Abstract
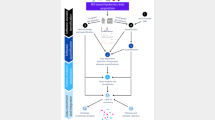
We present Mass Spectrometry-Data Independent Analysis software version 4 (MS-DIAL 4), a comprehensive lipidome atlas with retention time, collision cross-section and tandem mass spectrometry information. We formulated mass spectral fragmentations of lipids across 117 lipid subclasses and included ion mobility tandem mass spectrometry. Using human, murine, algal and plant biological samples, we annotated and semiquantified 8,051 lipids using MS-DIAL 4 with a 1–2% estimated false discovery rate. MS-DIAL 4 helps standardize lipidomics data and discover lipid pathways.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
All data resources are freely available at http://prime.psc.riken.jp/. Source data for Fig. 2 and Extended Data Figs. 4 and 5 are provided with the paper.
Code availability
All program packages and source code are freely available at http://prime.psc.riken.jp/.
References
Harayama, T. & Riezman, H. Nat. Rev. Mol. Cell Biol. 19, 281–296 (2018).
Kind, T. et al. Mass Spectrom. Rev. 37, 513–532 (2018).
Tsugawa, H., Ikeda, K. & Arita, M. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 1862, 762–765 (2017).
Liebisch, G. et al. Nat. Metab. 1, 745–747 (2019).
Hoffmann, N. et al. Anal. Chem. 91, 3302–3310 (2019).
Tsugawa, H. et al. Nat. Methods 16, 295–298 (2019).
Kind, T. et al. Nat. Methods 10, 755–758 (2013).
Pauling, J. K. et al. PLoS ONE 12, e0188394 (2017).
Fahy, E. et al. J. Lipid Res. 50(Suppl.), S9–S14 (2009).
Liebisch, G. et al. J. Lipid Res. 54, 1523–1530 (2013).
Bowden, J. A. et al. J. Lipid Res. 58, 2275–2288 (2017).
Burla, B. et al. J. Lipid Res. 59, 2001–2017 (2018).
Ulmer, C. Z. et al. Anal. Chem. 89, 13069–13073 (2017).
Vasilopoulou, C. G. et al. Nat. Commun. 11, 1–11 (2020).
Lintonen, T. P. I. et al. Anal. Chem. 86, 9662–9669 (2014).
Gorusupudi, A., Liu, A., Hageman, G. S. & Bernstein, P. S. J. Lipid Res. 57, 499–508 (2016).
Cohen, L. J. et al. Nature 549, 48–53 (2017).
Masukawa, Y. et al. J. Lipid Res. 49, 1466–1476 (2008).
Schleyer, G. et al. Nat. Microbiol. 4, 527–538 (2019).
Naoe, S., Tsugawa, H., Takahashi, M., Ikeda, K. & Arita, M. Metabolites 9, 241 (2019).
Tsugawa, H. et al. Nat. Methods 12, 523–526 (2015).
Yap, C. W. J. Comput. Chem. 32, 1466–1474 (2011).
Tsugawa, H. et al. J. Cheminform. 9, 1–12 (2017).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. Bioinformatics 24, 2534–2536 (2008).
Lai, Z. et al. Nat. Methods 15, 53–56 (2017).
Hartler, J. et al. Nat. Methods 14, 1171–1174 (2017).
Nakanishi, H., Iida, Y., Shimizu, T. & Taguchi, R. J. Biochem. 147, 245–256 (2010).
Tsugawa, H. et al. Anal. Chem. 88, 7946–7958 (2016).
Ni, Z., Angelidou, G., Hoffmann, R. & Fedorova, M. Sci. Rep. 7, 1–14 (2017).
Haimi, P., Uphoff, A., Hermansson, M. & Somerharju, P. Anal. Chem. 78, 8324–8331 (2006).
Song, H., Hsu, F. F., Ladenson, J. & Turk, J. J. Am. Soc. Mass Spectrom. 18, 1848–1858 (2007).
Alcoriza-Balaguer, M. I. et al. Anal. Chem. 91, 836–845 (2019).
Hutchins, P. D., Russell, J. D. & Coon, J. J. Cell Syst. 6, 621–625 (2018).
Koelmel, J. P. et al. BMC Bioinformatics 18, 1–11 (2017).
Kyle, J. E. et al. Bioinformatics 33, 1744–1746 (2017).
