Abstract
Hematopoietic stem cells (HSCs) regenerate after myeloablation, a procedure that adversely disrupts the bone marrow and drives leptin receptor-expressing cells, a key niche component, to differentiate extensively into adipocytes. Regeneration of the bone marrow niche is associated with the resolution of adipocytes, but the mechanisms remain poorly understood. Using Plin1-creER knock-in mice, we followed the fate of adipocytes in the regenerating niche in vivo. We found that bone marrow adipocytes were highly dynamic and dedifferentiated to leptin receptor-expressing cells during regeneration after myeloablation. Bone marrow adipocytes could give rise to osteolineage cells after skeletal injury. The cellular fate of steady-state bone marrow adipocytes was also plastic. Deletion of adipose triglyceride lipase (Atgl) from bone marrow stromal cells, including adipocytes, obstructed adipocyte dedifferentiation and led to severely compromised regeneration of HSCs as well as impaired B lymphopoiesis after myeloablation, but not in the steady state. Thus, the regeneration of HSCs and their niche depends on the cellular plasticity of bone marrow adipocytes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The snRNA-seq data generated by this study have been deposited in the NCBI Gene Expression Omnibus (GEO) repository under accession number GSE227255. Source data are provided with this paper.
Code availability
Standard bioinformatics pipelines used for analyzing snRNA-seq data were described in the ‘snRNA-seq’ section. All codes that have been used in the study are available at https://doi.org/10.5281/zenodo.8280830 (ref. 52) and https://github.com/LeiDingLab/Nature_Genetics_2023.
References
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Ding, L. & Morrison, S. J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
Lee, Y., Decker, M., Lee, H. & Ding, L. Extrinsic regulation of hematopoietic stem cells in development, homeostasis and diseases. Wiley Interdiscip. Rev. Dev. Biol. 6, https://doi.org/10.1002/wdev.279 (2017).
Comazzetto, S., Shen, B. & Morrison, S. J. Niches that regulate stem cells and hematopoiesis in adult bone marrow. Dev. Cell 56, 1848–1860 (2021).
Sugiyama, T., Kohara, H., Noda, M. & Nagasawa, T. Maintenance of the hematopoietic stem cell pool by CXCL12–CXCR4 chemokine signaling in bone marrow stromal cell niches. Immunity 25, 977–988 (2006).
Greenbaum, A. et al. CXCL12 in early mesenchymal progenitors is required for haematopoietic stem-cell maintenance. Nature 495, 227–230 (2013).
Cordeiro Gomes, A. et al. Hematopoietic stem cell niches produce lineage-instructive signals to control multipotent progenitor differentiation. Immunity 45, 1219–1231 (2016).
Zhou, B. O., Yue, R., Murphy, M. M., Peyer, J. G. & Morrison, S. J. Leptin-receptor-expressing mesenchymal stromal cells represent the main source of bone formed by adult bone marrow. Cell Stem Cell 15, 154–168 (2014).
Shu, H. S. et al. Tracing the skeletal progenitor transition during postnatal bone formation. Cell Stem Cell 28, 2122–2136.e3 (2021).
Baryawno, N. et al. A cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell 177, 1915–1932.e16 (2019).
Baccin, C. et al. Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization. Nat. Cell Biol. 22, 38–48 (2020).
Tikhonova, A. N. et al. The bone marrow microenvironment at single-cell resolution. Nature 569, 222–228 (2019).
Zhong, L. et al. Single cell transcriptomics identifies a unique adipose lineage cell population that regulates bone marrow environment. eLife 9, e54695 (2020).
Matsushita, Y. et al. A Wnt-mediated transformation of the bone marrow stromal cell identity orchestrates skeletal regeneration. Nat. Commun. 11, 332 (2020).
Chen, Q. et al. Apelin(+) endothelial niche cells control hematopoiesis and mediate vascular regeneration after myeloablative injury. Cell Stem Cell 25, 768–783.e6 (2019).
