Abstract
Adaptive evolution is a key feature of T cell immunity. During acute immune responses, T cells harboring high-affinity T cell antigen receptors (TCRs) are preferentially expanded, but whether affinity maturation by clonal selection continues through the course of chronic infections remains unresolved. Here we investigated the evolution of the TCR repertoire and its affinity during the course of infection with cytomegalovirus, which elicits large T cell populations in humans and mice. Using single-cell and bulk TCR sequencing and structural affinity analyses of cytomegalovirus-specific T cells, and through the generation and in vivo monitoring of defined TCR repertoires, we found that the immunodominance of high-affinity T cell clones declined during the chronic infection phase, likely due to cellular senescence. These data showed that under conditions of chronic antigen exposure, low-affinity TCRs preferentially expanded within the TCR repertoire, with implications for immunotherapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this article and its Supplementary Information files. For differentially expressed genes between high and low TCR avidity T cells (related to Fig. 6) see Supplementary Table 1. Additional raw data are available from the corresponding authors upon request.
Code availability
For algorithms and equations used for RNA sequencing and CDR3 analysis see Methods section. Additional information is available from the corresponding authors upon reasonable request.
References
Brodin, P. et al. Variation in the human immune system is largely driven by non-heritable influences. Cell 160, 37–47 (2015).
Klenerman, P. & Oxenius, A. T cell responses to cytomegalovirus. Nat. Rev. Immunol. 16, 367–377 (2016).
Karrer, U. et al. Memory inflation: continuous accumulation of antiviral CD8+ T cells over time. J. Immunol. 170, 2022–2029 (2003).
Tscharke, D. C., Croft, N. P., Doherty, P. C. & La Gruta, N. L. Sizing up the key determinants of the CD8+ T cell response. Nat. Rev. Immunol. 15, 705–716 (2015).
Busch, D. H. & Pamer, E. G. T cell affinity maturation by selective expansion during infection. J. Exp. Med. 189, 701–710 (1999).
Savage, P. A., Boniface, J. J. & Davis, M. M. A kinetic basis for T cell receptor repertoire selection during an immune response. Immunity 10, 485–492 (1999).
Day, E. K. et al. Rapid CD8+ T cell repertoire focusing and selection of high-affinity clones into memory following primary infection with a persistent human virus: human cytomegalovirus. J. Immunol. 179, 3203–3213 (2007).
Iancu, E. M. et al. Clonotype selection and composition of human CD8 T cells specific for persistent herpes viruses varies with differentiation but is stable over time. J. Immunol. 183, 319–331 (2009).
Trautmann, L. et al. Selection of T cell clones expressing high-affinity public TCRs within human cytomegalovirus-specific CD8 T cell responses. J. Immunol. 175, 6123–6132 (2005).
Price, D. A. et al. Avidity for antigen shapes clonal dominance in CD8+ T cell populations specific for persistent DNA viruses. J. Exp. Med. 202, 1349–1361 (2005).
Ouyang, Q. et al. Large numbers of dysfunctional CD8+ T lymphocytes bearing receptors for a single dominant CMV epitope in the very old. J. Clin. Immunol. 23, 247–257 (2003).
Griffiths, S. J. et al. Age-associated increase of low-avidity cytomegalovirus-specific CD8+ T cells that re-express CD45RA. J. Immunol. 190, 5363–5372 (2013).
Khan, N. et al. T cell recognition patterns of immunodominant cytomegalovirus antigens in primary and persistent infection. J. Immunol. 178, 4455–4465 (2007).
Khan, N. et al. Herpesvirus-specific CD8 T cell immunity in old age: cytomegalovirus impairs the response to a coresident EBV infection. J. Immunol. 173, 7481–7489 (2004).
Nauerth, M. et al. TCR-ligand k off rate correlates with the protective capacity of antigen-specific CD8+ T cells for adoptive transfer. Sci. Transl. Med. 5, 192ra87 (2013).
Dekhtiarenko, I., Jarvis, Ma, Ruzsics, Z. & Čičin-Šain, L. The context of gene expression defines the immunodominance hierarchy of cytomegalovirus antigens. J. Immunol. 190, 3399–3409 (2013).