Ni, Z., Angelidou, G., Lange, M., Hoffmann, R. & Fedorova, M. Anal. Chem. 89, 8800–8807 (2017).
Kochen, M. A. et al. Anal. Chem. 88, 5733–5741 (2016).
Pluskal, T., Castillo, S., Villar-Briones, A. & Oresic, M. BMC Bioinformatics 11, 395 (2010).
Zhou, Z., Tu, J., Xiong, X., Shen, X. & Zhu, Z. J. Anal. Chem. 89, 9559–9566 (2017).
Plante, P. L. et al. Anal. Chem. 91, 5191–5199 (2019).
Colby, S. M., Nuñez, J. R., Hodas, N. O., Corley, C. D. & Renslow, R. R. Anal. Chem. 92, 1720–1729 (2020).
Colby, S. M. et al. Anal. Chem. 91, 4346–4356 (2019).
Acknowledgements
This work was mainly supported by the JSPS Grant-in-Aid for Scientific Research on Innovative Areas ‘LipoQuality’ (grant nos. 15H05897, 15H05898 to Makoto Arita). This work was partially supported by AMED-LEAP under grant no. JP18gm0010003 (to Makoto Arita), RIKEN Pioneering Project ‘Glyco-lipidologue Initiative’ (to Masanori Arita and Makoto Arita), JSPS KAKENHI (grant nos. 18H02432, 18K19155 for H.T.), JST National Bioscience Database Center (NBDC to Masanori Arita), the Czech Science Foundation (grant no. 20-21114S to T.C.) and the Czech Ministry of Education, Youth and Sports (grant no. LTAUSA19124 to T.C.). We thank S. Madden and J. Fjeldsted (Agilent Technologies) for sharing the MIDAC SDK, S. Brehmer (Bruker) for sharing the tims SDK and the Waters developmental team for sharing the SDK for traveling wave ion mobility data. We thank X. Li for his help in validating MS-DIAL libraries. We also thank K. Takano and A. Hori (RIKEN) for LC–MS analysis, N. Hoffmann and S. Neumann (Leibniz Institute) for the discussion of mztab-M export and G. Liebisch (University Hospital Regensburg) for the discussion of lipid nomenclature.
Author information
Authors and Affiliations
Contributions
H.T., K.I., Masanori Arita and Makoto Arita designed the research. H.T. developed all computational programs, and M.T. and I.T. contributed to source coding for the lipid characterizations and IM-DIA-MS deconvolution, respectively. H.T. and M.T. curated most of the lipid mass spectra. H.T., K.I., Y.M., H.U. and Y.H. analyzed biological samples, and J.K., T.C., O.F. and K.S. provided LC–MS/MS data from NIST human plasma. H.T. and A.S. performed software comparisons, and H.T., A.S., P.B., Z.Z. and Z.-J.Z performed RT and CCS predictions for metabolites. H.T., M.T., H.U., N.O., Y.O. and J.K. performed the lipid MS/MS library creations, and Y.Y. created the lipidomics database. H.T. wrote the manuscript. H.T., K.I., Masanori Arita and Makoto Arita thoroughly discussed this project, and all authors helped to improve the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y.M., who performed LC–IM-MS/MS analyses, is an appreciation specialist at Bruker Japan.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Summary of the hybrid scoring system for lipid tandem mass spectrometry (MS/MS) annotations.
The MS/MS spectrum of the acetate adduct form of sphingomyelin (SM) 18:1;2O/16:0 (m/z 761.581) is shown as an example. The characteristic fragment ion of m/z 168.043 (C4H11NO4P–) and neutral loss of 74.037 Da (CH3COO + CH3) to specify the SM lipid class were detected whereas the product ion of m/z 449.315 as the neutral loss of acyl 16:0 to characterize the N-acyl chain moiety was observed in the MS/MS spectrum. The classical spectral similarity calculation using dot- and reverse dot product scores is executed to filter out noisy spectra: the MS/MS spectrum is recognized as unknown if the dot product is <0.1 or the reverse dot product is <0.5. For the decision tree algorithm for the acetate adduct form of SM lipid, the fragment existence of SM-specific fragment ions is evaluated, in which the relative abundance cut off is also utilized. If the neutral loss of N-acyl chain (as observed by m/z 449.315) is detected and the relative abundance is >0.1% when compared with the base peak of m/z 687.550 (recognized as 100%), the lipid structure of “molecular species level” is represented as SM 18:1;2O/16:0. If no acyl chain fragment exists, the description of “species level” is exported as SM 34:1;2O.
