Abstract
Immune-modulating therapies have revolutionized the treatment of chronic diseases, particularly cancer. However, their success is restricted and there is a need to identify new therapeutic targets. Here, we show that natural killer cell granule protein 7 (NKG7) is a regulator of lymphocyte granule exocytosis and downstream inflammation in a broad range of diseases. NKG7 expressed by CD4+ and CD8+ T cells played key roles in promoting inflammation during visceral leishmaniasis and malaria—two important parasitic diseases. Additionally, NKG7 expressed by natural killer cells was critical for controlling cancer initiation, growth and metastasis. NKG7 function in natural killer and CD8+ T cells was linked with their ability to regulate the translocation of CD107a to the cell surface and kill cellular targets, while NKG7 also had a major impact on CD4+ T cell activation following infection. Thus, we report a novel therapeutic target expressed on a range of immune cells with functions in different immune responses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The materials, data and any associated protocols that support the findings of this study are available from the corresponding author upon request. The RNA-seq and microarray data have been deposited in the NCBI Gene Expression Omnibus database (https://www.ncbi.nlm.nih.gov/geo/) with the accession codes GSE135965 (human RNA-seq data) and GSE135857 (mouse microarray data). The list of DEGs in CD4+ T cells isolated from PBMCs of patients with visceral leishmaniasis is available in Supplementary Table 1. Supplementary Table 2 contains the list of DEGs in liver CD4+ T cells of L. donovani-infected mice. Supplementary Table 3 contains the list of DEGs in splenic CD4+ T cells of L. donovani-infected mice.
Change history
15 February 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41590-024-01770-8
References
Netea, M. G. et al. A guiding map for inflammation. Nat. Immunol. 18, 826–831 (2017).
Trapani, J. A. & Smyth, M. J. Functional significance of the perforin/granzyme cell death pathway. Nat. Rev. Immunol. 2, 735–747 (2002).
Crawford, A. et al. Molecular and transcriptional basis of CD4+ T cell dysfunction during chronic infection. Immunity 40, 289–302 (2014).
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461 (2015).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Schett, G. & Neurath, M. F. Resolution of chronic inflammatory disease: universal and tissue-specific concepts. Nat. Commun. 9, 3261 (2018).
Tubo, N. J. & Jenkins, M. K. CD4+ T cells: guardians of the phagosome. Clin. Microbiol. Rev. 27, 200–213 (2014).
Engwerda, C. R., Ato, M. & Kaye, P. M. Macrophages, pathology and parasite persistence in experimental visceral leishmaniasis. Trends Parasitol. 20, 524–530 (2004).
Miller, L. H., Baruch, D. I., Marsh, K. & Doumbo, O. K. The pathogenic basis of malaria. Nature 415, 673–679 (2002).
Montes de Oca, M. et al. Blimp-1-dependent IL-10 production by Tr1 cells regulates TNF-mediated tissue pathology. PLoS Pathog. 12, e1005398 (2016).
Bald, T. et al. Immune cell–poor melanomas benefit from PD-1 blockade after targeted type I IFN activation. Cancer Discov. 4, 674–687 (2014).
O'Donnell, J. S., Teng, M. W. L. & Smyth, M. J. Cancer immunoediting and resistance to T cell-based immunotherapy. Nat. Rev. Clin. Oncol. 16, 151–167 (2019).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Fairfax, B. P. et al. Peripheral CD8+ T cell characteristics associated with durable responses to immune checkpoint blockade in patients with metastatic melanoma. Nat. Med. 26, 193–199 (2020).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Turman, M. A., Yabe, T., McSherry, C., Bach, F. H. & Houchins, J. P. Characterization of a novel gene (NKG7) on human chromosome 19 that is expressed in natural killer cells and T cells. Hum. Immunol. 36, 34–40 (1993).
Aschenbrenner, D. et al. An immunoregulatory and tissue-residency program modulated by c-MAF in human TH17 cells. Nat. Immunol. 19, 1126–1136 (2018).
Engwerda, C. R. & Kaye, P. M. Organ-specific immune responses associated with infectious disease. Immunol. Today 21, 73–78 (2000).
Engwerda, C. R., Ng, S. S. & Bunn, P. T. The regulation of CD4+ T cell responses during protozoan infections. Front. Immunol. 5, 498 (2014).