Nauerth, M. et al. Flow cytometry-based TCR-ligand K off-rate assay for fast avidity screening of even very small antigen-specific T cell populations ex vivo. Cytometry A 89, 816–825 (2016).
Dössinger, G. et al. MHC multimer-guided and cell culture-independent isolation of functional T cell receptors from single cells facilitates TCR identification for immunotherapy. PLoS One 8, e61384 (2013).
Munks, M. W. et al. Four distinct patterns of memory CD8 T cell responses to chronic murine cytomegalovirus infection. J. Immunol. 177, 450–458 (2006).
Enouz, S. et al. Autoreactive T cells bypass negative selection and respond to self-antigen stimulation during infection. J. Exp. Med. 209, 1769–1779 (2012).
Zehn, D., Lee, S. Y. & Bevan, M. J. Complete but curtailed T cell response to very low-affinity antigen. Nature 458, 211–214 (2009).
Rosette, C. et al. The impact of duration versus extent of TCR occupancy on T cell activation: a revision of the kinetic proofreading model. Immunity 15, 59–70 (2001).
Nauerth, M., Weissbrich, B. & Busch, D. H. The clinical potential for koff-rate measurement in adoptive immunotherapy. Expert Rev. Clin. Immunol. 9, 1151–1153 (2013).
Holst, J., Vignali, K. M., Burton, A. R. & Vignali, D. A. A. Rapid analysis of T cell selection in vivo using T cell-receptor retrogenic mice. Nat. Methods 3, 191–197 (2006).
Grassmann, S. et al. Distinct surface expression of activating receptor Ly49H drives differential expansion of NK cell clones upon murine cytomegalovirus infection. Immunity 50, 1391–1400.e4 (2019).
Obar, J. J., Khanna, K. M. & Lefrançois, L. Endogenous naive CD8+ T cell precursor frequency regulates primary and memory responses to infection. Immunity 28, 859–869 (2008).
Buchholz, V. R. et al. Disparate individual fates compose robust CD8+ T cell immunity. Science 340, 630–635 (2013).
Best, J. A. et al. Transcriptional insights into the CD8+ T cell response to infection and memory T cell formation. Nat. Immunol. 14, 404–412 (2013).
Akbar, A. N. & Henson, S. M. Are senescence and exhaustion intertwined or unrelated processes that compromise immunity? Nat. Rev. Immunol. 11, 289–295 (2011).
Čičin-Šain, L. & Arens, R. Exhaustion and inflation at antipodes of T cell responses to chronic virus infection. Trends Microbiol. 26, 498–509 (2018).
Marchi, E., Lee, L. N. & Klenerman, P. Inflation vs. exhaustion of antiviral CD8+ T cell populations in persistent infections: two sides of the same coin? Front. Immunol. 10, 197 (2019).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Davenport, M. P., Fazou, C., McMichael, A. J. & Callan, M. F. C. Clonal selection, clonal senescence, and clonal succession: the evolution of the T cell response to infection with a persistent virus. J. Immunol. 168, 3309–3317 (2002).
Akbar, A. N., Henson, S. M. & Lanna, A. Senescence of T lymphocytes: implications for enhancing human immunity. Trends Immunol. 37, 866–876 (2016).
Redeker, A. et al. The contribution of cytomegalovirus infection to immune senescence is set by the infectious dose. Front. Immunol. 8, 1953 (2018).
Wang, G. C., Dash, P., McCullers, J. A., Doherty, P. C. & Thomas, P. G. T cell receptor αβ diversity inversely correlates with pathogen-specific antibody levels in human cytomegalovirus infection. Sci. Transl. Med. 4, 128ra42 (2012).
Baumann, N. S. et al. Early primed KLRG1-CMV-specific T cells determine the size of the inflationary T cell pool. PLoS Pathog. 15, e1007785 (2019).
Stemberger, C. et al. A single naive CD8+ T cell precursor can develop into diverse effector and memory subsets. Immunity 27, 985–997 (2007).
Moon, J. J. et al. Naive CD4+ T cell frequency varies for different epitopes and predicts repertoire diversity and response magnitude. Immunity 27, 203–213 (2007).
