Abstract
Translation termination is an essential cellular process, which is also of therapeutic interest for diseases that manifest from premature stop codons. In eukaryotes, translation termination requires eRF1, which recognizes stop codons, catalyzes the release of nascent proteins from ribosomes and facilitates ribosome recycling. The small molecule SRI-41315 triggers eRF1 degradation and enhances translational readthrough of premature stop codons. However, the mechanism of action of SRI-41315 on eRF1 and translation is not known. Here we report cryo-EM structures showing that SRI-41315 acts as a metal-dependent molecular glue between the N domain of eRF1 responsible for stop codon recognition and the ribosomal subunit interface near the decoding center. Retention of eRF1 on ribosomes by SRI-41315 leads to ribosome collisions, eRF1 ubiquitylation and a higher frequency of translation termination at near-cognate stop codons. Our findings reveal a new mechanism of release factor inhibition and additional implications for pharmacologically targeting eRF1.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
EM maps and models are available under accession numbers PDB 8SCB, EMD-40344 and EMD-40345. All other data supporting the findings of this study are within the article and its Extended Data. Raw data files needed in other formats are available from the corresponding author upon reasonable request. Additional data used in this study are PDB 6SGC, 3JAG, 6MTE and 6XA1. Source data are provided with this paper.
References
Hellen, C. U. T. Translation termination and ribosome recycling in eukaryotes. Cold Spring Harb. Perspect. Biol. 10, a032656 (2018).
Frolova, L. et al. A highly conserved eukaryotic protein family possessing properties of polypeptide chain release factor. Nature 372, 701–703 (1994).
Brown, A., Shao, S., Murray, J., Hegde, R. S. & Ramakrishnan, V. Structural basis for stop codon recognition in eukaryotes. Nature 524, 493–496 (2015).
Matheisl, S., Berninghausen, O., Becker, T. & Beckmann, R. Structure of a human translation termination complex. Nucleic Acids Res. 43, 8615–8626 (2015).
Shao, S. et al. Decoding mammalian ribosome–mRNA states by translational GTPase complexes. Cell 167, 1229–1240.e15 (2016).
Frolova, L. Y. et al. Mutations in the highly conserved GGQ motif of class 1 polypeptide release factors abolish ability of human eRF1 to trigger peptidyl-tRNA hydrolysis. RNA 5, 1014–1020 (1999).
Pisarev, A. V. et al. The role of ABCE1 in eukaryotic posttermination ribosomal recycling. Mol. Cell 37, 196–210 (2010).
Shoemaker, C. J. & Green, R. Kinetic analysis reveals the ordered coupling of translation termination and ribosome recycling in yeast. Proc. Natl Acad. Sci. USA 108, E1392–E1398 (2011).
Kurosaki, T., Popp, M. W. & Maquat, L. E. Quality and quantity control of gene expression by nonsense-mediated mRNA decay. Nat. Rev. Mol. Cell Biol. 20, 406–420 (2019).
Karousis, E. D. & Mühlemann, O. The broader sense of nonsense. Trends Biochem. Sci. 47, 921–935 (2022).
Morais, P., Adachi, H. & Yu, Y.-T. Suppression of nonsense mutations by new emerging technologies. Int. J. Mol. Sci. 21, 4394 (2020).
Albers, S. et al. Engineered tRNAs suppress nonsense mutations in cells and in vivo. Nature 618, 842–848 (2023).
Sharma, J. et al. A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion. Nat. Commun. 12, 4358 (2021).
Gurzeler, L.-A. et al. Drug-induced eRF1 degradation promotes readthrough and reveals a new branch of ribosome quality control. Cell Rep. 42, 113056 (2023).
Jacoby, E., Reinhardt, J., Schmiedeberg, N. & Spanka, C. Pyrimido [4,5-B]quinoline- 4,5 (3H,10H)-diones as nonsense mutation suppressors. Patent WO/2014/091446 A1 (2014).