Jenner, R. G. et al. The transcription factors T-bet and GATA-3 control alternative pathways of T-cell differentiation through a shared set of target genes. Proc. Natl Acad. Sci. USA 106, 17876–17881 (2009).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Antignano, F. et al. Methyltransferase G9A regulates T cell differentiation during murine intestinal inflammation. J. Clin. Invest. 124, 1945–1955 (2014).
Stumhofer, J. S. et al. Interleukins 27 and 6 induce STAT3-mediated T cell production of interleukin 10. Nat. Immunol. 8, 1363–1371 (2007).
Stern, J. J., Oca, M. J., Rubin, B. Y., Anderson, S. L. & Murray, H. W. Role of L3T4+ and LyT-2+ cells in experimental visceral leishmaniasis. J. Immunol. 140, 3971–3977 (1988).
Dickinson, M. E. et al. High-throughput discovery of novel developmental phenotypes. Nature 537, 508–514 (2016).
Butler, N. S. et al. Therapeutic blockade of PD-L1 and LAG-3 rapidly clears established blood-stage Plasmodium infection. Nat. Immunol. 13, 188–195 (2011).
Mou, Z. et al. Identification of broadly conserved cross-species protective Leishmania antigen and its responding CD4+ T cells. Sci. Transl. Med. 7, 310ra167 (2015).
Engwerda, C., Belnoue, E., Gruner, A. C. & Renia, L. Experimental models of cerebral malaria. Curr. Top. Microbiol. Immunol. 297, 103–143 (2005).
Amante, F. H. et al. Immune-mediated mechanisms of parasite tissue sequestration during experimental cerebral malaria. J. Immunol. 185, 3632–3642 (2010).
Haque, A. et al. Granzyme B expression by CD8+ T cells is required for the development of experimental cerebral malaria. J. Immunol. 186, 6148–6156 (2011).
Valencia-Hernandez, A. M. et al. A natural peptide antigen within the Plasmodium ribosomal protein RPL6 confers liver TRM cell-mediated immunity against malaria in mice. Cell Host Microbe 27, 950–962.e7 (2020).
Lau, L. S. et al. CD8+ T cells from a novel T cell receptor transgenic mouse induce liver-stage immunity that can be boosted by blood-stage infection in rodent malaria. PLoS Pathog. 10, e1004135 (2014).
Betts, M. R. et al. Sensitive and viable identification of antigen-specific CD8+ T cells by a flow cytometric assay for degranulation. J. Immunol. Methods 281, 65–78 (2003).
Grulich, A. E., van Leeuwen, M. T., Falster, M. O. & Vajdic, C. M. Incidence of cancers in people with HIV/AIDS compared with immunosuppressed transplant recipients: a meta-analysis. Lancet 370, 59–67 (2007).
Cursons, J. et al. A gene signature predicting natural killer cell infiltration and improved survival in melanoma patients. Cancer Immunol. Res. 7, 1162–1174 (2019).
Smyth, M. J., Kelly, J. M., Baxter, A. G., Korner, H. & Sedgwick, J. D. An essential role for tumor necrosis factor in natural killer cell-mediated tumor rejection in the peritoneum. J. Exp. Med. 188, 1611–1619 (1998).
Smyth, M. J., Crowe, N. Y. & Godfrey, D. I. NK cells and NKT cells collaborate in host protection from methylcholanthrene-induced fibrosarcoma. Int. Immunol. 13, 459–463 (2001).
Hayakawa, Y. & Smyth, M. J. CD27 dissects mature NK cells into two subsets with distinct responsiveness and migratory capacity. J. Immunol. 176, 1517–1524 (2006).
Street, S. E., Cretney, E. & Smyth, M. J.Perforin and interferon-γ activities independently control tumor initiation, growth, and metastasis. Blood 97, 192–197 (2001).
Guillerey, C., Huntington, N. D. & Smyth, M. J. Targeting natural killer cells in cancer immunotherapy. Nat. Immunol. 17, 1025–1036 (2016).
Cheuk, S. et al. CD49a expression defines tissue-resident CD8+ T cells poised for cytotoxic function in human skin. Immunity 46, 287–300 (2017).
Medley, Q. G. et al. Characterization of GMP-17, a granule membrane protein that moves to the plasma membrane of natural killer cells following target cell recognition. Proc. Natl Acad. Sci. USA 93, 685–689 (1996).
Krzewski, K., Gil-Krzewska, A., Nguyen, V., Peruzzi, G. & Coligan, J. E. LAMP1/CD107a is required for efficient perforin delivery to lytic granules and NK-cell cytotoxicity. Blood 121, 4672–4683 (2013).
