Abstract
Regulatory myeloid immune cells, such as myeloid-derived suppressor cells (MDSCs), populate inflamed or cancerous tissue and block immune cell effector functions. The lack of mechanistic insight into MDSC suppressive activity and a marker for their identification has hampered attempts to overcome T cell inhibition and unleash anti-cancer immunity. Here, we report that human MDSCs were characterized by strongly reduced metabolism and conferred this compromised metabolic state to CD8+ T cells, thereby paralyzing their effector functions. We identified accumulation of the dicarbonyl radical methylglyoxal, generated by semicarbazide-sensitive amine oxidase, to cause the metabolic phenotype of MDSCs and MDSC-mediated paralysis of CD8+ T cells. In a murine cancer model, neutralization of dicarbonyl activity overcame MDSC-mediated T cell suppression and, together with checkpoint inhibition, improved the efficacy of cancer immune therapy. Our results identify the dicarbonyl methylglyoxal as a marker metabolite for MDSCs that mediates T cell paralysis and can serve as a target to improve cancer immune therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The microarray data generated from human MDSCs compared with monocytes have been deposited in the Gene Expression Omnibus with accession code GSE131679. Source data for Figs. 1, 2 and 4–7 and Extended Data Figs. 2–5 and 8–10 are presented with the paper. The data that support the findings of this study are available from the corresponding authors upon request.
References
Spitzer, M. H. et al. Systemic immunity is required for effective cancer immunotherapy. Cell 168, 487–502.e15 (2017).
Williams, M. A. & Bevan, M. J. Effector and memory CTL differentiation. Annu. Rev. Immunol. 25, 171–192 (2007).
Togashi, Y., Shitara, K. & Nishikawa, H. Regulatory T cells in cancer immunosuppression—implications for anticancer therapy. Nat. Rev. Clin. Oncol. 16, 356–371 (2019).
Veglia, F., Perego, M. & Gabrilovich, D. Myeloid-derived suppressor cells coming of age. Nat. Immunol. 19, 108–119 (2018).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Bilate, A. M. & Lafaille, J. J. Induced CD4+Foxp3+ regulatory T cells in immune tolerance. Annu. Rev. Immunol. 30, 733–758 (2012).
Sugiyama, D. et al. Anti-CCR4 mAb selectively depletes effector-type FoxP3+CD4+ regulatory T cells, evoking antitumor immune responses in humans. Proc. Natl Acad. Sci. USA 110, 17945–17950 (2013).
Bopp, T. et al. Cyclic adenosine monophosphate is a key component of regulatory T cell–mediated suppression. J. Exp. Med. 204, 1303–1310 (2007).
Nishikawa, H. & Sakaguchi, S. Regulatory T cells in cancer immunotherapy. Curr. Opin. Immunol. 27, 1–7 (2014).
Bronte, V. et al. Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat. Commun. 7, 12150 (2016).
Huang, L. R. et al. Intrahepatic myeloid-cell aggregates enable local proliferation of CD8+ T cells and successful immunotherapy against chronic viral liver infection. Nat. Immunol. 14, 574–583 (2013).
Pallett, L. J. et al. Metabolic regulation of hepatitis B immunopathology by myeloid-derived suppressor cells. Nat. Med. 21, 591–600 (2015).
Tcyganov, E., Mastio, J., Chen, E. & Gabrilovich, D. I. Plasticity of myeloid-derived suppressor cells in cancer. Curr. Opin. Immunol. 51, 76–82 (2018).
Kumar, V. et al. Cancer-associated fibroblasts neutralize the anti-tumor effect of CSF1 receptor blockade by inducing PMN-MDSC infiltration of tumors. Cancer Cell 32, 654–668.e5 (2017).
Hochst, B. et al. Activated human hepatic stellate cells induce myeloid derived suppressor cells from peripheral blood monocytes in a CD44-dependent fashion. J. Hepatol. 59, 528–535 (2013).