Germain, R. N. The art of the probable: system control in the adaptive immune system. Science 293, 240–245 (2001).
Schober, K., Buchholz, V. R. & Busch, D. H. TCR repertoire evolution during maintenance of CMV-specific T cell populations. Immunol. Rev. 283, 113–128 (2018).
Hebeisen, M. et al. Identifying individual T cell receptors of optimal avidity for tumor antigens. Front. Immunol. 6, 582 (2015).
Martinez, R. J. & Evavold, B. D. Lower affinity T cells are critical components and active participants of the immune response. Front. Immunol. 6, 468 (2015).
Anderton, S. M., Radu, C. G., Lowrey, P. A., Ward, E. S. & Wraith, D. C. Negative selection during the peripheral immune response to antigen. J. Exp. Med. 193, 1–12 (2001).
Viganò, S. et al. Functional avidity: a measure to predict the efficacy of effector T cells? Clin. Dev. Immunol. 2012, 153863 (2012).
Lin, M. Y. & Welsh, R. M. Stability and diversity of T cell receptor repertoire usage during lymphocytic choriomeningitis virus infection of mice. J. Exp. Med. 188, 1993–2005 (1998).
Lichterfeld, M. et al. Selective depletion of high-avidity human immunodeficiency virus type 1 (HIV-1)-specific CD8+ T cells after early HIV-1 infection. J. Virol. 81, 4199–4214 (2007).
Sherwood, A. M. et al. Tumor-infiltrating lymphocytes in colorectal tumors display a diversity of T cell receptor sequences that differ from the T cells in adjacent mucosal tissue. Cancer Immunol. Immunother. 62, 1453–1461 (2013).
Cui, J.-H. et al. TCR repertoire as a novel indicator for immune monitoring and prognosis assessment of patients with cervical cancer. Front. Immunol. 9, 2729 (2018).
Guedan, S., Ruella, M. & June, C. H. Emerging cellular therapies for cancer. Annu. Rev. Immunol. 37, 145–171 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Pope, C. et al. Organ-specific regulation of the CD8 T cell response to Listeria monocytogenes infection. J. Immunol. 166, 3402–3409 (2001).
Busch, D. H., Pilip, I. & Pamer, E. G. Evolution of a complex T cell receptor repertoire during primary and recall bacterial infection. J. Exp. Med. 188, 61–70 (1998).
Effenberger, M. et al. FLEXamers: a double tag for universal generation of versatile peptide-MHC multimers. J. Immunol. 202, 2164–2171 (2019).
Eltahla, A. A. et al. Linking the T cell receptor to the single cell transcriptome in antigen-specific human T cells. Immunol. Cell Biol. 94, 604–611 (2016).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Rényi, A. On measures of entropy and information. Proc. Fourth Berkeley Symp. Math. Stat. Probab. 1, 547–561 (1961).
Rempala, G. A. & Seweryn, M. Methods for diversity and overlap analysis in T cell receptor populations. J. Math. Biol. 67, 1339–1368 (2013).
Acknowledgements
We thank members of the Busch laboratory for experimental help and critical discussion, particularly M. Hammel, A. Hochholzer, F. Graml and E. d’Ippolito. We also thank A. Eltahla and F. Luciani for their advice on RNA-seq, A. Briggs for support with CDR3 sequencing and J. Leidinger for support with TRBV NMDS analysis. This work was mainly supported by the German Center for Infection Research (DZIF) and the Deutsche Forschungsgemeinschaft (DFG) (SFB 1321/TP17; SFB 1054/B09 and SFB 1371/TP04). L.C.S. was supported by the FOR2830/TP5 grant (DFG).
Author information
Authors and Affiliations
Contributions
K.S., V.R.B. and D.H.B. conceptualized the study; K.S., S.G. and D.H.B. developed methodology; P.L., P.H., E.N. and A.R. developed and performed software analyses; K.S., F.V., P.H., E.N. and L.F. conducted formal analysis of the data; K.S., F.V., S.G., T.R.M., J.E., S.J., B.W., L.M. and C.R.C. performed experiments; L.B., L.K., J.L., G.D., M.N., L.B., J.D.O., L.C.S. and D.H.B. contributed resources; K.S. and D.H.B. wrote the manuscript; all authors read and approved the manuscript; D.H.B. acquired most of the funding; K.S. and D.H.B. supervised the study and administered the project.