Extended Data Fig. 2 Concept of the ion mobility base framework (IBF) format.
The raw data of Bruker’s parallel accumulation-serial fragmentation (PASEF) are shown as an example. In the raw data format (.d), 400–800 spectral records along with the changes of trapped electric field are stored at each retention time (RT) point, and their recording is sequenced by the end of the liquid chromatography (LC) condition. The file size of our 20 min LC condition was approximately 1 GB. The IBF format (.ibf) was designed to rapidly access ion mobility (IM)-MS data and the accumulated ion signals summing the mass spectral data of IM axis at the same RT bin were also stored to achieve a rapid peak picking process in the RT and m/z dimensions. The calibrant information such as beta coefficient and intercept values for the Agilent single field method and t0, exponent, and coefficient values for the Waters collision cross section calculation were also stored in the IBF format. The file size was roughly equivalent to the original size, and data retrieval could be accomplished within 10 s.
Extended Data Fig. 3 Scheme of peak picking for liquid chromatography coupled with ion mobility tandem mass spectrometry (LC-IM-MS/MS).
The extracted ion chromatogram (EIC) for a certain m/z value with approximately 10 ppm mass tolerance is constructed by using accumulated MS1 spectral panels, which are constructed by summing spectra of the IM axis with three-decimal binning for m/z value. After the peak detection method is performed in the retention time (RT) axis, the extracted ion mobilogram (EIM) is constructed in the IM axis by accumulating ions from the left- and right edges of the detected peak in the RT axis, followed by the peak detection procedure applied to the EIM. For example, the average peak width on the RT dimension is 10–30 s whereas the average peak width on the IM dimension is 30–60 ms in our LC-IM-MS/MS condition. The MS/MS spectrum from data-dependent/independent acquisition methods is assigned to each peak on the IM axis. As a result, each peak spot in the RT and m/z dimension has more than one peak spot in the mobility and m/z dimension, and the peak contains peak height, peak area, RT, mobility value, m/z, and MS/MS as the peak properties. The collision cross-section (CCS) is calculated using the Mason–Schamp equation for Bruker trapped IM, the single field CCS method for Agilent drift tube IM, and the IM calibration function for Waters travelling wave IM.
Extended Data Fig. 4 Evaluations for the MS-DIAL 4 annotation pipeline.
The accuracy, precision, recall, specificity, and false discovery rate (FDR; %) were calculated by using the set of true positives (TPs) and true negatives (TNs), which are available as Supplementary Data 2. a, The set of 12,798 TPs and 64,121 TNs from liquid chromatography-electrospray ionization-positive-tandem mass spectrometry (LC-ESI(+)-MS/MS) (data dependent acquisition, DDA) data was processed where the minimum FDR value was 1.61% at 0.75 min retention time (RT) tolerance. b, The set of 10,600 TPs and 30,290 TNs from LC-ESI(−)-MS/MS-DDA was processed, where the minimum FDR value was 1.61% at 0.75 min RT tolerance. c, The set of 2,598 TPs and 30,131 TNs from LC-ESI(+)-IM-MS/MS (PASEF) was processed with various collision cross-section (CCS) tolerances without RT information, where the minimum FDR was 2.82% at 5 Å2 CCS tolerance. d, The set of 1,670 TPs and 20,737 TNs from LC-ESI(−)-IM-MS/MS (PASEF) was processed with no RT, where the minimum FDR was 2.22% at 2 Å2 CCS tolerance. e, The same set as used in (c) was processed with 1.5 min RT tolerance, where the minimum FDR was 1.31% at 7 Å2 CCS tolerance. f, The same set as in (d) was processed with 1.5 min RT tolerance, where the minimum FDR was 1.22% at 3 Å2 CCS tolerance.
Extended Data Fig. 5 Mapping lipid diversity in mammalian tissues and cells.