Ostrand-Rosenberg, S. & Fenselau, C. Myeloid-derived suppressor cells: immune-suppressive cells that impair antitumor immunity and are sculpted by their environment. J. Immunol. 200, 422–431 (2018).
Klein Geltink, R. I. et al. Mitochondrial priming by CD28. Cell 171, 385–397.e11 (2017).
Chang, C. H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Lu, Z. & Hunter, T. Metabolic kinases moonlighting as protein kinases. Trends Biochem. Sci. 43, 301–310 (2018).
Menk, A. V. et al. Early TCR signaling induces rapid aerobic glycolysis enabling distinct acute T cell effector functions. Cell Rep. 22, 1509–1521 (2018).
Buck, M. D., Sowell, R. T., Kaech, S. M. & Pearce, E. L. Metabolic instruction of immunity. Cell 169, 570–586 (2017).
Onfelt, B., Nedvetzki, S., Yanagi, K. & Davis, D. M. Cutting edge: membrane nanotubes connect immune cells. J. Immunol. 173, 1511–1513 (2004).
Watkins, S. C. & Salter, R. D. Functional connectivity between immune cells mediated by tunneling nanotubules. Immunity 23, 309–318 (2005).
O’Neill, L. A. & Pearce, E. J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 213, 15–23 (2016).
Ganeshan, K. & Chawla, A. Metabolic regulation of immune responses. Annu. Rev. Immunol. 32, 609–634 (2014).
El-Mir, M. Y. et al. Dimethylbiguanide inhibits cell respiration via an indirect effect targeted on the respiratory chain complex I. J. Biol. Chem. 275, 223–228 (2000).
Owen, M. R., Doran, E. & Halestrap, A. P. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem. J. 348, 607–614 (2000).
Wheaton, W. W. et al. Metformin inhibits mitochondrial complex I of cancer cells to reduce tumorigenesis. Elife 3, e02242 (2014).
Teeter, M. E., Baginsky, M. L. & Hatefi, Y. Ectopic inhibition of the complexes of the electron transport system by rotenone, piericidin A, demerol and antimycin A. Biochim. Biophys. Acta 172, 331–333 (1969).
Kinsky, O. R. et al. Metformin scavenges methylglyoxal to form a novel imidazolinone metabolite in humans. Chem. Res. Toxicol. 29, 227–234 (2016).
Beisswenger, P. & Ruggiero-Lopez, D. Metformin inhibition of glycation processes. Diabetes Metab. 29, 6S95–6S103 (2003).
Han, J., Gagnon, S., Eckle, T. & Borchers, C. H. Metabolomic analysis of key central carbon metabolism carboxylic acids as their 3-nitrophenylhydrazones by UPLC/ESI-MS. Electrophoresis 34, 2891–2900 (2013).
Allaman, I., Belanger, M. & Magistretti, P. J. Methylglyoxal, the dark side of glycolysis. Front. Neurosci. 9, 23 (2015).
Wang, T., Douglass, E. F. Jr., Fitzgerald, K. J. & Spiegel, D. A. A “turn-on” fluorescent sensor for methylglyoxal. J. Am. Chem. Soc. 135, 12429–12433 (2013).
Rabbani, N. & Thornalley, P. J. The dicarbonyl proteome: proteins susceptible to dicarbonyl glycation at functional sites in health, aging, and disease. Ann. NY Acad. Sci. 1126, 124–127 (2008).
Rabbani, N., Xue, M. & Thornalley, P. J. Methylglyoxal-induced dicarbonyl stress in aging and disease: first steps towards glyoxalase 1-based treatments. Clin. Sci. (Lond.) 130, 1677–1696 (2016).
Phillips, S. A. & Thornalley, P. J. The formation of methylglyoxal from triose phosphates. Investigation using a specific assay for methylglyoxal. Eur. J. Biochem. 212, 101–105 (1993).