Oltion, K. et al. An E3 ligase network engages GCN1 to promote the degradation of translation factors on stalled ribosomes. Cell 186, 346–362.e17 (2023).
Juette, M. F. et al. Didemnin B and ternatin-4 differentially inhibit conformational changes in eEF1A required for aminoacyl-tRNA accommodation into mammalian ribosomes. eLife 11, e81608 (2022).
Li, W., Chang, S. T.-L., Ward, F. R. & Cate, J. H. D. Selective inhibition of human translation termination by a drug-like compound. Nat. Commun. 11, 4941 (2020).
Florin, T. et al. An antimicrobial peptide that inhibits translation by trapping release factors on the ribosome. Nat. Struct. Mol. Biol. 24, 752–757 (2017).
Koller, T. O. et al. Structural basis for translation inhibition by the glycosylated drosocin peptide. Nat. Chem. Biol. 19, 1072–1081 (2023).
Feng, Q. & Shao, S. In vitro reconstitution of translational arrest pathways. Methods 137, 20–36 (2018).
Shao, S., von der Malsburg, K. & Hegde, R. S. Listerin-dependent nascent protein ubiquitination relies on ribosome subunit dissociation. Mol. Cell 50, 637–648 (2013).
Juszkiewicz, S. et al. Ribosome collisions trigger cis-acting feedback inhibition of translation initiation. eLife 9, e60038 (2020).
Sinha, N. K. et al. EDF1 coordinates cellular responses to ribosome collisions. eLife 9, e58828 (2020).
Kolosov, P. et al. Invariant amino acids essential for decoding function of polypeptide release factor eRF1. Nucleic Acids Res. 33, 6418–6425 (2005).
Bulygin, K. N. et al. Three distinct peptides from the N domain of translation termination factor eRF1 surround stop codon in the ribosome. RNA 16, 1902–1914 (2010).
Seit-Nebi, A., Frolova, L. & Kisselev, L. Conversion of omnipotent translation termination factor eRF1 into ciliate-like UGA-only unipotent eRF1. EMBO Rep. 3, 881–886 (2002).
Manuvakhova, M., Keeling, K. & Bedwell, D. M. Aminoglycoside antibiotics mediate context-dependent suppression of termination codons in a mammalian translation system. RNA 6, 1044–1055 (2000).
Namy, O., Hatin, I. & Rousset, J. Impact of the six nucleotides downstream of the stop codon on translation termination. EMBO Rep. 2, 787–793 (2001).
Cridge, A. G., Crowe-McAuliffe, C., Mathew, S. F. & Tate, W. P. Eukaryotic translational termination efficiency is influenced by the 3′ nucleotides within the ribosomal mRNA channel. Nucleic Acids Res. 46, 1927–1944 (2018).
Cao, S. et al. Defining molecular glues with a dual-nanobody cannabidiol sensor. Nat. Commun. 13, 815 (2022).
Faust, T. B., Donovan, K. A., Yue, H., Chamberlain, P. P. & Fischer, E. S. Small-molecule approaches to targeted protein degradation. Annu. Rev. Cancer Biol. 5, 181–201 (2020).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Holm, M. et al. mRNA decoding in human is kinetically and structurally distinct from bacteria. Nature 617, 200–207 (2023).
Paternoga, H. et al. Structural conservation of antibiotic interaction with ribosomes. Nat. Struct. Mol. Biol. 30, 1380–1392 (2023).
Lin, J., Zhou, D., Steitz, T. A., Polikanov, Y. S. & Gagnon, M. G. Ribosome-targeting antibiotics: modes of action, mechanisms of resistance, and implications for drug design. Annu. Rev. Biochem 87, 451–478 (2018).
Li, W. et al. Structural basis for selective stalling of human ribosome nascent chain complexes by a drug-like molecule. Nat. Struct. Mol. Biol. 26, 501–509 (2019).