Moreira-Teixeira, L. et al. Mouse transcriptome reveals potential signatures of protection and pathogenesis in human tuberculosis. Nat. Immunol. 21, 464–476 (2020).
Karwacz, K. et al. Critical role of IRF1 and BATF in forming chromatin landscape during type 1 regulatory cell differentiation. Nat. Immunol. 18, 412–421 (2017).
Mombaerts, P. et al. RAG-1-deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Song, J. et al. A mouse model for the human pathogen Salmonella typhi. Cell Host Microbe 8, 369–376 (2010).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Hochheiser, K., Kueh, A. J., Gebhardt, T. & Herold, M. J. CRISPR/Cas9: a tool for immunological research. Eur. J. Immunol. 48, 576–583 (2018).
Bradley, D. J. & Kirkley, J. Regulation of Leishmania populations within the host. I. The variable course of Leishmania donovani infections in mice. Clin. Exp. Immunol. 30, 119–129 (1977).
Franke-Fayard, B. et al. Murine malaria parasite sequestration: CD36 is the major receptor, but cerebral pathology is unlinked to sequestration. Proc. Natl Acad. Sci. USA 102, 11468–11473 (2005).
Putz, E. M. et al. NK cell heparanase controls tumor invasion and immune surveillance. J. Clin. Invest. 127, 2777–2788 (2017).
Markey, K. A., Gartlan, K. H., Kuns, R. D., MacDonald, K. P. & Hill, G. R. Imaging the immunological synapse between dendritic cells and T cells. J. Immunol. Methods 423, 40–44 (2015).
Du, P., Kibbe, W. A. & Lin, S. M. Lumi: a pipeline for processing Illumina microarray. Bioinformatics 24, 1547–1548 (2008).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Schmittgen, T. D., Lee, E. J. & Jiang, J. High-throughput real-time PCR. Methods Mol. Biol. 429, 89–98 (2008).
Gabrysova, L. et al. c-Maf controls immune responses by regulating disease-specific gene networks and repressing IL-2 in CD4+ T cells. Nat. Immunol. 19, 497–507 (2018).
Chow, M. et al. NLRP3 suppresses NK cell–mediated responses to carcinogen-induced tumors and metastases. Cancer Res. 72, 5721–5732 (2012).
Hayakawa, Y. et al. Cutting edge: tumor rejection mediated by NKG2D receptor–ligand interaction is dependent upon perforin. J. Immunol. 169, 5377–5381 (2002).
Ferrari de Andrade, L. et al. Natural killer cells are essential for the ability of BRAF inhibitors to control BRAFV600E-mutant metastatic melanoma. Cancer Res. 74, 7298–7308 (2014).
Gao, Y. et al. Tumor immunoevasion by the conversion of effector NK cells into type 1 innate lymphoid cells. Nat. Immunol. 18, 1004–1015 (2017).
Young, A. et al. Co-inhibition of CD73 and A2AR adenosine signaling improves anti-tumor immune responses. Cancer Cell 30, 391–403 (2016).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Wagner, A. H. et al. DGIdb 2.0: mining clinically relevant drug–gene interactions. Nucleic Acids Res. 44, D1036–D1044 (2016).
Eppig, J. T., Motenko, H., Richardson, J. E., Richards-Smith, B. & Smith, C. L. The International Mouse Strain Resource (IMSR): cataloging worldwide mouse and ES cell line resources. Mamm. Genome 26, 448–455 (2015).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Colaprico, A. et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 44, e71 (2016).
Acknowledgements
We thank the staff at the Kala-Azar Medical Research Centre (KAMRC), Muzaffarpur, India for help with the collection of blood samples, as well as patients and volunteers for allowing the use of blood samples. We thank staff at the QIMR Berghofer flow cytometry laboratory for assistance, and staff at the QIMR Berghofer animal facility for animal husbandry. We acknowledge the facilities, and the scientific and technical assistance of the MAGEC, Walter and Eliza Hall Institute of Medical Research. The MAGEC is supported by the Australian Phenomics Network (APN) and the APN is supported by the Australian government through the National Collaborative Research Infrastructure Strategy program. We thank the NIH tetramer facility (Atlanta, GA, United States) for production of the I-Ab-PEPCK335–351 tetramer used to detect L. donovani PEPCK-specific CD4+ T cells in these studies. This work was made possible through Queensland State Government funding and grants and fellowships from the National Health and Medical Research Council of Australia (NHMRC; grant numbers 1037304, 1058685, 1078671, 1132519, 1132975 and 1154265). Funding was also provided through a National Institutes of Health Tropical Medicine Research Centre (TMRC) grant (U19 AI074321), as well as Australian post-graduate awards through Griffith University’s Institute of Glycomics and School of Natural Sciences to P.T.B. and S.S.N., respectively; a Dr. Mildred Scheel Stiftung für Krebsforschung scholarship from Deutsche Krebshilfe to M.B.; and an INSPIRE Faculty grant (LSBM-109/IF-14) provided by the Indian government Department of Science and Technology (DST), Banaras Hindu University and University Grants Commission (M-14-70) to R.K. B.S. was supported by a Junior Research Fellowship from the Indian Council of Medical Research.
