Abstract
Quantitative mass spectrometry reveals how CD4+ and CD8+ T cells restructure proteomes in response to antigen and mammalian target of rapamycin complex 1 (mTORC1). Analysis of copy numbers per cell of >9,000 proteins provides new understanding of T cell phenotypes, exposing the metabolic and protein synthesis machinery and environmental sensors that shape T cell fate. We reveal that lymphocyte environment sensing is controlled by immune activation, and that CD4+ and CD8+ T cells differ in their intrinsic nutrient transport and biosynthetic capacity. Our data also reveal shared and divergent outcomes of mTORC1 inhibition in naïve versus effector T cells: mTORC1 inhibition impaired cell cycle progression in activated naïve cells, but not effector cells, whereas metabolism was consistently impacted in both populations. This study provides a comprehensive map of naïve and effector T cell proteomes, and a resource for exploring and understanding T cell phenotypes and cell context effects of mTORC1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All proteomics data are available for interrogation using the EPD (https://peptracker.com/epd). Analysed proteomics data used to generate the figures are available in Supplementary Data 1–5. Raw mass spectrometry data files and MaxQuant analysis files are available from the ProteomeXchange data repository (http://proteomecentral.proteomexchange.org/cgi/GetDataset) and can be accessed with the identifier PXD012058. Flow cytometry data that support the findings of this study are available from the corresponding author upon request.
References
Araki, K. et al. Translation is actively regulated during the differentiation of CD8+ effector T cells. Nat. Immunol. 18, 1046–1057 (2017).
Sinclair, L. V. et al. Control of amino-acid transport by antigen receptors coordinates the metabolic reprogramming essential for T cell differentiation. Nat. Immunol. 14, 500–508 (2013).
Ricciardi, S. et al. The translational machinery of human CD4+ T cells is poised for activation and controls the switch from quiescence to metabolic remodeling. Cell Metab. 28, 895–906 (2018).
Geiger, R. et al. l-arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842 (2016).
Hukelmann, J. L. et al. The cytotoxic T cell proteome and its shaping by the kinase mTOR. Nat. Immunol. 17, 104–112 (2016).
Rieckmann, J. C. et al. Social network architecture of human immune cells unveiled by quantitative proteomics. Nat. Immunol. 18, 583–593 (2017).
Tan, H. Y. et al. Integrative proteomics and phosphoproteomics profiling reveals dynamic signaling networks and bioenergetics pathways underlying T cell activation. Immunity 46, 488–503 (2017).
Procaccini, C. et al. The proteomic landscape of human ex vivo regulatory and conventional T cells reveals specific metabolic requirements. Immunity 44, 406–421 (2016).
Duguet, F. et al. Proteomic analysis of regulatory T cells reveals the importance of themis1 in the control of their suppressive function. Mol. Cell. Proteomics 16, 1416–1432 (2017).
Cuadrado, E. et al. Proteomic analyses of human regulatory T cells reveal adaptations in signaling pathways that protect cellular identity. Immunity 48, 1046–1059 (2018).
Valvezan, A. J. & Manning, B. D. Molecular logic of mTORC1 signalling as a metabolic rheostat. Nat. Metab. 1, 321–333 (2019).
Finlay, D. K. et al. PDK1 regulation of mTOR and hypoxia-inducible factor 1 integrate metabolism and migration of CD8+ T cells. J. Exp. Med. 209, 2441–2453 (2012).
Zeng, H. et al. mTORC1 and mTORC2 kinase signaling and glucose metabolism drive follicular helper T cell differentiation. Immunity 45, 540–554 (2016).
Pollizzi, K. N. & Powell, J. D. Regulation of T cells by mTOR: the known knowns and the known unknowns. Trends Immunol. 36, 13–20 (2015).
Araki, K., Youngblood, B. & Ahmed, R. The role of mTOR in memory CD8+ T-cell differentiation. Immunol. Rev. 235, 234–243 (2010).
Terada, N. et al. Rapamycin blocks cell-cycle progression of activated T-cells prior to events characteristic of the middle to late G1 phase of the cycle. J. Cell. Physiol. 154, 7–15 (1993).