Zavialov, A. V., Mora, L., Buckingham, R. H. & Ehrenberg, M. Release of peptide promoted by the GGQ motif of class 1 release factors regulates the GTPase activity of RF3. Mol. Cell 10, 789–798 (2002).
Korostelev, A. et al. Crystal structure of a translation termination complex formed with release factor RF2. Proc. Natl Acad. Sci. USA 105, 19684–19689 (2008).
Laurberg, M. et al. Structural basis for translation termination on the 70S ribosome. Nature 454, 852–857 (2008).
Weixlbaumer, A. et al. Insights into translational termination from the structure of RF2 bound to the ribosome. Science 322, 953–956 (2008).
Scolnick, E., Tompkins, R., Caskey, T. & Nirenberg, M. Release factors differing in specificity for terminator codons. Proc. Natl Acad. Sci. USA 61, 768–774 (1968).
Yip, M. C. J. & Shao, S. Detecting and rescuing stalled ribosomes. Trends Biochem. Sci. 46, 731–743 (2021).
Juszkiewicz, S. et al. ZNF598 is a quality control sensor of collided ribosomes. Mol. Cell 72, 469–481.e7 (2018).
Wu, C. C.-C., Peterson, A., Zinshteyn, B., Regot, S. & Green, R. Ribosome collisions trigger general stress responses to regulate cell fate. Cell 182, 404–416.e14 (2020).
Garshott, D. M. et al. iRQC, a surveillance pathway for 40S ribosomal quality control during mRNA translation initiation. Cell Rep. 36, 109642 (2021).
Garzia, A., Meyer, C. & Tuschl, T. The E3 ubiquitin ligase RNF10 modifies 40S ribosomal subunits of ribosomes compromised in translation. Cell Rep. 36, 109468 (2021).
Ikeuchi, K. et al. Collided ribosomes form a unique structural interface to induce Hel2‐driven quality control pathways. EMBO J. 38, e100276 (2019).
Juszkiewicz, S. & Hegde, R. S. Initiation of quality control during poly(A) translation requires site-specific ribosome ubiquitination. Mol. Cell 65, 743–750.e4 (2017).
Sugiyama, T. et al. Sequential ubiquitination of ribosomal protein uS3 triggers the degradation of non-functional 18S rRNA. Cell Rep. 26, 3400–3415.e7 (2019).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Chandrasekaran, V. et al. Mechanism of ribosome stalling during translation of a poly(A) tail. Nat. Struct. Mol. Biol. 26, 1132–1140 (2019).
Brown, A., Baird, M. R., Yip, M. C., Murray, J. & Shao, S. Structures of translationally inactive mammalian ribosomes. eLife 7, e40486 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Taoka, M. et al. Landscape of the complete RNA chemical modifications in the human 80S ribosome. Nucleic Acids Res. 46, 9289–9298 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Acknowledgements
Cryo-EM screening and data collection were performed at the Harvard Center for Cryo-Electron Microscopy (HC2EM). Data processing was supported by SBGrid. We thank A. Brown, Y. Peng and Shao laboratory members for helpful discussions. This work was supported by the Packard Foundation (S.S.), NIH DP2GM137415 (S.S.) and the UCSF Invent Fund (J.T.). M.C.J.Y. was supported by the American Heart Association (predoctoral fellowship 287375208). K.O. was supported by a UCSF Genentech Fellowship.
Author information
Authors and Affiliations
Contributions
J.P.L.C., M.C.J.Y. and S.S. performed biochemical analyses. K.O. performed flow cytometry experiments. M.C.J.Y. collected and processed cryo-EM data. S.S. and J.T. conceived the project. S.S. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
J.T. is a founder of Kezar Life Sciences and Terremoto Biosciences and is a scientific advisor to Entos. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Yury Polikanov, Daniel Wilson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of SRI-41315 in an in vitro translation system.
