Abstract
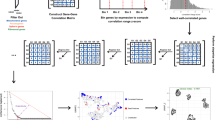
As single-cell datasets grow in sample size, there is a critical need to characterize cell states that vary across samples and associate with sample attributes, such as clinical phenotypes. Current statistical approaches typically map cells to clusters and then assess differences in cluster abundance. Here we present co-varying neighborhood analysis (CNA), an unbiased method to identify associated cell populations with greater flexibility than cluster-based approaches. CNA characterizes dominant axes of variation across samples by identifying groups of small regions in transcriptional space—termed neighborhoods—that co-vary in abundance across samples, suggesting shared function or regulation. CNA performs statistical testing for associations between any sample-level attribute and the abundances of these co-varying neighborhood groups. Simulations show that CNA enables more sensitive and accurate identification of disease-associated cell states than a cluster-based approach. When applied to published datasets, CNA captures a Notch activation signature in rheumatoid arthritis, identifies monocyte populations expanded in sepsis and identifies a novel T cell population associated with progression to active tuberculosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data analyzed during this study are available in three previously published articles6,7,8. Source data are provided with this paper.
Code availability
An open-source repository containing code for running CNA is available at https://github.com/immunogenomics/cna; an open-source repository containing code underlying all figures and tables is available at https://github.com/immunogenomics/cna-display; and an open-source repository containing code underlying all simulations is available at https://github.com/immunogenomics/cna-sim. Source data are provided with this paper.
References
Kashima, Y. et al. Single-cell sequencing techniques from individual to multiomics analyses. Exp. Mol. Med. 52, 1419–1427 (2020).
Andrews, T. S. et al. Tutorial: guidelines for the computational analysis of single-cell RNA sequencing data. Nat. Protoc. 16, 1–9 (2021).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Burkhardt, D. B. et al. Quantifying the effect of experimental perturbations at single-cell resolution. Nat. Biotechnol. 39, 619–629 (2021).
Nathan, A. et al. Multimodally profiling memory T cells from a tuberculosis cohort identifies cell state associations with demographics, environment and disease. Nat. Immunol. 22, 781–793 (2021).
Reyes, M. et al. An immune-cell signature of bacterial sepsis. Nat. Med. 26, 333–340 (2020).
Wei, K. et al. Notch signalling drives synovial fibroblast identity and arthritis pathology. Nature 582, 259–264 (2020).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Butler, A. et al. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Haghverdi, L. et al. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Liu, J. et al. Jointly defining cell types from multiple single-cell datasets using LIGER. Nat. Protoc. 15, 3632–3662 (2020).
Fonseka, C. Y. et al. Mixed-effects association of single cells identifies an expanded effector CD4+ T cell subset in rheumatoid arthritis. Sci. Transl. Med. 10, eaaq0305 (2018).
Millard, N. et al. Maximizing statistical power to detect clinically associated cell states with scPOST. Preprint at https://www.biorxiv.org/content/10.1101/2020.11.23.390682v1 (2020).
Liu, Z. et al. Notch signaling in postnatal joint chondrocytes, but not subchondral osteoblasts, is required for articular cartilage and joint maintenance. Osteoarthritis Cartilage 24, 740–751 (2016).
Wang, X. & Astrof, S. Neural crest cell-autonomous roles of fibronectin in cardiovascular development. Development 143, 88–100 (2016).
Zhang, F. et al. Defining inflammatory cell states in rheumatoid arthritis joint synovial tissues by integrating single-cell transcriptomics and mass cytometry. Nat. Immunol. 20, 928–942 (2019).
Sanlioglu, S. et al. Lipopolysaccharide induces Rac1-dependent reactive oxygen species formation and coordinates tumor necrosis factor-α secretion through IKK regulation of NF-κB. J. Biol. Chem. 276, 30188–30198 (2001).
Pan, C. et al. Suppression of the RAC1/MLK3/p38 signaling pathway by β-elemene alleviates sepsis-associated encephalopathy in mice. Front. Neurosci. 13, 358 (2019).
von Knethen, A. & Brüne, B. Histone deacetylation inhibitors as therapy concept in sepsis. Int. J. Mol. Sci. 20, 346 (2019).