Terada, N., Franklin, R. A., Lucas, J. J., Blenis, J. & Gelfand, E. W. Failure of rapamycin to block proliferation once resting cells have entered the cell-cycle despite inactivation of p70 S6 kinase. J. Biol. Chem. 268, 12062–12068 (1993).
Wisniewski, J. R., Hein, M. Y., Cox, J. & Mann, M. A “proteomic ruler” for protein copy number and concentration estimation without spike-in standards. Mol. Cell. Proteomics 13, 3497–3506 (2014).
Mehta, M. M. et al. Hexokinase 2 is dispensable for T cell-dependent immunity. Cancer Metab. 6, 10 (2018).
Varanasi, S. K., Jaggi, U., Hay, N. & Rouse, B. T. Hexokinase II may be dispensable for CD4 T cell responses against a virus infection. PLoS ONE 13, e0191533 (2018).
Swamy, M. et al. Glucose and glutamine fuel protein O-GlcNAcylation to control T cell self-renewal and malignancy. Nat. Immunol. 17, 712–720 (2016).
Sinclair, L. V., Neyens, D., Ramsay, G., Taylor, P. M. & Cantrell, D. A. Single cell analysis of kynurenine and System L amino acid transport in T cells. Nat. Commun. 9, 1981 (2018).
So, L. et al. The 4E-BP–eIF4E axis promotes rapamycin-sensitive growth and proliferation in lymphocytes. Sci. Signal. 9, ra57 (2016).
Lingel, H. et al. CTLA-4-mediated posttranslational modifications direct cytotoxic T-lymphocyte differentiation. Cell Death Differ. 24, 1739–1749 (2017).
Loh, P. G. et al. Structural basis for translational inhibition by the tumour suppressor Pdcd4. Embo J. 28, 274–285 (2009).
Suzuki, C. et al. PDCD4 inhibits translation initiation by binding to elF4A using both its MA3 domains. Proc. Natl Acad. Sci. USA 105, 3274–3279 (2008).
Cham, C. M., Driessens, G., O’Keefe, J. P. & Gajewski, T. F. Glucose deprivation inhibits multiple key gene expression events and effector functions in CD8+ T cells. Eur. J. Immunol. 38, 2438–2450 (2008).
Doedens, A. L. et al. Hypoxia-inducible factors enhance the effector responses of CD8+ T cells to persistent antigen. Nat. Immunol. 14, 1173–1182 (2013).
Jabara, H. H. et al. A missense mutation in TFRC, encoding transferrin receptor 1, causes combined immunodeficiency. Nat. Genet. 48, 74–78 (2016).
Pakos-Zebrucka, K. et al. The integrated stress response. EMBO Rep. 17, 1374–1395 (2016).
Wyant, G. A. et al. mTORC1 activator SLC38A9 is required to efflux essential amino acids from lysosomes and use protein as a nutrient. Cell 171, 642–654 (2017).
Saucedo, L. J. et al. Rheb promotes cell growth as a component of the insulin/TOR signalling network. Nat. Cell Biol. 5, 566–571 (2003).
Bar-Peled, L. et al. A tumor suppressor complex with GAP activity for the Rag GTPases that signal amino acid sufficiency to mTORC1. Science 340, 1100–1106 (2013).
Wolfson, R. L. et al. Sestrin2 is a leucine sensor for the mTORC1 pathway. Science 351, 43–48 (2016).
Chantranupong, L. et al. The CASTOR proteins are arginine sensors for the mTORC1 pathway. Cell 165, 153–164 (2016).
Yang, J. L. et al. Critical roles of mTOR complex 1 and 2 for T follicular helper cell differentiation and germinal center responses. eLife. 5, e17936 (2016).
Lee, K. et al. Mammalian target of rapamycin protein complex 2 regulates differentiation of TH1 and TH2 cell subsets via distinct signaling pathways. Immunity 32, 743–753 (2010).
