Abstract
Natural killer (NK) cells can recognize virus-infected and stressed cells1 using activating and inhibitory receptors, many of which interact with HLA class I. Although early studies also suggested a functional impact of HLA class II on NK cell activity2,3, the NK cell receptors that specifically recognize HLA class II molecules have never been identified. We investigated whether two major families of NK cell receptors, killer-cell immunoglobulin-like receptors (KIRs) and natural cytotoxicity receptors (NCRs), contained receptors that bound to HLA class II, and identified a direct interaction between the NK cell receptor NKp44 and a subset of HLA-DP molecules, including HLA-DP401, one of the most frequent class II allotypes in white populations4. Using NKp44ζ+ reporter cells and primary human NKp44+ NK cells, we demonstrated that interactions between NKp44 and HLA-DP401 trigger functional NK cell responses. This interaction between a subset of HLA-DP molecules and NKp44 implicates HLA class II as a component of the innate immune response, much like HLA class I. It also provides a potential mechanism for the described associations between HLA-DP subtypes and several disease outcomes, including hepatitis B virus infection5,6,7, graft-versus-host disease8 and inflammatory bowel disease9,10.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All figures have associated raw data. All primary data files are available upon request from the corresponding author.
Code availability
All codes are available upon request from the corresponding author
References
Jost, S. & Altfeld, M. Control of human viral infections by natural killer cells. Annu. Rev. Immunol. 31, 163–194 (2013).
Jiang, Y. Z. et al. Interaction of natural killer cells with MHC class II: reversal of HLA-DR1-mediated protection of K562 transfectant from natural killer cell-mediated cytolysis by brefeldin-A. Immunology 87, 481–486 (1996).
Lobo, P. I., Chang, M. Y. & Mellins, E. Mechanisms by which HLA-class II molecules protect human B lymphoid tumour cells against NK- and LAK-mediated cytolysis. Immunology 88, 625–629 (1996).
al-Daccak, R. et al. Gene polymorphism of HLA-DPB1 and DPA1 loci in caucasoid population: frequencies and DPB1-DPA1 associations. Hum. Immunol. 31, 277–285 (1991).
Guo, X. et al. Strong influence of human leukocyte antigen (HLA)-DP gene variants on development of persistent chronic hepatitis B virus carriers in the Han Chinese population. Hepatology 53, 422–428 (2011).
Kamatani, Y. et al. A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat. Genet. 41, 591–595 (2009).
Thomas, R. et al. A novel variant marking HLA-DP expression levels predicts recovery from hepatitis B virus infection. J. Virol. 86, 6979–6985 (2012).
Petersdorf, E. W. et al. High HLA-DP expression and graft-versus-host disease. N. Engl. J. Med. 373, 599–609 (2015).
Hadley, D. et al. HLA-DPB1*04:01 protects genetically susceptible children from celiac disease autoimmunity in the TEDDY study. Am. J. Gastroenterol. 110, 915–920 (2015).
Goyette, P. et al. High-density mapping of the MHC identifies a shared role for HLA-DRB1*01:03 in inflammatory bowel diseases and heterozygous advantage in ulcerative colitis. Nat. Genet. 47, 172–179 (2015).
Garcia-Beltran, W. F. et al. Open conformers of HLA-F are high-affinity ligands of the activating NK-cell receptor KIR3DS1. Nat. Immunol. 17, 1067–1074 (2016).
Triebel, F. et al. LAG-3, a novel lymphocyte activation gene closely related to CD4. J. Exp. Med. 171, 1393–1405 (1990).
Berry, R. et al. Targeting of a natural killer cell receptor family by a viral immunoevasin. Nat. Immunol. 14, 699–705 (2013).
Deuss, F. A., Watson, G. M., Fu, Z., Rossjohn, J. & Berry, R. Structural basis for CD96 immune receptor recognition of nectin-like protein-5, CD155. Structure 27, 219–228.e3 (2019).
Rossjohn, J. et al. T cell antigen receptor recognition of antigen-presenting molecules. Annu. Rev. Immunol. 33, 169–200 (2015).
Vivian, J. P. et al. Killer cell immunoglobulin-like receptor 3DL1-mediated recognition of human leukocyte antigen B. Nature 479, 401–405 (2011).
Vitale, M. et al. NKp44, a novel triggering surface molecule specifically expressed by activated natural killer cells, is involved in non-major histocompatibility complex-restricted tumor cell lysis. J. Exp. Med. 187, 2065–2072 (1998).
Alter, G., Malenfant, J. M. & Altfeld, M. CD107a as a functional marker for the identification of natural killer cell activity. J. Immunol. Methods 294, 15–22 (2004).