Corresponding authors
Ethics declarations
Competing interests
D.H.B. is co-founder of STAGE Cell Therapeutics GmbH (now Juno Therapeutics/Celgene) and T Cell Factory B.V. (now Kite/Gilead). D.H.B. has a consulting contract with and receives sponsored research support from Juno Therapeutics/Celgene. C.R.C. and A.R. are employees of Juno Therapeutics/Celgene. E.N. and L.F. are employees of ENPICOM B.V. The other authors declare no competing interests.
Additional information
Editor recognition statement Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
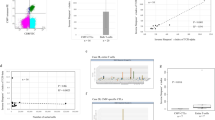
a, pMHC multimer+ CD8+ populations from PBMCs of hCMV+ healthy donors specific for HLA A2/pp65495-503, HLA B7/pp65417-426, HLA B8/IE188-96, HLA B8/IE1199-207KM and HLA A2/IE1316-324. Depicted is population mean and individual t1/2 of single-cell pMHC-TCR koff-rates measured by confocal microscopy (left) and correlation of t1/2 with population size (right). D2, D6 and D11 populations refer to the same donor. n = 15, 17, 8, 22, 50, 51, 11, 62, 20, 26, 22, 57, 12, 29, 26, 18 replicate measurements (from left to right). b, Phenotype of H2Kb/OVA257–264 pMHC multimer+ CD8+ T cells after infection with mCMVIE2-SL in the spleen. Depicted is population mean ± SD of n = 12 mice from two independent experiments. Statistical testing by mixed-effects model (REML) analysis (time and phenotype each ***) followed by Tukey’s multiple comparisons test.
Extended Data Fig. 2
a, Simpson index of TRBV repertoires of H2Kb/OVA257–264 pMHC multimer+ or multimer- CD8+ T cells after infection with mCMVM45-SL or mCMVIE2-SL in the spleen. Depicted is mean ± SEM. n = 12 mice from two independent experiments. Statistical testing by mixed-effects model (REML) analysis (time, cohorts and time x cohorts all ****) followed by Tukey’s multiple comparisons test. Shown asterisks indicate comparisons against Mult- populations from the same infection system. b, Structural affinity and population sizes of H2Kb/OVA257–264 pMHC multimer+ CD8+ T cells after infection with mCMVIE2-SL in the spleen. Mean ± SEM of population size is depicted versus population mean ± SEM of t1/2 of population pMHC-TCR koff-rates measured by confocal microscopy with n 2-7 mice per time point ((d8 3, d15 7, d30 2, d58 4, d > 200 7) and random adjustment for equal cell numbers for each population). No error bars for d30 t1/2 values since n = 2 for this time point. For analogous results using flow cytometry also see Fig. 1b. c, Structural affinity of H2Kb/M38316–323 pMHC multimer+ CD8+ T cells after infection with mCMVWT in peripheral blood as measured by flow cytometry (left; mean and measurements from n = 4 mice of t1/2 of population pMHC-TCR koff-rates; statistical testing by one-way ANOVA (*) and Tukey’s multiple comparisons test) or in spleens as measured by confocal microscopy (right; with 9 (d 15) and 5 (d > 177) mice per time point and random adjustment for equal cell numbers for each population, n = 335 (d 15) and 122 (d > 177) single-cell measurements per time point; statistical testing by two-tailed Mann-Whitney test). d, Single-cell pMHC-TCR koff-rates of H2Kb/OVA257–264 pMHC multimer+ CD8+ T cells after infection with mCMVIE2-SL measured by confocal microscopy after sorting by CD8 positivity or CD4 negativity. n = 61 (no CD8) and 61 (CD8 clone 5H10) measurements. Statistical testing by two-tailed Mann-Whitney test. e, Single-cell pMHC-TCR koff-rates measured by confocal microscopy of naïve OT-I T cells or inflationary memory OT-I T cells isolated from peripheral blood 390 days post mCMVIE2-SL infection. n = 85 (OT-I naive) and 11 (OT-I day 390 p.i.) measurements. Statistical testing by two-tailed Mann-Whitney test. f, pMHC-TCR koff-rates measured by confocal microscopy (PB, peripheral blood; LN, lymph node) or flow cytometry (BM, bone marrow; liver) of inflationary endogenous H2Kb/OVA257–264 pMHC multimer+ CD8+ T cells at day > 200 post mCMVIE2-SL infection. In order to make measurements