Sinclair, L. V. et al. Phosphatidylinositol-3-OH kinase and nutrient-sensing mTOR pathways control T lymphocyte trafficking. Nat. Immunol. 9, 513–521 (2008).
Brenes, A., Afzal, V., Kent, R. & Lamond, A. I. The Encyclopedia of Proteome Dynamics: a big data ecosystem for (prote)omics. Nucleic Acids Res. 46, D1202–D1209 (2018).
Ben-Sahra, I. & Manning, B. D. mTORC1 signaling and the metabolic control of cell growth. Curr. Opin. Cell Biol. 45, 72–82 (2017).
Pause, A. et al. Insulin-dependent stimulation of protein synthesis by phosphorylation of a regulator of 5′-cap function. Nature 371, 762–767 (1994).
Altan-Bonnet, G. & Germain, R. N. Modeling T cell antigen discrimination based on feedback control of digital ERK responses. PLoS. Biol. 3, 1925–1938 (2005).
Klein-Hessling, S. et al. NFATc1 controls the cytotoxicity of CD8+ T cells. Nat. Commun. 8, 511 (2017).
Pircher, H., Burki, K., Lang, R., Hengartner, H. & Zinkernagel, R. M. Tolerance induction in double specific T-cell receptor transgenic mice varies with antigen. Nature 342, 559–561 (1989).
Barnden, M. J., Allison, J., Heath, W. R. & Carbone, F. R. Defective TCR expression in transgenic mice constructed using cDNA-based alpha- and beta-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Hughes, C. S. et al. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 10, 757 (2014).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotech. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Acknowledgements
The authors thank members of the Cantrell laboratory for comments on the manuscript, T. Ly for discussions on the data, A. Whigham and R. Clarke from the Flow Cytometry Facility for cell sorting and advice on flow cytometry, and the biological sciences research unit at the University of Dundee. This research was supported by a Wellcome Trust Principal Research Fellowship to D.A.C. (205023/Z/16/Z), a Wellcome Trust Strategic Award to D.A.C. and A.I.L. (105024/Z/14/Z) and a Wellcome Trust Equipment Award to D.A.C. (202950/Z/16/Z). This work is dedicated to Olivia Mason.
Author information
Authors and Affiliations
Contributions
A.J.M.H., L.S., J.L.H. and L.V.S. performed the experiments. J.L.H. performed the liquid chromatography–mass spectrometry analysis. A.J.M.H., J.L.H. and A.B. analyzed the proteomics data. A.B. designed and implemented the EPD. A.I.L. and D.A.C. conceived the project and discussed the data. A.J.M.H. and D.A.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Laurie A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 The cumulative abundance of proteins within T cell populations.
(a) Proteins ranked according to their abundance and plotted against their cumulative abundance. The number of proteins that comprise 25%, 50%, 75% and 100% of the total cellular protein mass is provided adjacent to graphs. (b) Direct comparisons of CD4+ and CD8+ naïve populations and CD4+ and CD8+ effector populations. Proteins that make up the top 75% of naïve and effector proteomes (identified in a), are highlighted with red circles. For a and b, n = 6 biologically independent samples for CD8+ naïve cells and 3 biologically independent samples for each of the other T cell populations. For b, proteins were deemed to change significantly if they had a P value <0.05 (two-tailed t-test with unequal variance) and a fold change > 2 standard deviations from the mean fold change between populations (fold change cut-off indicated with dashed lines).
Supplementary Figure 2 Scaling versus enrichment during T cell differentiation.