Author information
Authors and Affiliations
Contributions
S.S.N. and C.R.E. conceived of and designed the study with input from M.J.S., S.S., R. Kumar, G.R.H., K.N., S.-K.T., T.B., M.D.-E., M.W.L.T., A.G.B., W.C.D. and A.H. and led and coordinated the study with M.J.S. and S.S. S.S.N. and C.R.E. co-wrote the manuscript together with M.J.S. and G.R.H. S.S.N. performed the bioinformatics analysis with D.C., S.W. and N.C. S.B.C., B.S., S.S.S., O.P.S. and R. Kumar collected and processed samples from patients with visceral leishmaniasis under the coordination of S.S. I.D., P.Z., and R. Kuns performed the early experiments in inflammatory models. F.D.L.R., T.C.M.F., E.M., J.N., J.A.E., M.S.F.S., M.M.d.O., P.T.B., Y.W., F.H.A. and C.L.E. performed all of the experimental malaria and visceral leishmaniasis experiments. J.Y., J.H., X.-Y.L., A.R.A., M.C., M.B., K.N. and M.J.S. performed all of the cancer model experiments. A.J.K. and M.J.H. generated the C57BL/6J-Nkg7em1(cre)WEHI mouse. A.L. and N.L.D. performed modeling of the NKG7 tertiary protein structure. B.A.M., S.-K.T. and T.C.M.F. performed all of the retrovirus transductions and confocal microscopy. J.U. developed the PEPCK tetramer and provided advice on its use. N.G. and W.R.H. produced the Plasmodium peptide–MHC I tetramer and helped design the PbT-I cell-killing assays. W.C.D., A.G.B. and M.D.-E. provided important discussions for the project and critical feedback on the manuscript. All co-authors read, reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Nkg7 is only expressed by NK cells and a subset of CD8+ TCRβ+ cells in the spleen in naive mice. t-SNE plot of splenocytes from a naive mouse, pre-gated to exclude doublets and dead cells.
The remaining cells were clustered using TCRβ-BUV737, CD4-BUV395, CD8α-PE/Cy7, CD11b-PerCP/Cy5.5, CD11c-BV785, MHC-II-Pacific Blue, B220-BV650, NK1.1-APC/Cy7 and Ly-6C-BV605. Equal numbers of cells (15,000 cells) are shown for Cre– and Cre+ plots. The black oval indicates the GFP+ population. n = 1 per genotype, performed once. BV, brilliant violet.
Extended Data Fig. 2 Changes in the frequencies of Nkg7-expressing NK cells and CD8+ T cells during Leishmania donovani infection.
a, The gating strategy used to assess changes in the key immune cell subsets including NK cells, CD4+ T cells, CD8+ T cells, B cells, cDCs, pDCs, CD11bhi Ly6Cint monocytes, inflammatory monocytes, macrophages, and NKT cells. b, The graphs show changes in GFP within each of the key immune cell subsets in the spleen and liver during L. donovani infection. Statistical significance was determined using the Kruskal–Wallis one-way analysis of variance (ANOVA) with Dunn’s multiple comparisons test. c, The frequencies of GFP+ TH1 (gated on NK1.1– TCRβ+ CD8– CD4+ IFN-γ+ IL-10–) and TR1 cells (gated on NK1.1– TCRβ+ CD8– CD4+ IFN-γ+ IL-10+) in the spleen and liver during the course of infection are shown. A two-way ANOVA with Sidak’s multiple comparisons test was performed to test for statistical significance. d, Changes in the frequency and total number of GFP+ cells in the spleen and liver over the course of infection are shown. p value is indicated where * p < 0.05. Error bars represent mean ± SEM. The data shown is representative of two independent experiments, each with n = 3 mice per genotype, per timepoint.