Hooper, A. T. et al. Engraftment and reconstitution of hematopoiesis is dependent on VEGFR2-mediated regeneration of sinusoidal endothelial cells. Cell Stem Cell 4, 263–274 (2009).
Zhou, B. O. et al. Bone marrow adipocytes promote the regeneration of stem cells and haematopoiesis by secreting SCF. Nat. Cell Biol. 19, 891–903 (2017).
Bowers, E. et al. Granulocyte-derived TNFalpha promotes vascular and hematopoietic regeneration in the bone marrow. Nat. Med. 24, 95–102 (2018).
Emont, M. P. et al. A single-cell atlas of human and mouse white adipose tissue. Nature 603, 926–933 (2022).
Hagberg, C. E. et al. Flow cytometry of mouse and human adipocytes for the analysis of browning and cellular heterogeneity. Cell Rep. 24, 2746–2756.e5 (2018).
Ye, R. et al. Impact of tamoxifen on adipocyte lineage tracing: inducer of adipogenesis and prolonged nuclear translocation of Cre recombinase. Mol. Metab. 4, 771–778 (2015).
Jeffery, E. et al. The adipose tissue microenvironment regulates depot-specific adipogenesis in obesity. Cell Metab. 24, 142–150 (2016).
Kalajzic, Z. et al. Directing the expression of a green fluorescent protein transgene in differentiated osteoblasts: comparison between rat type I collagen and rat osteocalcin promoters. Bone 31, 654–660 (2002).
Matsushita, Y. et al. CCN3 protein participates in bone regeneration as an inhibitory factor. J. Biol. Chem. 288, 19973–19985 (2013).
Naveiras, O. et al. Bone-marrow adipocytes as negative regulators of the haematopoietic microenvironment. Nature 460, 259–263 (2009).
Scheller, E. L. et al. Region-specific variation in the properties of skeletal adipocytes reveals regulated and constitutive marrow adipose tissues. Nat. Commun. 6, 7808 (2015).
Haemmerle, G. et al. Defective lipolysis and altered energy metabolism in mice lacking adipose triglyceride lipase. Science 312, 734–737 (2006).
Ahmadian, M. et al. Desnutrin/ATGL is regulated by AMPK and is required for a brown adipose phenotype. Cell Metab. 13, 739–748 (2011).
Logan, M. et al. Expression of Cre recombinase in the developing mouse limb bud driven by a Prxl enhancer. Genesis 33, 77–80 (2002).
Horowitz, M. C. et al. Bone marrow adipocytes. Adipocyte 6, 193–204 (2017).
Sanchez-Gurmaches, J., Hsiao, W. Y. & Guertin, D. A. Highly selective in vivo labeling of subcutaneous white adipocyte precursors with Prx1-Cre. Stem Cell Rep. 4, 541–550 (2015).
Martinez-Botas, J. et al. Absence of perilipin results in leanness and reverses obesity in Lepr(db/db) mice. Nat. Genet. 26, 474–479 (2000).
Snoeck, H. W. Aging of the hematopoietic system. Curr. Opin. Hematol. 20, 355–361 (2013).
Rossi, D. J. et al. Cell intrinsic alterations underlie hematopoietic stem cell aging. Proc. Natl Acad. Sci. USA 102, 9194–9199 (2005).
Verovskaya, E. V., Dellorusso, P. V. & Passegue, E. Losing sense of self and surroundings: hematopoietic stem cell aging and leukemic transformation. Trends Mol. Med. 25, 494–515 (2019).
Ambrosi, T. H. et al. Adipocyte Accumulation in the Bone Marrow during Obesity and Aging Impairs Stem Cell-Based Hematopoietic and Bone Regeneration. Cell Stem Cell 20, 771–784.e6 (2017).
Grabner, G. F., Xie, H., Schweiger, M. & Zechner, R. Lipolysis: cellular mechanisms for lipid mobilization from fat stores. Nat. Metab. 3, 1445–1465 (2021).