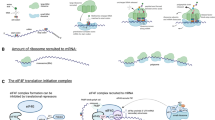
(a) The protein content of cellular compartments and processes during T cell differentiation. The protein content of ribosomes (KEGG 03010), mitochondria (GO:0005739), nuclear envelope (GO:0005635) and the glycolytic pathway was calculated using estimates of protein copy numbers per cell as described in the methods section. Data is also presented showing the proportion of the cell that constitutes ribosomes, mitochondria, nuclear envelope and the glycolytic pathway (presented as a % of the total cellular protein content). (b) Copy numbers and concentration of hexokinase 1 and 2 (HK1 and HK2) in CD4+ and CD8+ cells. (c) Expression profile for tRNA synthetase enzymes in CD8+ T cells. The volcano plot compares the expression profile of enzymes in naïve versus effector CD8+ cells (CTL/naïve copy numbers). The horizontal dashed line indicates a P value = 0.05 (two-tailed t-test with unequal variance), vertical dashed line indicates the mean fold change between populations. The protein mass of these enzymes is also presented. (d) Copy numbers for components of the EIF2 complex – subunits alpha (EIF2S1), beta (EIF2S2) and gamma (EIF2S3). For a-d, n = 6 biologically independent samples for CD8+ naïve cells and 3 biologically independent samples for each of the other T cell populations. Histogram bars represent the mean +/- SD.
Supplementary Figure 3 Environmental sensing in T cells.
(a) The impact of immune activation on the lysosomal arginine sensor SLC38A9, the cytosolic arginine sensor CASTOR1, the leucine sensor SESTRIN2 and the mTORC1 activating GTPase RHEB. Histogram bars represent the mean +/- SD. (b) Copy numbers for GATOR complex members in naïve (N), TCR activated (T) and effector (E) CD8+ populations. Copy numbers are the average of replicates. For a and b, n = 6 biologically independent samples for CD8+ naïve cells and 3 biologically independent samples for each of the other T cell populations.
Supplementary Figure 4 The impact of rapamycin on effector molecules, transcription factors, transporters and fatty acid metabolism.
(a) Volcano plots showing the expression profile of effector molecules in T cells in response to mTORC1 inhibition: Granzyme B, C, D, E and N (GZMB, C, D, E and N); perforin (PRF1); interferon-γ (IFN-γ), lymphotoxin alpha (LTA); lymphotoxin beta (LTB); interleukin 2 (IL2); TNF Superfamily Member 11 (TNFSF11); TNF Superfamily Member 8 (TNFSF8); CD40 ligand (CD40LG). (b) The impact of inhibiting mTORC1 on key transcription factors in T cells – T-Box 21 (TBX21/T-bet), Proto-Oncogene C-Myc (MYC), Basic Leucine Zipper ATF-Like Transcription Factor (BATF), Interferon Regulatory Factor 4 (IRF4) and PR Domain Containing 1 (PRDM1/BLIMP1). (c) Abundance of Hypoxia Inducible Factor 1 Subunit Alpha (HIF-1α) in response to rapamycin. (d) The expression profile of glucose transporters SLC2A1 and SLC2A3 and the lactate transporter SLC16A3 in response to mTORC1 inhibition. (e) The impact of mTORC1 inhibition on proteins involved in fatty acid/sterol metabolism: Hydroxy-3-Methylglutaryl-CoA Synthase 1 (HMGCS1); Fatty Acid Desaturase 1 and 2 (FADS1 and FADS2); Stearoyl-CoA Desaturase 2/3 (SCD2/3). For a, b and e fold change calculated as +rapamycin/control using protein copy numbers. The horizontal dashed line on volcano plots indicates a P value = 0.05 (two-tailed t-test with unequal variance) while vertical dashed lines indicate a fold change of 0.67, 1 and 1.5. For a-e, n = 6 biologically independent samples for CD8+ naïve cells and 3 biologically independent samples for each of the other T cell populations. Histogram bars represent the mean +/- SD.
Supplementary Figure 5 The impact of mTORC1 inhibition on mitochondrial processes and the EIF4A1:PDCD4 complex.
(a) The expression profile for all mitochondrial proteins, (b) mitochondrial ribosome proteins and (c) proteins implicated in oxidative phosphorylation. For a, b and c fold change calculated as +rapamycin/control using protein copy numbers. The horizontal dashed line on volcano plots indicates a P value = 0.05 (two-tailed t-test with unequal variance) while vertical dashed lines indicate a fold change of 0.67, 1 and 1.5. (d) The abundance of Programmed Cell Death 4 (PDCD4) and Eukaryotic Translation Initiation Factor 4A1 (EIF4A1) in CD8+ and CD4+ T cells. Histogram bars represent the mean +/- SD. For a-d, n = 6 biologically independent samples for CD8+ naïve cells and 3 biologically independent samples for each of the other T cell populations.