Extended Data Fig. 3 Nkg7 deficiency results in reduced CD4+ T cell responses during Leishmania donovani infection.
a, The liver and spleen weights of WT and Nkg7–/– mice during L. donovani infection. A two-way ANOVA with Sidak’s multiple comparisons test was used to determine statistical significance. Data is representative of two experiments, where n = 3 naive WT and Nkg7–/– mice, and n = 5 WT and 4 Nkg7–/– mice at days 14, 28 and 58 p.i. groups. b and c, The frequency and total number of conventional (Foxp3–) CD4+ T cells in the liver (b) and spleen (c) at day 14 p.i. are shown. Statistical significance was determined using a two-way ANOVA with Tukey’s multiple comparisons test. The data shown is representative of two independent experiments, each with n = 3 naive WT and Nkg7–/– mice, and n = 5 WT and 4 Nkg7–/– mice at day 14 p.i. d, The expression of Ifng and Tnf mRNA by spleen or liver CD4+ T cells in naive or infected (day 14 p.i.) mice was determined by RT-qPCR. A two-way ANOVA with Sidak’s multiple comparisons test was used to determine statistical significance. n = 4 naive and 5 infected mice in each group. e, The representative histograms show PD-1, CTLA-4, and ICOS staining on CD4+ T cells in the liver at day 14 p.i. The graphs indicate the frequencies of PD-1+, CTLA-4+, and ICOS+ CD4+ T cells. Statistical significance was determined using the Mann–Whitney test. Data is derived from one experiment, where n = 3 naive WT and Nkg7–/– mice, and n = 5 infected WT and 4 infected Nkg7–/– mice. f, The frequency and total number of I-AbPEPCK335-351 (tetramer)-PE+ cells in the liver of naive and infected (day 14 p.i.) mice are shown. A two-way ANOVA with multiple comparisons test was used to test for statistical significance. n = 4 WT or Nkg7–/– naïve, and 5 WT or 6 Nkg7–/– infected mice. Data is representative of two independent experiments. g, Representative plots depict the differences in TH1 (IFN-γ+ T-bet+ cells) frequencies in WT or Nkg7–/– I-AbPEPCK335-351 (tetramer)-PE+ cells. The frequencies and numbers are shown in the accompanying graphs below. Statistical significance was determined using the Mann–Whitney test. n = 5 mice per group. Data is representative of two independent experiments. h, The frequency of WT or Nkg7–/– CD4+ TCRβ+ cells and I-AbPEPCK335-351 (tetramer)-PE+ cells expressing IL-6-stimulated phosphorylated (p)-STAT3 are shown. Statistical significance was determined using the Mann–Whitney test. n = 5 mice per group. Data is representative of two independent experiments. p value is indicated where * p < 0.05. Error bars represent mean ± SEM.
Extended Data Fig. 4 The absence of NKG7 results in decreased CD8+ T cell cytotoxicity during PbA infection.
a, The graphs show the frequency and total number of NK cells, CD4+ T cells and CD8+ T cells in the brain of naive and infected (peak of ECM) WT (n = 3 naive and 5 infected) and Nkg7–/– (n = 3 naive and 5 infected) mice. Cell subsets frequencies are expressed as a percentage of CD45+ cells. Data is representative of two independent experiments. b, Representative flow cytometry plots were gated on lymphocytes, singlets, live cells, NK1.1-APC/Cy7– TCRβ-BUV737+, CD8α-PerCP/Cy5.5+ cells and the frequencies of CD11a+ CD49d+ cells and Granzyme B+ cells are shown. n = 3 naïve and 4 infected mice per strain. Data is representative of two independent experiments. c, The graph shows the proportions of PbT-IWT and PbT-IΔNkg7 cells in the spleen of naive (n = 4) or infected mice at peak of ECM (n = 5). d, The frequencies of splenic PbT-IWT and PbT-IΔNkg7 transgenic CD8+ T cells expressing Granzyme B or Perforin, in the presence of monensin, are shown. n = 4 naïve and 5 infected mice at peak of ECM. e, The representative histograms show differences in the expression of CD107a by splenic PbT-IWT and PbT-IΔNkg7 cells incubated with monensin or stimulated with PMA and ionomycin in the presence of monensin. The frequency and MFI of CD107a expression is shown in the accompanying graphs. n = 4 naïve and 5 infected mice at peak of ECM. Statistical significance in all graphs was determined using a two-way ANOVA with Tukey’s (a) or Sidak’s (b–e) multiple comparisons test.