Perry, R. J. et al. Hepatic acetyl CoA links adipose tissue inflammation to hepatic insulin resistance and type 2 diabetes. Cell 160, 745–758 (2015).
Ono, T. et al. IL-17-producing gammadelta T cells enhance bone regeneration. Nat. Commun. 7, 10928 (2016).
Justesen, J. et al. Adipocyte tissue volume in bone marrow is increased with aging and in patients with osteoporosis. Biogerontology 2, 165–171 (2001).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Hamilton, T. G., Klinghoffer, R. A., Corrin, P. D. & Soriano, P. Evolutionary divergence of platelet-derived growth factor alpha receptor signaling mechanisms. Mol. Cell. Biol. 23, 4013–4025 (2003).
Haemmerle, G. et al. ATGL-mediated fat catabolism regulates cardiac mitochondrial function via PPAR-alpha and PGC-1. Nat. Med. 17, 1076–1085 (2011).
Fan, Y. et al. Parathyroid hormone directs bone marrow mesenchymal cell fate. Cell Metab. 25, 661–672 (2017).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender.Nat. Methods 20, 1323–1335 (2023).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e4 (2019).
Sacma, M. et al. Haematopoietic stem cells in perisinusoidal niches are protected from ageing. Nat. Cell Biol. 21, 1309–1320 (2019).
Gao, L., Decker, M., Chen, H. & Ding, L. Thrombopoietin from hepatocytes promotes hematopoietic stem cell regeneration after myeloablation. eLife 10, e69894 (2021).
Lee, Y., Leslie, J., Yang, Y. & Ding, L. Hepatic stellate and endothelial cells maintain hematopoietic stem cells in the developing liver. J. Exp. Med. 218, e20200882 (2021).
Lee, Y., DiMaulo-Milk, E., Leslie, J. & Ding, L. Hematopoietic stem cells temporally transition to thrombopoietin dependence in the fetal liver. Sci. Adv. 8, eabm7688 (2022).
Sarkaria, S. M. et al. Systematic dissection of coordinated stromal remodeling identifies Sox10(+) glial cells as a therapeutic target in myelofibrosis. Cell Stem Cell 30, 832–850.e6 (2023).
Gao, L. snRNA-seq analysis LeiDingLab-Nature_Genetics_2023:v1. Zenodo https://doi.org/10.5281/zenodo.8280830 (2023).
Acknowledgements
This work was supported by the Rita Allen Foundation, the Schaefer Scholar program, and the National Heart, Lung and Blood Institute (grant nos. R01HL132074 and R01HL153487). L.D. was also supported by a Scholar Award from the Leukemia and Lymphoma Society, grant no. R01HL155868, and grant no. R01GM146061 from the National Institutes of Health (NIH). H.H. was supported by the Uehara Memorial Foundation and the Japan Society for the Promotion of Science. L.G. was supported by a NYSTEM training grant and an American Heart Association postdoctoral fellowship. We thank M. Kissner and R. Gordon-Schneider at the Columbia Stem Cell Initiative for help on flow cytometry. We thank E. DiMaulo-Milk for critically reading the manuscript. This research was funded in part through the NIH/National Cancer Institute Cancer Center Support Grant, grant nos. P30CA013696 and P41EB027062.
Author information
Authors and Affiliations
Contributions
H.H. performed all of the experiments. L.G. performed analysis on the snRNA-seq data. D.N.T. and G.V.-N. helped with the adipocyte culture experiments and editing of the manuscript. H.H. and L.D. designed the experiments, interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bone marrow regenerates after irradiation.
a-j, Kinetics of maturing bone marrow hematopoietic cells after sublethal irradiation (6 Gy). Irradiated wild-type mice were analyzed at 2 weeks (2 w), 5 weeks (5 w), and 8 weeks (8 w) post-irradiation. Non-irradiated mice were used as controls (Con). The frequencies and numbers of CD3+ cells (a, b), B220+ cells (c, d), Gr1+ Mac1+ cells (e, f), Ter119+ cells (g, h), and CD41+ cells (i, j) are shown (n = 4 for each group). All data represent mean ± SD. One-way ANOVAs followed by Dunnett’s test were used to assess statistical significance.