Supplementary Figure 6 The impact of mTORC1 inhibition on DNA replication proteins and cell cycle protein complexes.
(a) The impact of mTORC1 inhibition on proteins implicated in DNA replication (KEGG annotation 03030 plus the addition of thymidine kinase 1 and thymidine kinase 2). Fold change calculated as +rapamycin/control using protein copy numbers. The horizontal dashed line indicates a P value = 0.05 (two-tailed t-test with unequal variance) while vertical dashed lines indicate a fold change of 0.67, 1 and 1.5. (b) Stochiometric model for cell cycle entry and progression in CD4+ T cells. Protein copy numbers are presented for cyclin D2 (CCND2), cyclin D3 (CCND3), cyclin dependent kinase 4 (CDK4), cyclin dependent kinase 6 (CDK6) and the cyclin dependent kinase inhibitor CDKN1B (P27). (c) The impact of rapamycin on the cyclin D/P27 model in CD4+ cells TCR triggered for 24 h in the presence of rapamycin, and effector TH1 cells incubated with rapamycin for 24 h on day 5 of in vitro culture. For a, b and c, n = 3 biologically independent samples for each T cell populations. For b and c, copy numbers are rounded to the nearest thousand and are the average of biological replicates.
Supplementary Figure 7 Representative gating strategy for DNA synthesis data presented in Fig. 8a and system L amino acid transport assay presented in Fig. 4c.
Supplementary Figure 8 Representative flow cytometry data and gating strategy for sorted CD8+ and CD4+ naïve cells.
(a) Pure populations of naïve CD8+ cells (a) and CD4+ cells (b) were generated by cell sorting before processing for proteomics. Representative flow cytometry plots are shown.
Supplementary Figure 9 Representative flow cytometry data and gating strategy for sorted TCR activated CD8+ cells treated with rapamycin.
(a) Pure populations of 24 h TCR activated CD8+ cells without rapamycin (a) and with rapamycin treatment (b) were generated by cell sorting before processing for proteomics. Representative flow cytometry plots are shown.
Supplementary Figure 10 Representative flow cytometry data and gating strategy for sorted TCR activated CD4+ cells treated with rapamycin.
(a) Pure populations of 24 h TCR activated CD4+ cells without rapamycin (a) and with rapamycin treatment (b) were generated by cell sorting before processing for proteomics. Representative flow cytometry plots are shown.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10
Supplementary Data 1
Data used for generating heat maps.
Supplementary Data 2
Proteomics data for examining T cell differentiation.
Supplementary Data 3
Proteomics data for comparing CD8 and CD4 cells.
Supplementary Data 4
Proteomics data examining the impact of mTORC1 inhibition.
Supplementary Data 5
Protein categories used in this study.
Rights and permissions
About this article
Cite this article
Howden, A.J.M., Hukelmann, J.L., Brenes, A. et al. Quantitative analysis of T cell proteomes and environmental sensors during T cell differentiation. Nat Immunol 20, 1542–1554 (2019). https://doi.org/10.1038/s41590-019-0495-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0495-x
This article is cited by
-
Integrative temporal multi-omics reveals uncoupling of transcriptome and proteome during human T cell activation
npj Systems Biology and Applications (2024)
-
A Novel Homozygous Germline Mutation in Transferrin Receptor 1 (TfR1) Leads to Combined Immunodeficiency and Provides New Insights into Iron-Immunity Axis
Journal of Clinical Immunology (2024)
-
An integrated proteome and transcriptome of B cell maturation defines poised activation states of transitional and mature B cells
Nature Communications (2023)
-
Regulation of CD8+ T memory and exhaustion by the mTOR signals
Cellular & Molecular Immunology (2023)
-
CD47 masks pro-phagocytic ligands in cis on tumor cells to suppress antitumor immunity
Nature Immunology (2023)