using two different assays comparable, data were normalized to t1/2 from spleen and pooled. Statistical testing by one-way ANOVA. n = 2 (PB), 4 (LN), 3 (BM) and 3 (liver). Mean ± SEM of population size is depicted, no error bars for PB since n = 2. g, Absolute and relative (% of CD8) cell numbers of OT-I and endogenous pMHC multimer+ T cells at day 15 and 199 after mCMVIE2-SL infection (and transfer of 100 OT-I cells) as well as ratio of OT-I over endogenous pMHC multimer+ T cells, in salivary glands and spleen. n = 3 replicates. Statistical testing by one-way ANOVA. Mean ± SEM of population size is depicted. h, ‘Relative’ and ‘absolute distribution’ of pMHC-TCR koff-rates from Fig. 2e (mCMVIE2-SL). Relative distribution indicates percentage of pMHC multimer+ T cells within indicated t1/2 categories. Absolute distribution additionally takes size of pMHC multimer+ population into account (A.U., arbitrary unit, indicates percentage of pMHC multimer+ T cells within indicated t1/2 categories multiplied by size of pMHC multimer+ population). i, Peptide sensitivity in terms of dose-dependent antigen-specific IFNγ release of H2Kb/OVA257–264 pMHC multimer+ CD8+ T cells from spleen at day 15 or day > 200 post mCMVIE2-SL infection. Representative of two independent experiments. ns, not significant. ** p value < 0.01, *** p value < 0.001, **** p value < 0.0001.
Extended Data Fig. 3
a, Representative sorting scheme and purity control of H2Kb/OVA257–264 pMHC multimer+ CD8+ T cells from peripheral blood for cDNA generation. b, Analysis as in Fig. 3c, but with centroids for mean position for time point repertoires of n = 12 mice in two-dimensional space. Ellipses indicate standard deviation of V1 and V2 (left). Quantification of V1 x V2 standard deviations for different time point repertoires (right). c, Abundance-weighted Morisita-Horn similarity index between repertoires of a representative individual mouse at a specific time point and all previous repertoires in the same mouse. Reference was created by analysis with all possible permutations of time orderings (n=5040 permutations). Experimental: Dot represents value. Reference: Boxes and whiskers indicate median with 5-95 % CI of n = 5040 iterations. d, Delta CT values of TCR excision circles (TREC) normalized over Tcra reads after qPCR of gDNA from CD8+ T cells of 7-week old mice thymectomized at the age of 4 weeks (Thy) or not-thymectomized mice (WT). Mean thymic output of thymectomized animals is 44% of WT animals. n = 10 mice. Depicted is mean ± SEM. Statistical testing by unpaired two-tailed t-test. **** p value < 0.0001. e, Population sizes of H2Kb/M38 and H2Kb/M45 pMHC multimer+ CD8+ T cells after thymectomy (Thy) or no thymectomy (WT) and WT mCMV infection in peripheral blood. n = 8-20 mice from 2-4 independent experiments (WT M38: d7 18, d10 8, d15 17, d60 12, d > 200 20; Thy M38: d7 14, d10 8, d15 18, d60 19, d > 200 19; WT M45: d7 18, d10 8, d15 16, d60 12, d > 200 11; Thy M45: d7 12, d10 8, d15 16, d60 19, d > 200 9). Depicted is mean ± SEM. Sham operation is equivalent to no thymectomy (data not shown). Statistical testing by mixed-effects model (REML) analysis (time, treatment and time x treatment all ****) followed by Tukey’s multiple comparisons test. Specific statistical testing between treatments and antigen-specificities is indicated in the graph. ****, p value < 0.0001. f, Simpson index of TRBV repertoires of cells from (e). Depicted is mean ± SEM. n = 12 mice from two independent experiments. Statistical testing by mixed-effects model (REML) analysis (time ***, cohorts and time x cohorts ****) followed by Tukey’s multiple comparisons test. Specific statistical testing between treatments and antigen-specificities is indicated in the graph. *** p value < 0.001, *** p value < 0.001. g, Longitudinal tracking of TRBV usage in peripheral blood of 2 representative non-thymectomized (WT) and 2 representative thymectomized (Thy) mice from two independent experiments.