Ohmori, S., Mori, M., Shiraha, K. & Kawase, M. Biosynthesis and degradation of methylglyoxal in animals. Prog. Clin. Biol. Res. 290, 397–412 (1989).
Ray, S. & Ray, M. Formation of methylglyoxal from aminoacetone by amine oxidase from goat plasma. J. Biol. Chem. 258, 3461–3462 (1983).
Lyles, G. A. & Chalmers, J. The metabolism of aminoacetone to methylglyoxal by semicarbazide-sensitive amine oxidase in human umbilical artery. Biochem. Pharmacol. 43, 1409–1414 (1992).
Mizukoshi, E. et al. Myeloid-derived suppressor cells correlate with patient outcomes in hepatic arterial infusion chemotherapy for hepatocellular carcinoma. Cancer Immunol. Immunother. 65, 715–725 (2016).
Gao, X. H. et al. Circulating CD14+ HLA-DR−/low myeloid-derived suppressor cells predicted early recurrence of hepatocellular carcinoma after surgery. Hepatol. Res. 47, 1061–1071 (2017).
Arihara, F. et al. Increase in CD14+HLA-DR−/low myeloid-derived suppressor cells in hepatocellular carcinoma patients and its impact on prognosis. Cancer Immunol. Immunother. 62, 1421–1430 (2013).
Bronte, V., Serafini, P., Mazzoni, A., Segal, D. M. & Zanovello, P. l-arginine metabolism in myeloid cells controls T-lymphocyte functions. Trends Immunol. 24, 302–306 (2003).
Geiger, R. et al. l-Arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842.e13 (2016).
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nat. Rev. Immunol. 12, 253–268 (2012).
Gabrilovich, D. I. Myeloid-derived suppressor cells. Cancer Immunol. Res. 5, 3–8 (2017).
Rabbani, N. & Thornalley, P. J. Dicarbonyl stress in cell and tissue dysfunction contributing to ageing and disease. Biochem. Biophys. Res. Commun. 458, 221–226 (2015).
Nemet, I. & Varga-Defterdarovic, L. Methylglyoxal-derived β-carbolines formed from tryptophan and its derivates in the Maillard reaction. Amino Acids 32, 291–293 (2007).
Murray, P. J. Amino acid auxotrophy as a system of immunological control nodes. Nat. Immunol. 17, 132–139 (2016).
Chantranupong, L. et al. The CASTOR proteins are arginine sensors for the mTORC1 pathway. Cell 165, 153–164 (2016).
Saxton, R. A., Chantranupong, L., Knockenhauer, K. E., Schwartz, T. U. & Sabatini, D. M. Mechanism of arginine sensing by CASTOR1 upstream of mTORC1. Nature 536, 229–233 (2016).
Wang, S. et al. Lysosomal amino acid transporter SLC38A9 signals arginine sufficiency to mTORC1. Science 347, 188–194 (2015).
Cheng, C. T. et al. Arginine starvation kills tumor cells through aspartate exhaustion and mitochondrial dysfunction. Commun. Biol. 1, 178 (2018).
Taheri, F. et al. l-Arginine regulates the expression of the T-cell receptor ζ chain (CD3ζ) in Jurkat cells. Clin. Cancer Res. 7, 958s–965s (2001).
Qiu, F. et al. Arginine starvation impairs mitochondrial respiratory function in ASS1-deficient breast cancer cells. Sci. Signal. 7, ra31 (2014).
Clausen, B. E., Burkhardt, C., Reith, W., Renkawitz, R. & Forster, I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 8, 265–277 (1999).
Agarwal, A. et al. Transient opening of the mitochondrial permeability transition pore induces microdomain calcium transients in astrocyte processes. Neuron 93, 587–605.e7 (2017).
Knier, B. et al. Myeloid-derived suppressor cells control B cell accumulation in the central nervous system during autoimmunity. Nat. Immunol. 19, 1341–1351 (2018).