Hiltbold, E. M. & Roche, P. A. Trafficking of MHC class II molecules in the late secretory pathway. Curr. Opin. Immunol. 14, 30–35 (2002).
Wen, F., Esteban, O. & Zhao, H. Rapid identification of CD4+ T-cell epitopes using yeast displaying pathogen-derived peptide library. J. Immunol. Methods 336, 37–44 (2008).
Birnbaum, M. E. et al. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157, 1073–1087 (2014).
Lo, W.-L. et al. An endogenous peptide positively selects and augments the activation and survival of peripheral CD4+ T cells. Nat. Immunol. 10, 1155–1161 (2009).
Dai, S. et al. Crystal structure of HLA-DP2 and implications for chronic beryllium disease. Proc. Natl Acad. Sci. USA 107, 7425–7430 (2010).
Stevanovic, S. et al. HLA class II upregulation during viral infection leads to HLA-DP-directed graft-versus-host disease after CD4+ donor lymphocyte infusion. Blood 122, 1963–1973 (2013).
Holzemer, A. et al. Selection of an HLA-C*03:04-restricted HIV-1 p24 Gag sequence variant is associated with viral escape from KIR2DL3+ natural killer cells: data from an observational cohort in South Africa. PLoS Med. 12, e1001900 (2015).
Naiyer, M. M. et al. KIR2DS2 recognizes conserved peptides derived from viral helicases in the context of HLA-C. Sci. Immunol. 2, eaal5296 (2017).
O’Connor, G. M. et al. Peptide-dependent recognition of HLA-B*57:01 by KIR3DS1. J. Virol. 89, 5213–5221 (2015).
O’Connor, G. M. et al. Mutational and structural analysis of KIR3DL1 reveals a lineage-defining allotypic dimorphism that impacts both HLA and peptide sensitivity. J. Immunol. 192, 2875–2884 (2014).
Chapel, A. et al. Peptide-specific engagement of the activating NK cell receptor KIR2DS1. Sci. Rep. 7, 2414 (2017).
Rajagopalan, S. & Long, E. O. The direct binding of a p58 killer cell inhibitory receptor to human histocompatibility leukocyte antigen (HLA)-Cw4 exhibits peptide selectivity. J. Exp. Med. 185, 1523–1528 (1997).
Holling, T. M., Schooten, E., Langerak, A. W. & van den Elsen, P. J. Regulation of MHC class II expression in human T-cell malignancies. Blood 103, 1438–1444 (2004).
Thompson, J. A. et al. Tumor cells transduced with the MHC class II transactivator and CD80 activate tumor-specific CD4+ T cells whether or not they are silenced for invariant chain. Cancer Res. 66, 1147–1154 (2006).
Takayama, T. et al. Imbalance of NKp44+NKp46− and NKp44−NKp46+ natural killer cells in the intestinal mucosa of patients with Crohn’s disease. Gastroenterology 139, 882–892.e3 (2010).
Glatzer, T. et al. RORgammat+ innate lymphoid cells acquire a proinflammatory program upon engagement of the activating receptor NKp44. Immunity 38, 1223–1235 (2013).
Campbell, K. S., Yusa, S., Kikuchi-Maki, A. & Catina, T. L. NKp44 triggers NK cell activation through DAP12 association that is not influenced by a putative cytoplasmic inhibitory sequence. J. Immunol. 172, 899–906 (2004).
Cantoni, C. et al. NKp44, a triggering receptor involved in tumor cell lysis by activated human natural killer cells, is a novel member of the immunoglobulin superfamily. J. Exp. Med. 189, 787–796 (1999).
Siewiera, J. et al. Natural cytotoxicity receptor splice variants orchestrate the distinct functions of human natural killer cell subtypes. Nat. Commun. 6, 10183 (2015).
Arnon, T. I. et al. Recognition of viral hemagglutinins by NKp44 but not by NKp30. Eur. J. Immunol. 31, 2680–2689 (2001).
Ho, J. W. et al. H5-type influenza virus hemagglutinin is functionally recognized by the natural killer-activating receptor NKp44. J. Virol. 82, 2028–2032 (2008).
Rosental, B. et al. Proliferating cell nuclear antigen is a novel inhibitory ligand for the natural cytotoxicity receptor NKp44. J. Immunol. 187, 5693–5702 (2011).
Baychelier, F. et al. Identification of a cellular ligand for the natural cytotoxicity receptor NKp44. Blood 122, 2935–2942 (2013).