Extended Data Fig. 4
a, Progeny of 100 CD45.1+ OT-I T cells in peripheral blood transferred one day before mCMVIE2-SL infection (with N4, T4 or V4 APL). Representative data at day 7 and 452 p.i. (left). Transferred cells over time (right). Depicted is replicates with mean. n = 4 mice per group (except N4 d279-425, T4 d169 and d425, and V4 d425: n = 3 mice). Statistical testing by mixed-effects model (REML) analysis with fixed effects (type III). ns, not significant. *, p value < 0.05. Representative of two independent experiments. b, TCR library for H2Kb/OVA257–264 SIINFEKL. From left to right: CDR3α (above) and CDR3β (below) amino acid sequences of 15 TCRs; their respective dissociation kinetics during flow-cytometric pMHC-TCR koff-rate measurement; splenic progeny of 100 CD45.1+ CD8+ T cells from retrogenic mice with the respective TCRs at day 8 after transfer and infection of recipients with Listeria monocytogenes OVA; peptide sensitivity in terms of dose-dependent antigen-specific IFNγ release of cells from previous column.
Extended Data Fig. 5
Endogenous H2Kb/OVA257–264 pMHC multimer+ CD8+ T cells and progeny of 100/300/1000 CD45.1+ CD8+ T cells from TCR 33 retrogenic mice at day 8 after transfer and infection of recipients with mCMVIE2-SL in peripheral blood. n = 3 mice. Depicted is mean ± SEM. Statistical testing by two-way RM ANOVA analysis followed by Tukey’s multiple comparisons test. ns not significant.
Extended Data Fig. 6
a, Representative sorting scheme and purity control of progeny of TCR 33 and 28 retrogenic CD8+ T cells from peripheral blood for cDNA generation (for further experimental details see Fig. 6). b, Percentage of shared genes that are enriched in TCR 33 or TCR 28, and genes that are enriched in phenotypic clusters from28. c, Relative mRNA amount of Sirt1 and CD244 normalized over Actb expression based on qPCR, plotted versus normalized counts from RNA sequencing of same cDNA from Fig. 6.
Extended Data Fig. 7 Gating scheme.
Representative gating scheme for flow-cytometry experiments.
Supplementary information
Supplementary Table 1
Differentially expressed genes between high and low TCR avidity T cells (related to Fig. 6).
Supplementary Video 1
Representative video of day 15 dissociation (related to Fig. 2d,e).
Supplementary Video 2
Representative video of day >200 dissociation (related to Fig. 2d,e).
Rights and permissions
About this article
Cite this article
Schober, K., Voit, F., Grassmann, S. et al. Reverse TCR repertoire evolution toward dominant low-affinity clones during chronic CMV infection. Nat Immunol 21, 434–441 (2020). https://doi.org/10.1038/s41590-020-0628-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0628-2
This article is cited by
-
Interferon-γ couples CD8+ T cell avidity and differentiation during infection
Nature Communications (2023)
-
CD83 expression characterizes precursor exhausted T cell population
Communications Biology (2023)
-
Unique roles of co-receptor-bound LCK in helper and cytotoxic T cells
Nature Immunology (2023)
-
Time-dependent analysis of the impact on early cytomegalovirus reactivation of HLA mismatch and acute graft-versus-host disease after allogeneic hematopoietic cell transplantation from related donors in acquired aplastic anemia
Annals of Hematology (2023)
-
Non-viral precision T cell receptor replacement for personalized cell therapy
Nature (2023)