a, Effect of SRI-41315 on protein synthesis. Representative titrations of SRI-41315 into in vitro translation reactions of a radiolabeled 3xFlag-tagged model protein ending with the indicated stop codon or a nonstop (NS) control, analyzed by SDS-PAGE and autoradiography on film (top) or by phosphorimaging (bottom). NC, expected nascent protein chain; NC-tRNA, nonhydrolyzed peptidyl-tRNA adduct. Orange dots, smaller products specifically observed with SRI-41315, representative of 5 replicates with similar results. b, SRI-41315 does not trap nascent proteins on ribosomes. In vitro translation reactions as in panel a without or with 0.5 µM eRF1(AAQ) and/or 100 µM SRI-41315 added after 5 min were size fractionated on 10-50% sucrose gradients. The total (T) and eleven fractions collected from the top of each gradient were analyzed by SDS-PAGE and autoradiography. Note: eRF1(AAQ) but not SRI-41315 retains NC-tRNAs and NCs hydrolyzed from tRNAs during SDS-PAGE in ribosomal fractions, representative of 3 replicates with similar results.
Extended Data Fig. 2 SRI-41315 traps ubiquitylated eRF1 on ribosomes.
a, RNF14 levels in different mammalian lysates. Two-fold dilutions of rabbit reticulocyte lysate (RRL) or HEK293T cell lysate analyzed by SDS-PAGE and immunoblotting. Note: low levels of RNF14 in RRL relative to GCN1 and ribosomal proteins, representative of 2 replicates with similar results. b, RNF14 mediates eRF1 ubiquitylation with SRI-41315. In vitro translation reactions containing 10 µM His-tagged ubiquitin without or with 50 nM wildtype (WT) or catalytically inactive (C417A) recombinant RNF14 (rRNF14) and 100 µM SRI-41315 were analyzed directly (total) or after denaturing pulldowns of His-tagged ubiquitin (His-Ub PD) by SDS-PAGE and immunoblotting. Note: WT but not C417A rRNF14 enhances SRI-41315-dependent ubiquitylation of eRF1, representative of 3 replicates with similar results. Residual eRF1 ubiquitylation is likely due to endogenous RNF14 in RRL. c, Titration of SRI-41315 into in vitro translation reactions containing 50 nM recombinant RNF14 analyzed directly (total) or after His-Ub PD by SDS-PAGE and immunoblotting, representative of 3 replicates with similar results. d, eRF1 ubiquitylation is slower than translation. Timecourses assaying eRF1 ubiquitylation in the presence of 50 nM recombinant RNF14, 10 µM His-tagged ubiquitin, and 25 µM SRI-41315 (top) compared to timecourses of radiolabeled nascent protein (NC) synthesis (bottom) in in vitro translation reactions, representative of 2 replicates with similar results. e, SRI-41315 traps eRF1 on ribosomes. Translation reactions as in Fig. 1c were size fractionated over 10-50% sucrose gradients. A total (T) sample and eleven fractions collected from the top of each gradient were analyzed directly or after His-Ub PD by SDS-PAGE and immunoblotting for eRF1. Note: SRI-41315 retains both unmodified and ubiquitylated eRF1 in ribosomal fractions, representative of 3 replicates with similar results.
Extended Data Fig. 3 SRI-41315 recruits collision sensors to ribosomes.
a,b, Effects of SRI-41315 in cells. Lysates of Flp-In 293 T-REx cells treated without or with 30 µM SRI-41315, 1.8 µM emetine, a concentration of the translation elongation inhibitor demonstrated to cause ribosome collisions, and/or 1 µM MLN-7243, an inhibitor of the E1 ubiquitin-activating enzyme, for 2 hr were analyzed by SDS-PAGE and immunoblotting a, directly or b, after size fractionation on sucrose gradients, representative of 2 replicates with similar results. Note: ubiquitylated eRF1 (Ub-eRF1) is detected specifically with SRI-41315 and suppressed by MLN-7243. SRI-41315 and the low dose of emetine both lead to the recruitment of the ribosome collision sensor EDF1 to ribosomal fractions. c, SRI-41315 induces the recruitment of EDF1 to ribosomes in vitro. SDS-PAGE and immunoblotting for the ribosome collision sensor EDF1 in total, soluble, or ribosomal fractions from translation reactions containing the indicated concentrations of SRI-41315, representative of 2 replicates with similar results. d, SRI-41315 induces eRF1 degradation in Flp-In 293 T-REx cells after 20 hr, representative of 3 replicates with similar results.