Wu, H.-P. et al. Serial increase of IL-12 response and human leukocyte antigen-DR expression in severe sepsis survivors. Crit. Care 15, R224 (2011).
Steinhauser, M. L. et al. Multiple roles for IL-12 in a model of acute septic peritonitis. J. Immunol. 162, 5437–5443 (1999).
Oliveira, N. M. et al. Sepsis induces telomere shortening: a potential mechanism responsible for delayed pathophysiological events in sepsis survivors? Mol. Med. 22, 886–891 (2016).
Gutierrez-Arcelus, M. et al. Lymphocyte innateness defined by transcriptional states reflects a balance between proliferation and effector functions. Nat. Commun. 10, 687 (2019).
Cano-Gamez, E. et al. Single-cell transcriptomics identifies an effectorness gradient shaping the response of CD4+ T cells to cytokines. Nat. Commun. 11, 1801 (2020).
Luecken, M. et al. Benchmarking atlas-level data integration in single-cell genomics. Preprint at https://www.biorxiv.org/content/10.1101/2020.05.22.111161v1 (2020).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Klein, S. L. & Flanagan, K. L. Sex differences in immune responses. Nat. Rev. Immunol. 16, 626–638 (2016).
Silva, C. L. et al. Cytotoxic T cells and mycobacteria. FEMS Microbiol. Lett. 197, 11–18 (2001).
Li, M. et al. Age related human T cell subset evolution and senescence. Immun. Ageing 16, 24 (2019).
Shirai, T. et al. TH1-biased immunity induced by exposure to Antarctic winter. J. Allergy Clin. Immunol. 111, 1353–1360 (2003).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at https://www.biorxiv.org/content/10.1101/060012v2 (2021).
Acknowledgements
We thank A. Gupta, D. Kotliar, Y. Luo, N. Millard, M. Reyes, S. Sakaue, F. Zhang, the members of the CGTA discussion group and the Raychaudhuri lab for helpful discussions and feedback. This work is supported, in part, by funding from the National Institutes of Health (NIH) including UH2AR067677, U19 AI111224, U01 HG009379 and 1R01AR063759. S.A. was supported by the Swiss National Science Foundation postdoctoral mobility fellowships P2ELP3_172101 and P400PB_183823 and NIH grant T32HG010464. L.R. was supported, in part, by NIH 5T32HG2295-17. J.K. was supported by NIH grant T32GM007753. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Y.A.R., L.R. and S.R. designed and conceptualized the study. Y.A.R. and L.R. designed and implemented the algorithm and performed simulations. Y.A.R., L.R. and J.K. performed analysis of real data. A.N., S.A. and I.K. provided input on methodologic design and real data analysis. D.B.M., M.M., A.N. and I.K. provided dataset-specific expertise. Y.A.R., L.R. and S.R. wrote the manuscript with input from the remaining authors.
Corresponding author
Ethics declarations
Competing interests
S.R. serves as a consultant for Gilead, Pfizer, Janssen and Rheos Medicines and is a founder of Mestag Therapeutics. I.K. serves as a consultant for Mestag Therapeutics. The other authors declare no competing interests.
Additional information
Peer review information Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–20 and Supplementary Figs. 1–13
Source data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Rights and permissions
About this article
Cite this article
Reshef, Y.A., Rumker, L., Kang, J.B. et al. Co-varying neighborhood analysis identifies cell populations associated with phenotypes of interest from single-cell transcriptomics. Nat Biotechnol 40, 355–363 (2022). https://doi.org/10.1038/s41587-021-01066-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01066-4
This article is cited by
-
Benchmarking differential abundance methods for finding condition-specific prototypical cells in multi-sample single-cell datasets
Genome Biology (2024)
-
Cellograph: a semi-supervised approach to analyzing multi-condition single-cell RNA-sequencing data using graph neural networks
BMC Bioinformatics (2024)
-
A concerted neuron–astrocyte program declines in ageing and schizophrenia
Nature (2024)
-
CD200+ fibroblasts form a pro-resolving mesenchymal network in arthritis
Nature Immunology (2024)
-
Single-cell genomics meets human genetics
Nature Reviews Genetics (2023)