Hoechst, B. et al. A new population of myeloid-derived suppressor cells in hepatocellular carcinoma patients induces CD4+CD25+Foxp3+ T cells. Gastroenterology 135, 234–243 (2008).
Weiskirchen, R. et al. Genetic characteristics of the human hepatic stellate cell line LX-2. PLoS ONE 8, e75692 (2013).
Du, P., Kibbe, W. A. & Lin, S. M. lumi: a pipeline for processing Illumina microarray. Bioinformatics 24, 1547–1548 (2008).
Smyth, G. in Bioinformatics and Computational Biology Solutions Using R and Bioconductor (eds Gentleman, R., Carey, V., Huber, W., Irizarry, R. & Dudoi, S.) 397–420 (Springer, 2005).
Tripathi, S. et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Bausch-Fluck, D. et al. A mass spectrometric-derived cell surface protein atlas. PLoS ONE 10, e0121314 (2015).
Ahmed, N., Argirov, O. K., Minhas, H. S., Cordeiro, C. A. A. & Thornalley, P. J. Assay of advanced glycation endproducts (AGEs): surveying AGEs by chromatographic assay with derivatization by 6-aminoquinolyl-N-hydroxysuccinimidyl-carbamate and application to Nepsilon-carboxymethyl-lysine- and Nepsilon-(1-carboxyethyl)lysine-modified albumin. Biochem. J. 364, 1–14 (2002).
Salomón, T. et al. Ketone body acetoacetate buffers methylglyoxal via a non-enzymatic conversion during diabetic and dietary ketosis. Cell Chem. Biol. 24, 935–943.e7 (2017).
Frank, O., Kreissl, J. K., Daschner, A. & Hofmann, T. Accurate determination of reference materials and natural isolates by means of quantitative 1H NMR spectroscopy. J. Agric. Food Chem. 62, 2506–2515 (2014).
Acknowledgements
We thank J. Schulze, M. Beyer and S. Schmitt (Life and Medical Science Institute, University of Bonn, Germany) for isolating RNA and carrying out Illumina whole-genome arrays; R. Weisskirchen (Institute of Molecular Pathobiochemistry, Experimental Gene Therapy and Clinical Chemistry, RWTH University Hospital Aachen, Germany) and J. Trebicka (Department of Internal Medicine I, University Clinic Bonn, Germany) for providing LX2 cells; R. Berger (Institute of Molecular Immunology and Experimental Oncology, Technische Universität München, Munich, Germany) for performing the seahorse experiments; and C. Llanto, S. Michailidou and S. Hegenbarth for technical support. P.A.K. and M. Heikenwälder were supported by the German Research Council (SFB TRR179) and German Center for Infection Research, Munich site. B.H. was supported by German Cancer Aid. T.K. was supported by the German Research Council (SFB1054, TR128, TR274 and SyNergy (EXC 2145; ID 390857198)) and the ERC (CoG 647215).
Author information
Authors and Affiliations
Contributions
T. Baumann, A.D., C.S., S.S., M. Hiltensperger, K.L., V.L., U.A., B.L.-D., J.S., L.S., N.K., T. Bauer, M.L., K.E., S.E., J.E.H., M.A., M.S. and A.H. performed the experiments and analyzed the data. S.D., J.S. and U.A. performed the bioinformatics analyses. N.H., D.H., B.S., D.S., F.A., T.W., C.F., M.S., T.M., H.Z., M. Heikenwälder, T.K., C.F., C.D. and T.H. contributed specific technologies and reagents. B.S., D.S., T.M., H.Z., M. Heikenwälder, T.K., C.D., T.H., P.J.M., P.A.K. and B.H. designed the experiments. P.J.M., P.A.K. and B.H. wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Differentially expressed genes in human MDSCs compared to monocytes.
a, heatmap of differentially expressed genes in stromal cell-induced human MDSCs compared to monocytes. b, no difference in expression of genes coding for surface molecules between MDSCs and monocytes (n = 3).