Barrow, A. D. et al. Natural killer cells control tumor growth by sensing a growth factor. Cell 172, 534–548.e19 (2018).
De Maria, A. et al. NKp44 expression, phylogenesis and function in non-human primate NK cells. Int. Immunol. 21, 245–255 (2009).
Ting, J. P.-Y. & Trowsdale, J. Genetic control of MHC class II expression. Cell 109, S21–S33 (2002).
Allcock, R. J. N., Barrow, A. D., Forbes, S., Beck, S. & Trowsdale, J. The human TREM gene cluster at 6p21.1 encodes both activating and inhibitory single IgV domain receptors and includes NKp44. Eur. J. Immunol. 33, 567–577 (2003).
Slierendregt, B. L., Otting, N., Kenter, M. & Bontrop, R. E. Allelic diversity at the Mhc-DP locus in rhesus macaques (Macaca mulatta). Immunogenetics 41, 29–37 (1995).
Nizetic, D., Figueroa, F., Dembic, Z., Nevo, E. & Klein, J. Major histocompatibility complex gene organization in the mole rat Spalax ehrenbergi: evidence for transfer of function between class II genes. Proc. Natl Acad. Sci. USA 84, 5828–5832 (1987).
Vilches, C. & Parham, P. KIR: diverse, rapidly evolving receptors of innate and adaptive immunity. Annu. Rev. Immunol. 20, 217–251 (2002).
Petersen, J. et al. T-cell receptor recognition of HLA-DQ2-gliadin complexes associated with celiac disease. Nat. Struct. Mol. Biol. 21, 480–488 (2014).
Henderson, K. N. et al. A structural and immunological basis for the role of human leukocyte antigen DQ8 in celiac disease. Immunity 27, 23–34 (2007).
Acknowledgements
This work has been funded in part by the Pathogenesis and the Viral Latency Programs of the Heinrich Pette Institute, Leibniz Institute for Experimental Virology and the German Center for Infection Research (DZIF) through TTU 04.810. This project has been funded in part with federal funds from the Frederick National Laboratory for Cancer Research, under contract no. HHSN261200800001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government. This research was supported in part by the Intramural Research Program of the National Institutes of Health, Frederick National Laboratory, Center for Cancer Research. W.F.G.-B. was supported by National Institute of General Medical Sciences (T32GM007752) and the NIH (P01-AI104715 and F31AI116366). P.J.N. was supported by NIH U19 NS095774. A.H. was supported by the German Center for Infection Research (DZIF) through an MD/PhD Stipend (TI 07.002) and via the Clinician Scientist Program of the Faculty of Medicine, University Medical Center Hamburg-Eppendorf, Hamburg, Germany. J.R was supported by an Australian Research Council Laureate Fellowship (FL160100049) and R.B was supported by a Career Development Fellowship from the National Health and Medical Research Council of Australia (APP1109901). We would like to thank H. Reid and K. Loh for their kind gift of HLA-DQ2 viral stocks. We would like to thank the NIH Tetramer Core Facility for all provided HLA class II monomers.
Author information
Authors and Affiliations
Contributions
A.N. performed reporter cell and primary NK cell experiments and analyzed the data. A.C. and A.N. conducted and analyzed the HLA class II-coated bead assay. A.N., W.F.G.-B. and A.H. designed and generated the Jurkat reporter cell lines. G.M.W., R.B. and J.R. conducted the SPR measurements. P.J.N., M.O. and J.-H.L. provided the single HLA-DP antigens and gave important intellectual input. L.R. performed mixed effects linear regression models and provided important statistical guidance. A.N., W.F.G.-B. and M.A. designed the experiments. G.M., C.K., A.P.-R. and M.C. gave important intellectual input throughout the process. M.A. supervised the study. A.N. wrote the first draft of the manuscript and M.A. revised and edited the manuscript. All authors revised the manuscript and approved it for publication.
Corresponding author
Ethics declarations
Competing interests
A.N., W.F.G.-B. and M.A. filed a patent application (EP18174760.1) regarding the therapeutic use of anti-NKp44 antibodies for the treatment and/or prevention of graft-versus-host disease. M.O. and J.-H.L. are current employees of OneLambda Inc., a part of Thermo Fisher Scientific. All other authors declare no competing interest.
Additional information
Peer review information: Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 KIR3DL1 and KIR2DL4 Fc constructs exhibit unspecific binding to HLA class II coated beads at high concentrations.