Extended Data Fig. 4 Cryo-EM data processing.
a, Ribosome-nascent protein complexes (RNCs) from translation reactions containing 0.5 µM eRF1(AAQ) and 100 µM SRI-41315 added at 5 min were affinity purified via the 3xFlag tag encoded in the nascent chain (NC) and analyzed by SDS-PAGE and Coomassie staining (top) or immunoblotting (bottom). NC-tRNA, nonhydrolyzed peptidyl-tRNA adduct. b, Representative cryo-EM micrograph of RNCs from panel a. c, Summary of cryo-EM data processing and classification strategy.
Extended Data Fig. 5 Quality of cryo-EM maps and model.
a, Fourier shell correlation (FSC) vs. resolution (1/Å) curves for the indicated cryo-EM maps. b, The indicated cryo-EM map colored by local resolution. c, Model vs. map FSC curves. d, Density of the defined nascent protein sequence in the ribosomal exit tunnel in the sharpened cryo-EM map contoured at 2.8σ. e, The cryo-EM map of the SRI-41315 binding site as in Fig. 2b colored by local resolution (left) or by entity (right). Magnesium ions are in green. f, Coordination of the magnesium ion at the SRI-41315 binding site compared to density in the sharpened cryo-EM map contoured at 4.0σ.
Extended Data Fig. 6 eRF1 secondary structure and alignments.
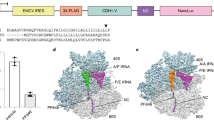
Sequence alignments of eRF1 from the indicated species (H. sapiens UniProt P62495; O. cuniculus UniProt P62497; M. musculus UniProt Q8BWY3; X. laevis UniProt P35615; D. rerio UniProt Q803E5; D. melanogaster UniProt Q9VPH7; C. elegans UniProt O16520; S. cerevisiae UniProt P12385) with secondary structure designations above colored based on their presence in the N (blue), M (green), and C (orange) domains of eRF1. In the accommodated state of eRF1, α8 of the M domain and α9 of the C domain form a continuous helix (yellow). eRF1 also contains a minidomain insertion (gray) in the C domain that is not present in structurally similar decoding factors. In the N domain, Met51 (purple arrowhead) involved in SRI-41315 binding, as well as the NIKS and YxCxxxF motifs (gray lines) and Glu55 (gray arrowhead) involved in stop codon recognition, are indicated below. Note: human, rabbit, and mouse eRF1 are 100% identical. Part of the C-terminal extension in C. elegans eRF1 is not shown.
Extended Data Fig. 7 Structural comparisons.
a, Validation of SRI-41315 density. The model of the SRI-41315 binding site as in Fig. 2b docked into cryo-EM maps of the rabbit ribosome bound to eRF1(AAQ) without SRI-41315 (EMD-3038; left), of the human ribosome bound to eRF1 trapped by PF846 (EMD-22085; middle), or generated from particles selected for occupancy of the recycling factor ABCE1 (right), contoured at the indicated levels. Note: the left and middle maps lack SRI-41315, while the right map retains strong density corresponding to SRI-41315 coexisting with ABCE1 binding. b, The overall conformation of accommodated eRF1 is unchanged with SRI-41315. The model of eRF1(AAQ) bound to SRI-41315 (purple) aligned to eRF1(AAQ) without SRI-41315 (pink; PDB 3JAG) or eRF1 trapped on ribosomes by PF846 (blue; 6XA1). The N, M, and C domains (left), the GGQ motif, and the SRI-41315 binding site (right) are indicated. c, Docking of related small molecule eRF1 degraders in the SRI-41315 binding site. d, SRI-41315 binding region of eRF1 (PDB 1DT9) colored by conservation. Note: Met51 is less conserved than Tyr125 and other residues required for stop codon decoding. e, SRI-41315 does not change ABCE1 conformation. Alignment of ABCE1 on termination complexes without (pink; PDB 3JAG) or with (dark blue) SRI-41315. Iron-sulfur clusters are colored by heteroatoms. f, ABCE1 is slightly stabilized on ribosomes with SRI-41315. Total, soluble, and ribosomal fractions of translation reactions without or with 100 µM SRI-41315 as in Fig. 1c were analyzed by SDS-PAGE and immunoblotting for ABCE1, representative of 3 replicates with similar results.