Extended Data Fig. 2 The generation of a Plin1creER knockin allele.
a, Clustering of bone marrow cells based on t-Distributed Stochastic Neighbor Embedding (t-SNE) from published data12. b, Gene expression levels of Lepr, Adipoq, and Plin1 overlaid on t-SNE visualization. c, Clustering of bone marrow cells based on t-SNE from published data11. d, Gene expression levels of Lepr, Adipoq, and Plin1 overlaid on t-SNE visualization. e, RT-PCR analyses revealed that Plin1 is expressed by adipose tissues but not bone marrow mesenchymal stromal cells. Adipoq is expressed by bone marrow mesenchymal stromal cells while Ucp1 is only expressed by brown adipose tissue. β-Actin was used as a control for RT-PCR. Lane 1: Brown Adipose Tissue (BAT), Lane 2: White Adipose Tissue (WAT), Lane 3: sorted CD45/Ter119/CD31−PDGFRα+ bone marrow mesenchymal stromal cells (n = 1). f, Gene targeting strategy. Plin1creER was generated by inserting P2A-creER into the ninth exon of Plin1 using homologous recombination in ES cells. g, Representative genotyping PCR results confirmed the germline transmission of the Plin1creER allele (Lane 1: creER/creER, Lane 2: creER/+, Lane 3: +/+) (n = 3). h, Representative image of the bone marrow from creER/creER homozygous mice (n = 2). Adipocytes were stained with an anti-perilipin antibody (in green). Nuclei were stained with DAPI (in blue). The square indicates an enlarged region. Scale bar, 100 μm. i-k, Sublethally irradiated (6 Gy) wild-type mice were treated with three doses of corn oil or tamoxifen on days 10, 12, and 14 and analyzed on day 15 post-irradiation. Experimental scheme (i). Representative images of the bone marrow (j). Quantification of bone marrow adipocytes stained with an anti-perilipin antibody (n = 3 for each group) (k). Scale bars, 50 μm. l, Representative whole-mount confocal images and frequency of perilipin+ adipocytes that were tdTomato+ (n = 3) in the bone marrow from Plin1creER; Rosa26LSL-tdTomato mice treated with tamoxifen on days 10, 12, and 14 and analyzed on day 15 post-irradiation. Scale bar, 100 μm. m, Individual channels of Fig. 2c (15 d). Scale bar, 100 μm. All data represent mean ± SD. A two-sided Student’s t-test was used to assess statistical significance in k.
Extended Data Fig. 3 Prominent dedifferentiation of bone marrow adipocyte was evident with Plin1-creER induction by a low dose of tamoxifen.
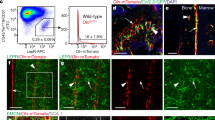
a, Experimental scheme. b, c, Irradiated Plin1creER; Rosa26LSL-tdTomato mice were administered 0.2 mg tamoxifen on day 14 post-irradiation and analyzed 36 hours (36 hr) later (n = 3). Representative images of the bone marrow are shown. Scale bar, 50 μm. d, A representative flow cytometric plot showing no labeling of stromal cells in enzymatically digested bone marrow from irradiated Plin1creER; Rosa26LSL-tdTomato mice treated with 0.2 mg tamoxifen on day 14 post-irradiation and analyzed 36 hours (36 hr) later (n = 3). e, f, Irradiated Plin1creER; Rosa26LSL-tdTomato mice were administered 0.2 mg tamoxifen on day 14 and analyzed 6 weeks (6 w) later (n = 5). Representative images of the bone marrow are shown. Scale bar, 50 μm. g, A representative flow cytometric plot of enzymatically digested bone marrow cells of the tibia from irradiated Plin1creER; Rosa26LSL-tdTomato mice treated as in e and f (n = 6). h, Frequency of tdTomato+ cells that were perilipin− in the bone marrow of irradiated Plin1creER; Rosa26LSL-tdTomato mice treated with 0.2 mg tamoxifen on day 14 post-irradiation and analyzed 6 weeks later (n = 5). i, Representative image showing adipocyte-derived tdTomato+ bone marrow stromal cells from irradiated Plin1creER; Rosa26LSL-tdTomato mice expressed LepR (arrows) (n = 2). The mice were treated with 0.2 mg tamoxifen on day 14 after irradiation and analyzed 6 weeks later. Scale bar, 50 μm. All data represent mean ± SD.