Extended Data Fig. 2 Mechanistic characterization of stromal-cell induced MDSC-mediated suppression of T cell proliferation.
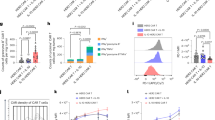
a-i, characterization of stromal cell-induced, human MDSCs and monocytes from the same donor. a, arginase activity measured by urea production (106 cells; n = 6 biological independent samples). b, nitric oxide-production (106 cells; n = 5 biological independent samples). c, ROS production measured by 5 µM 2,7-Dichlorodihydrofluorescein diacetate (H2DCFDA). d, CD274 (PD-L1) expression (n = 6). e, IL-10 concentration in cell culture supernatant (106 cells/ml; n = 5 biological independent samples). f and g, mRNA expression levels of TGF-beta (n = 5 biological independent samples) and Indolamine-2,3-Dioxygenase-I (IDO-I; n = 5 biological independent samples). h-i, proliferation of anti-CD3/CD28 activated CD8 T cells after co-culture with monocytes or MDSCs in presence of inhibitors: L-NOHA, arginase-I inhibitor (10 µM); L-NMMA, NO-synthase inhibitor (10 µM); MnTBAP, superoxide dismutase mimetic and peroxynitrite scavenger (40 µM); 1-MT, IDO-inhibitor (20 µM) or blocking antibodies against TGF-beta, IL-10 or PD-1 (40 µg/ml each) or transwell-insert (0.4 µm) (triplicates; n = 3). h, (n = 8 biological independent samples). j, inhibition of T cell proliferation by tumor-derived MDSCs (CD14+HLADR-/low cells; n = 3 biological independent replicates; proliferation index is plotted). ns = not significant; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; two-way unpaired t-test.
Extended Data Fig. 3 Contact with MDSCs impairs T cell receptor cell signaling.
a-d, human CD8 T cells were activated with anti-CD3/CD28 in co-culture with MDSCs or monocytes (ratio 1:1) for 30 minutes followed by separation of CD8 T cells by FACSorting. a, pan-kinase activity in activated T cells after contact with MDSCs for 30 minutes (n = 3 biological independent samples). b, immunoblot of T cell kinase phosphorylation at indicated time points after anti-CD3/CD28 activation (n = 3 biological independent experiments). c, purity ≥ 99 % of FACSorted CD8 T cells (n = 3 biological independent experiments). d, time kinetics of T cell kinase phosphorylation detected by intracellular staining with phosphorylation-specific antibodies by flow cytometry (n = 3 biological independent samples). ns = not significant; *p < 0.05; **p < 0.01; ***p < 0.001; two-way unpaired t-test.
Extended Data Fig. 4 CD8 T cells show metabolic phenocopies of monocytes and MDSCs after co-culture.
a-f, analysis of anti-CD3/CD28 activated human CD8 T cells after coculture with MDSCs or monocytes (ratio 1:1) for 30 minutes. a, cell surface Glut-1 expression levels (n = 3 biological independent samples). b, glucose (2-NBDG) uptake (n = 3 biological independent samples). c, hexokinase activity from FACSorted CD8 T cells (triplicates; n = 3). d, glycolytic rates in FACSorted CD8 T cells by extracellular flux analysis (n = 3 biological independent samples). e, oxygen consumption rates (OCR) of FACSorted CD8 T cells (n = 3 biological independent samples). f, ATP content of CD8 T cells after FACSorting (n = 3 biological independent samples;). ns = not significant; *p < 0.05; **p < 0.01; ***p < 0.001; two-way unpaired t-test plotted as SEM.
Extended Data Fig. 5 MDSCs suppress T cell effector functions.