Extended Data Fig. 8 Flow cytometry analysis.
a, Cells were initially gated by FSC-A vs. SSC-A to exclude debris. b, Cells from the gate in panel a were gated by FSC-A vs. FSC-W to exclude doublets. c, Cells from the gate in panel b were gated by GFP positivity (left) as judged by comparison to an untransduced reference (right) to exclude untransduced cells from analysis.
Extended Data Fig. 9 Cryptic stop codon analysis.
a, Transcript (top line) and polypeptide (bottom line) sequence of the reporter designed to test translation termination at cryptic stop codons. The reporter contains an N-terminal 8xHis tag (blue), a modified calmodulin sequence (light orange), the test codon position (dark orange; stop1), a modified sequence encoding the autonomously folding villin headpiece (VHP) domain followed by the cytosolic domain of Sec61β (light green), and the UAA stop codon (black; stop2). The reporter sequence was modified to remove all codons, except for one UGG codon (pink), that start with a U and contain a purine (A/G) in the second and/or third positions. Additional cryptic codons that start with a C and contain purines in the second and third positions (purple) were tested by mutations (see panel c). b, SRI-41315 induces translation termination at specific cryptic stop codons. Quantification of the ratio of stop1 to stop2 products from reporters containing UUA (green) or UAU (orange) in the test codon position synthesized in vitro with increasing concentrations of SRI-41315. The normalized average (line) and individual ratios (dots) from three independent experiments are shown. c, Mutagenesis of additional cryptic stop codons. Assays as in Fig. 4a with UGA (lane 1) or UGG (lanes 2–5) in the test codon position and additional mutation of CAG and CAA codons (Qmut; purple) upstream of the stop1 position and/or of the UGG codon (Wmut; pink) downstream of the stop1 position as indicated in panel a. Radiolabeled reporter products generated without (left) or with (right) 100 µM SRI-41315 are shown. Note: changes in products upon mutation of these codons occur specifically with SRI-41315, representative of 3 replicates with similar results. Purple dots denote products abolished by mutating the CAG/CAA codons; pink dot denotes the lower band of a doublet abolished by mutating the UGG codon. Yellow dot denotes a band with increased intensity after mutation of the CAG/CAA codons.
Supplementary information
Supplementary Information
Supplementary Table 1.
Source data
Source Data Fig. 1
Unprocessed images of stained gels, western blots, autoradiography and phosphorimaging.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed autoradiography image.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed images of autoradiography and phosphorimaging.
Source Data Extended Data Fig. 2
Unprocessed images of stained gels, western blots, autoradiography and phosphorimaging.
Source Data Extended Data Fig. 3
Unprocessed images of Ponceau stained blots and western blots.
Source Data Extended Data Fig. 4
Unprocessed images of stained gels and western blots.
Source Data Extended Data Fig. 7
Unprocessed image of western blot.
Source Data Extended Data Fig. 9
Unprocessed images of autoradiography.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Coelho, J.P.L., Yip, M.C.J., Oltion, K. et al. The eRF1 degrader SRI-41315 acts as a molecular glue at the ribosomal decoding center. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-023-01521-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-023-01521-0