Extended Data Fig. 4 Dedifferentiation of bone marrow adipocytes following 5FU challenge.
a, Representative images of the bone marrow showing adipocyte content from the non−treated control (Con), 10 days (10 d), 4 weeks (4 w), or 6 weeks (6 w) after 5FU (150 mg kg-1) administration in wild-type mice. Adipocytes were stained with an anti-perilipin antibody (in green). Scale bars, 100 μm. b, Quantification of bone marrow adipocytes following 5FU treatment (n = 3 for each group). c-e, Plin1creER; Rosa26LSL-tdTomato mice were treated with 3 doses of tamoxifen on days 6, 8, and 10 and analyzed 4 weeks (4 w) and 6 weeks (6 w) after 5FU treatment. Representative images of the bone marrow (c), the frequency of adipocytes that were tdTomato+ (n = 3 for each group) (d), and the frequency of tdTomato+ cells that were perilipin- (n = 3 for each group) (e) are shown. Scale bars, 100 μm. f, Representative image showing that adipocyte-derived stromal cells express LepR (arrows) in the bone marrow from Plin1creER; Rosa26LSL-tdTomato mice after 5FU treatment (n = 3). The mice were treated with tamoxifen on days 6, 8, and 10, and analyzed 4 weeks after 5FU treatment. The square indicates an enlarged region. Scale bar, 50 μm. All data represent mean ± SD. A one-way ANOVA followed by Dunnett’s test was used in b. Two-sided Student’s t-tests were used in d and e.
Extended Data Fig. 5 snRNA-seq reveals stromal contribution by bone marrow adipocytes during regeneration.
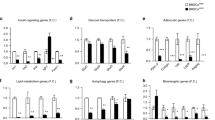
a, Heatmap showing the signature genes of two distinct mesenchymal cell clusters in the regenerating bone marrow (Lepr+ and Adipocyte). The Lepr+ cluster is further divided into two clusters (Lepr+_Osteo and Lepr+_Adipo). b, Violin plots showing the expression levels of Lepr, Adipoq, and Alpl. c, d, Expression levels of Perilipin family genes (Plin1, Plin2, Plin3, Plin4, and Plin5) and three genes coding lipases (Atgl, Lipe, and Mgll) are shown in violin plots (c) and dot plot (d). e, Dot plot showing the expression levels of Adrb1, Adrb2, Adrb3, and Il6ra.
Extended Data Fig. 6 Ex vivo properties of mature bone marrow adipocytes and their derivatives.