CD8 T cell subpopulations (naïve, effector, central memory, effector memory; from left to right) were isolated by FACS-Sorting and activated with anti-CD3/CD28 in co-culture with MDSCs or monocytes or left alone as control. a and b, intracellular staining for IFNγ, TNF and granzyme B (n = 3 independent biological experiments). c and d, quantification from (a,b). e, Granzyme B release from FACSorted CD8 T cell populations stimulated with CEF (105 T cells/ well; 2 µg/ml CEF) quantified by ELISPOT (n = 3 independent biological experiments). f and g, proliferation of FACS-Sorted CD8 T cell subpopulations measured by CFSE-dilution and quantification (n = 3 biological independent samples). Two-way unpaired t-test plotted as SEM.
Extended Data Fig. 6 Transfer of cytosolic constituents from MDSCs to CD4 T cells and NK cells.
a, transfer of cytosolic constituents from MDSCs into NKT cells and CD4 T cells (n = 3). b, gating strategy used in c and d. adoptive transfer of CD45.1+CD8 T cells (106) into CD45.2+ LysM-Cre/Rosa-mito-GFP mice tumor bearing mice (B16 melanoma cells), and flow cytometric analysis for GFP fluorescence in transferred CD45.1+CD8+ cells isolated from tumor tissue and spleen directly ex vivo at 24 hours after transfer (n = 4). d, proliferation of isolated and activated CD8 T cells from tumor tissue and spleen was analyzed on day 3 using flow cytometry (c, d, representative plots of n = 4 biological independent samples).
Extended Data Fig. 7 Detection of methylglyoxal and its generation and neutralization.
a, derivatization of carbonyl compounds with 3-Nitrophenylhydrazine (3-NPH) for detection by mass spectrometry. b, Methylglyoxal content measurement in different immune cells from the peripheral blood of HCC patients (n = 3). c, Chemical formulas for molecules with guanidine-groups that are targets of methylglyoxal (red boxes).
Extended Data Fig. 8 Guanidine-treatment of MDSCs abrogates their suppressive activity on CD8 T cell effector functions.
a-c and e,f, human CD8 T cells were cultures in the presence of MDSCs that were partially pretreated with indicated compounds. a, T cells were stimulated alone, treated with DMBG, in the presence of MDSCs (pretreated with DMBG, methylguanidine, aminoguanidine or rodenidine (200 µM each). Proliferation (a) or cytotoxic activity (b) was measured by the dilution of CFSE (n = 3) or by ELISpot (n = 3 biological independent samples). d, Suppressive activity of CD11b+Ly6C+ or CD11b+Ly6G+ cells from the central nerves system during the recovery phase of experimental autoimmune encephalomyelitis (EAE; day 22 after immunization partially pretreated with DMBG (n = 3). e activated CD8 T cells were cocultured with CD4+CD25+CD127- regulatory T cells. Were Indicated, DMBG (200 µM) were added and proliferation was measured on day 3 (n = 3 biological independent samples). f, CD15+ cells were isolated from blood from the same patient were isolated an cocultured with CFSE labeled, activated CD8 T cells. Proliferation was measured by the dilution of CFSE (n = 2 biological independent samples). g, h, T cells were cocultivated with MDSCs for 30 min and reisolated. The glucose uptake capacity (2-NBDG) was measured at 30 min intervals (n = 3 biological independent samples). h, continuous glucose uptake measurement after addition of DMBG (n = 3), live-dead discrimination of CD8 T cells cultured alone, with monocytes or with MDSCs (n = 5 biological independent samples). j, cell count of CD8 T cells cultured alone, with monocytes or with MDSCs (n = 3 biological independent samples). *p < 0.05; **p < 0.01; ***p < 0.001; two-way unpaired t-test.
Extended Data Fig. 9 Monocytic cells affect the amino acid composition of CD8 T cells.
Free amino acids and advanced glycation products were measured using SIDA-UHPLC-MS/MSMRM in T cells after co-culture with MDSCs or monocytes. (n = 4 biological independent samples (3 biological samples for T cells without stimulation)). ns = not significant; two-way unpaired t-test; indicating no significant differences between any groups.
Extended Data Fig. 10 DMBG restores effector function of vaccination-induced CD8 T cells in cancer tissue.