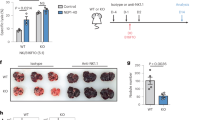
Extended Data Fig. 5 Nkg7-deficiency results in increased cancer metastasis in experimental models.
a, A correlation (r) between the moving average of a 19-gene natural killer (NK) cell signature genes and NKG7 expression in n = 472 samples from TCGA:SKCM dataset. b, The graphs indicate differences in the number of lung metastases between WT and Nkg7–/– mice following injection of RM-1 prostate carcinoma cells (n = 6 WT and 7 Nkg7–/–). The survival of WT and Nkg7–/– mice treated with either cIg (n = 7 WT and 13 Nkg7–/–) or α-asGM1 (n = 6 WT and 12 Nkg7–/–) in an intraperitoneal RMA-s lymphoma model was also assessed. Statistical significance between groups was tested using the Log-rank (Mantel–Cox) test. Additionally, the difference in percentage of tumour-free mice between WT (n = 17) and Nkg7–/– (n = 14) mice following MCA-induced fibrosarcoma generation is shown. The log-rank (Mantel–Cox) test was used to determine statistical significance. ** and *** represents p < 0.01 and 0.001 respectively. ns, not significant. c, The lungs of WT and Nkg7–/– mice, injected with LWT1 cells, were assessed for differences in the frequency and total cell number of hematopoietic cells, at 14 days post-injection. d, The frequency and total cell number of NK cells and T cells were quantified in the lungs of mice injected with LWT1 cells, at 14 days post-injection. e, The differences in the frequency of NK cells at different stages of maturation, based on CD27 and CD11b expression at 14 days post-injection of LWT1 cells is shown. The data shown in c–e is representative of two independent experiments where n = 5 mice per group. The Mann–Whitney test was used to determine statistical significance. p values are shown as follows: * p < 0.05 and ** p < 0.01.
Extended Data Fig. 6 Nkg7–/– NK cells do not have reduced abilities to conjugate with target cells or to form synapses.
a and b, The representative histograms show expression of DNAM-1 (CD226), NKG2D (CD314), CD11a, Granzyme B, and Perforin by WT (n = 6) or Nkg7–/– (n = 5) NK cells in the naive state (a) or when activated with rIL-2 (b). The MFI for the expression of each marker in naive NK cells is also shown. Data is pooled from 2 independent experiments. Sv, Streptavidin. c, Representative plots depict the frequency of cell conjugates formed when CellTrace Violet (CTV)-labelled WT or Nkg7–/– NK cells were co-cultured with carboxyfluorescein succinimidyl ester (CFSE)-labelled YAC-1 target cells for 30 minutes at an E:T ratio of 1:2. The frequency of conjugated NK cells at 5, 15, 30, and 60 minutes is shown in the accompanying graph. n = 3 mice per group. Data is pooled from 2 independent experiments. d, Representative images of effector NK cell–YAC-1 target cell conjugates visualised using an Amnis® ImageStream®XMark II after in vitro co-culture of WT or Nkg7–/– cells with target cells for 15 minutes. The graphs show the MFI of Phalloidin or LFA-1 at the interface between effector and target cells. Data obtained from one experiment.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Supplementary Tables 1 and 5–7.
Supplementary Tables
Supplementary Tables 2–5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ng, S.S., De Labastida Rivera, F., Yan, J. et al. The NK cell granule protein NKG7 regulates cytotoxic granule exocytosis and inflammation. Nat Immunol 21, 1205–1218 (2020). https://doi.org/10.1038/s41590-020-0758-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0758-6
This article is cited by
-
CTLs heterogeneity and plasticity: implications for cancer immunotherapy
Molecular Cancer (2024)
-
T cell expressions of aberrant gene signatures and Co-inhibitory receptors (Co-IRs) as predictors of renal damage and lupus disease activity
Journal of Biomedical Science (2024)
-
CD4+ T cell immunity is dependent on an intrinsic stem-like program
Nature Immunology (2024)
-
Effects of methionine deficiency on B7H3-DAP12-CAR-T cells in the treatment of lung squamous cell carcinoma
Cell Death & Disease (2024)
-
Single-cell transcriptomic profiling reveals immune cell heterogeneity in acute myeloid leukaemia peripheral blood mononuclear cells after chemotherapy
Cellular Oncology (2024)