a, Experimental scheme for monitoring bone marrow adipocyte culture in vitro. Individual bone marrow adipocytes isolated from Plin1creER; Rosa26LSL-tdTomato mice at 15 days after sublethal irradiation (tamoxifen administration on day 14) were followed by time-lapse imaging in culture for 7 days. Lipid-filled adipocytes adhering to the ceiling of the flask on day 2 were tracked by confocal microscopy. b, Representative images of live cell imaging of bone marrow tdTomato+ adipocytes and their derivatives from day 2 to 7 (n = 2). Arrows point to adipocyte-derived cells (tdTomato+) with a stromal morphology but lacking lipid droplets (LipiDye II negative). Lipid droplets were stained with LipiDye II (in green). Scale bars, 50 μm. c, Experimental scheme for testing stromal cell function in vitro. d, Sublethally irradiated Plin1creER; Rosa26LSL-tdTomato mice were administered tamoxifen on days 10, 12, and 14 post-irradiation. At 8 weeks post-irradiation, CFU-F assays were performed with enzymatically digested bone marrow cells. Attached cells at clonal density were counted one day after plating and colonies were counted 12 days later. A representative image of a CFU-F colony (>25 cells) stained by Crystal Violet is shown. Scale bar, 500 μm. e, A representative image showing that tdTomato+ stromal cells are proliferative. The cells were from single tdTomato+ cells. Scale bar, 200 μm. f, Quantifications of CFU-F frequency from attached tdTomato− and tdTomato+ stromal cells (n = 4). g, Representative images of live cell imaging of a single tdTomato+ cell for 5 days. Lipid droplets were stained by LipiDye II (in green, arrow) on day 5 to show that lipid droplets are accumulating in a tdTomato+ cell. Scale bars, 50 μm. h, Quantification of adipocyte frequency from attached tdTomato− and tdTomato+ stromal cells (n = 4). After initiation of cell culture as in c, cells were cultured in a normal media without differentiation induction for 12 days and then LipiDye II+ adipocytes were counted. i, A representative image of a colony stained by Alizarin Red S showing osteoblastic differentiation. Scale bar, 500 μm. j, Quantification of CFU-osteoblasts (CFU-Ob) frequency from attached tdTomato- and tdTomato+ stromal cells (n = 4). After initiation of cell culture as in c, cells were cultured in a normal media for 12 days and then cultured in osteolineage differentiation media for 14 days. All data represent mean ± SD. Two-sided Student’s t-tests were used to assess statistical significance.
Extended Data Fig. 7 Perturbed lymphopoiesis in the bone marrow of Prx1-cre; Atglfl/fl mice after irradiation.
a-e, The frequencies and numbers of CD3+ cells (a), B220+ cells (b), Gr1+ Mac1+ cells (c), Ter119+ cells (d) and CD41+ cells (e) in the bone marrow of Prx1-cre; Atglfl/fl (Mut) and control mice (Con) (n = 7 for Con, n = 6 for Mut). f-i, Representative flow cytometric plots of LK (Lin− c-Kit+) cells from Prx1−cre; Atglfl/fl (Mut) and control mice (Con) at 8 weeks post-irradiation (f). Quantifications of CMP number (g), GMP number (h), and MEP number (i) in the bone marrow of Mut and Con at 15 days (n = 4 for each group) and 8 weeks (n = 6 for each group) post-irradiation. j, The number of CD45/Ter119/CD31−PDGFRα+ bone marrow mesenchymal stromal cells of Prx1−cre; Atglfl/fl (Mut) and control mice (Con) at 8 weeks post-irradiation (n = 5 for each group). A one-sided Student’s t-test was used. k-o, Representative flow cytometric plots and frequencies of bone marrow cells that are positive for CD3 (k), B220 (l), Gr1 and Mac1 (m), Ter119 (n), and CD41 (o). Prx1-cre; Atglfl/fl (Mut) and control mice (Con) were analyzed at 8 weeks post-irradiation (n = 8 for Con, n = 9 for Mut). p, Experimental scheme for transplantation assays in Fig. 7m. Competitive transplantation of 106 bone marrow cells (CD45.2) from Prx1-cre; Atglfl/f or control mice with 106 competitor bone marrow cells (CD45.1) into lethally irradiated recipients. Donor and competitor cells were harvested from sublethally irradiated mice at 8 weeks post-irradiation. All data represent mean ± SD. Two-sided Student’s t-tests were used in a-e, g-i, and k-o.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Uncropped gel image.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Uncropped gel image.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hirakawa, H., Gao, L., Tavakol, D.N. et al. Cellular plasticity of the bone marrow niche promotes hematopoietic stem cell regeneration. Nat Genet 55, 1941–1952 (2023). https://doi.org/10.1038/s41588-023-01528-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01528-2