At d10 after s.c. B16-OVA cancer cell inoculation, mice received ovalbumin adjuvanted with CpG/alphaGalCer, anti-PD-1 (200 µg/every 3. Day) and/or DMBG in drinking water (40 mM), and analyses were performed at d17 (5 animals per group; n = 3). a, Kaplan-Meier survival curves of tumor bearing mice (n = 5). b, the expression of ovalbumin was tested in B16-melanoma in cell culture (1), ex vivo after vaccination on day 17 (2) and on day 34 after combined therapy using anti-PD-1 and DMBG alone (3,4) or in combination with vaccination (5,6) (n = 3 independen biological samples). c - e, MBo-fluorescence and glucose uptake of CD11b+ cells from spleen. f, g CD8 T cell proliferation (CFSE-dilution) in co-culture with CD11b+Ly6C+ cells or CD11b+Ly6G+ (FACSorted) from cancer tissue or spleen (numbers denote division indices). h, i, MBo-fluorescence and glucose uptake of CD8+ T cells from the spleen (a, c – e: n = 6 biological independent samples; g – i: n = 5 biological identical samples). ns = not significant; indicating no significant differences between any groups.; *p < 0.05; **p < 0.01; ***p < 0.001; two-way unpaired t-test or 2-sided Mantel-Cox test (survival curve).
Supplementary information
Supplementary Tables
Supplementary Table 1: Patient classification. Supplementary Table 2: All DEGs in MDSCs compared with monocytes. Supplementary Table 3: Immune regulatory genes in MDSCs compared with monocytes. Supplementary Table 4: Significantly DEGs in MDSCs compared with monocytes. Supplementary Table 5: All differentially expressed surface markers in MDSCs compared with monocytes (Cell Surface Protein Atlas dataset)66. Supplementary Table 6: All differentially expressed glycolytic enzyme genes in MDSCs compared with monocytes. Supplementary Table 7: Metabolites in MDSCs compared with monocytes. Supplementary Table 8: MRM transitions and optimized MS/MS parameters for AGP analysis. Supplementary Table 9: MRM transitions and optimized MS/MS parameters for amino acid analysis.
Supplementary Video 1
Three-dimensional reconstruction from high-resolution confocal imaging of the interaction between a MitoTracker Green–labeled MDSC (right cell in image) interacting with a T cell (left cell in image), illustrating transfer of the MitoTracker Green label or labeled cytosolic constituents from the MDSC to the T cell and the formation of a physical connection between the two cells that might allow for transfer of cytosolic constituents.
Supplementary Video 2
Time-lapse confocal microscopy of the interaction between anti-CD8-eF670-labeled T cells and MitoTracker Green–labeled MDSCs, showing that eF670-positive T cells acquire MitoTracker Green fluorescence only when located in the direct vicinity of MDSCs, to allow for physical contact between the two cell populations.
Source data
Source Data Fig. 1
Additional statistical information
Source Data Fig. 2
Additional statistical information
Source Data Fig. 4
Additional statistical information
Source Data Fig. 5
Additional statistical information
Source Data Fig. 6
Additional statistical information
Source Data Fig. 7
Additional statistical information
Source Data Extended Data Fig. 2
Additional statistical information
Source Data Extended Data Fig. 3
Additional statistical information
Source Data Extended Data Fig. 4
Additional statistical information
Source Data Extended Data Fig. 5
Additional statistical information
Source Data Extended Data Fig. 8
Additional statistical information
Source Data Extended Data Fig. 9
Additional statistical information
Source Data Extended Data Fig. 10
Additional statistical information
Rights and permissions
About this article
Cite this article
Baumann, T., Dunkel, A., Schmid, C. et al. Regulatory myeloid cells paralyze T cells through cell–cell transfer of the metabolite methylglyoxal. Nat Immunol 21, 555–566 (2020). https://doi.org/10.1038/s41590-020-0666-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0666-9