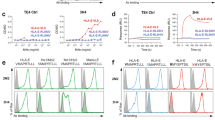
a Binding of KIR3DL1 Fc construct at various concentrations (1–100 µg/mL) to HLA-DR (blue), HLA-DQ (yellow) and HLA-DP (red) coated beads as well as positive (grey) and negative (black) control beads is plotted as median fluorescence intensity (MFI). Each dot represents the measured MFI for binding of the KIR3DL1 Fc construct at a specific concentration to a certain HLA class II coated bead or negative/positive control beads. Lines connect matching MFI values for one specific HLA class II allotype or negative/positive control beads measured at different KIR3DL1 Fc construct concentrations. Data is representative for one single experiment (n = 1). b MFI values of KIR2DL4 Fc construct binding at varying concentrations (1–100 µg/mL; left panel) and at 10 µg/mL to HLA-DR (blue), HLA-DQ (yellow) and HLA-DP (red) coated beads as well as positive (grey) and negative (black) control beads (right panel) are depicted. Each dot in both panels represents one HLA class II molecule. Lines in the left panel connect matching MFI values for one specific HLA class II allotype or negative/positive control beads measured at different KIR2DL4 Fc construct concentrations. Horizontal line in right panel indicates median, error bars indicate interquartile range. Data is representative for one single experiment (n = 1).
Supplementary Figure 2 Untransduced Jurkat cells do not show functional responses to HLA class II molecules.
Activation of untransduced Jurkat cells in response to anti-KIR2DL3, anti-NKp46, anti-NKp44 as well as HLA-DR7 and HLA-DP401 CLIP monomers was assessed by the expression of CD69. Plot represents one of eight independent experiments (left panel). The percentage of CD69+ cells following incubation on non-coated wells (blank) was subtracted from all samples. Corrected values are illustrated as median with interquartile range as determined in eight independent biological replicates (n = 8).
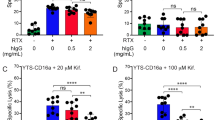
Supplementary Figure 3 Unstimulated NK cells do not express NKp44 and do not degranulate upon co-incubation with HLA-DP401.
a Surface expression of NKp44 was determined in freshly isolated untreated (black) and IL-2 plus IL-15 treated (red) primary NK cells. Plot represents one of seven individual donors. MFI of NKp44 expression from untreated and cytokine-treated primary NK cells was determined in seven individual donors (n = 7). Each dot represents one donor. Horizontal line indicates the median, error bars display interquartile range. Two-tailed Wilcoxon matched-pairs signed rank test was used to assess differences in the surface expression of NKp44 between untreated and cytokine-treated primary NK cells. *p = 0.02. b Freshly isolated unstimulated NK cells were isolated from seven individual donors and co-incubated with plate-coated anti-NKp44, HLA-DR7 CLIP, HLA-DP401 CLIP or non-coated wells (blank) in the presence of purified mouse IgG1 isotype or purified anti-human NKp44 antibody (both at a final concentration of 10 µg/mL). The percentage of CD107a+ cells was determined. Each dot represents one individual donors (n = 7) and lines connect responses from one individual donor.
Supplementary Figure 4 HLA-DP surface expression of JE6.1-DP transduced cell lines is increased by CLIP pulsing.
a HLA-DP surface expression of HLA-DP transduced cell lines (red and blue histograms) is depicted. The HLA-DP expression of the untransduced parental cell line is displayed in grey. Plots represent one of seven individual experiments. b HLA-DP (left panel) and CLIP (right panel) surface expression following CLIP pulsing in comparison to DMSO-treated cells is depicted as fold change in MFI [MFI CLIP pulsed/MFI DMSO] for the four indicated JE6.1-DP expressing cell lines. Each dot represents one individual biological replicate as determined in seven independent experiments (n = 7). Boxes represent 25th to 75th percentiles. Whiskers indicate minimum and maximum values, horizontal line indicates the median.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and Supplementary Tables 1 and 2
Rights and permissions
About this article
Cite this article
Niehrs, A., Garcia-Beltran, W.F., Norman, P.J. et al. A subset of HLA-DP molecules serve as ligands for the natural cytotoxicity receptor NKp44. Nat Immunol 20, 1129–1137 (2019). https://doi.org/10.1038/s41590-019-0448-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0448-4
This article is cited by
-
Advances of medical nanorobots for future cancer treatments
Journal of Hematology & Oncology (2023)
-
Roles of natural killer cells in immunity to cancer, and applications to immunotherapy
Nature Reviews Immunology (2023)
-
The impact of HLA polymorphism on herpesvirus infection and disease
Immunogenetics (2023)
-
The role of natural killer cells in liver inflammation
Seminars in Immunopathology (2021)
-
Variation and expression of HLA-DPB1 gene in HBV infection
Immunogenetics (2021)