, NR: non-responder; D: donor.
Supplementary information
Supplementary Information
Supplementary Figs.1–3 and Supplementary Tables 1–4.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Routy, B., Lenehan, J.G., Miller, W.H. et al. Fecal microbiota transplantation plus anti-PD-1 immunotherapy in advanced melanoma: a phase I trial. Nat Med 29, 2121–2132 (2023). https://doi.org/10.1038/s41591-023-02453-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-023-02453-x
This article is cited by
-
Longitudinal analysis shows how microbiome changes might influence responses to immune checkpoint blockade
Nature Medicine (2024)
-
Identification of dynamic microbiota signatures in patients with melanoma receiving ICIs: opportunities and challenges
Nature Reviews Clinical Oncology (2024)
-
Mapping the single cell spatial immune landscapes of the melanoma microenvironment
Clinical & Experimental Metastasis (2024)
-
The Microbiome Matters: Its Impact on Cancer Development and Therapeutic Responses
Journal of Microbiology (2024)
-
Longitudinal gut microbiome changes in immune checkpoint blockade-treated advanced melanoma
Nature Medicine (2024)