Hierarchical clustering analysis was performed using the data matrix containing the count of molecules categorized to each lipid subclass; the count was scaled from −1 to 1. Overall, 112 lipid subclasses in addition to PC, CE, DG, TG, Cer-NS and SM subclasses containing very-long-chain poly unsaturated fatty acid (VLCPUFA) are described in the x-axis; total counts of molecular species annotated in each lipid subclass are in brackets. The LipidMaps category and a specialized lipid class containing VLCPUFA are given. Asterisks indicate the lipid subclasses that only MS-DIAL 4 can characterize when compared with existing lipidomics software tools evaluated in this study. The lipid nomenclature is detailed in Supplementary Table 2. Note that although potentially quantitative, no detection does not evince the nonexistence of certain lipid subclasses because of technical limitations.
Extended Data Fig. 6 Prediction of putative lipid structures by untangling the tandem mass spectrometry (MS/MS) spectra.
a, Fragment assignments for acylated hexosyl ceramide (AHexCer) containing sphingosine (18:1;2O), N-acyl chain (22:0;O), and O-acyl chain (O-16:0) are shown. Compared to the O-acyl chain positional isomer (HexCer-EOS), the fragment ion of Hex-O-acyl 16:0 (m/z 401.29) is observed in positive ion mode, and no hexosyl loss (162 Da) is found before the O-acyl 16:0 loss (238 Da) in negative ion mode. The behaviors were reproducible in all of the 24 different molecular species of AHexCer annotated in this study. b, Fragment assignments for acylated sphingomyelin (ASM) containing sphingosine (18:1;2O), N-acyl chain (24:1), and O-acyl chain (O-16:0) are shown although the sphingobase and N-acyl chain moieties cannot be determined as the molecular species level basically at least in our experimental condition. The SM specific fragment ions can be observed in both positive and negative ion MS/MS spectra. In addition, the product ion derived from the fatty acid moiety (O-acyl 16:0) was detected in both ion modes although they cannot be observed in usual SM lipid species. These behaviors were also true in all of the eight different molecules of ASM lipid subclass found in this study. c, An example of acylated uronosyldiacylglycerol (ADGGA) containing 16:0 and 18:2 acyl chains in the glycerol moiety and O-16:0 acyl chain in the uronosyl moiety is described. The fragment ions of HexA-O-acyl 16:0 (m/z 415.269) and acyl 16:0 are observed in positive ion mode where HexA denotes the uronic acid moiety. These behaviors were true in all of the 28 different molecules of the ADGGA lipid subclass found in this study.
Extended Data Fig. 7 Validation for the proposed structure of acylated SM (ASM) by the complete synthesis.
a, The extracted ion chromatogram (EIC) of m/z 1109.921 (±0.015) in negative ion mode data for mouse kidney sample. The retention time of chromatogram peak top is also described. b, The EIC of m/z 1109.921 (±0.015) in negative ion mode data for the synthesized compound sample of ASM (O-16:0)18:1(4E);(1OH,3OH)/24:1(15Z) which should be detected by m/z 1109.921 as acetate adduct form. c, The EIC of m/z 1109.921 (±0.015) in negative ion mode data for the mixture of mouse kidney sample and synthesize compound sample (1:1, v/v). d, The experimental ESI(−)-MS/MS spectrum of m/z 1109.923 at 14.5 min in LC-ESI(−)-MS/MS data of mouse kidney tissue. e, The experimental ESI(+)-MS/MS spectrum of m/z 1051.915 at 14.5 min in LC-ESI(+)-MS/MS data of mouse kidney tissue. f, The experimental ESI(−)-MS/MS spectrum of m/z 1109.916 at 14.5 min in LC-ESI(−)-MS/MS data of ASM (O-16:0)18:1(4E);(1OH,3OH)/24:1(15Z). g, The experimental ESI(+)-MS/MS spectrum of m/z 1051.915 at 14.5 min in LC-ESI(+)-MS/MS data of ASM (O-16:0)18:1(4E);(1OH,3OH)/24:1(15Z). The characteristic product ion to annotate the sphingobase moiety (SPB 18:1;2O) was also detected, and it was also observed in kidney data for the additional experiment as described in (e). The experiments were repeated independently (n=3) with similar results.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2 and Fig. 1
Supplementary Table
Supplementary Tables 1–8. Full descriptions for each table are provided in the Excel file.
Supplementary Data 1
Results of lipid profiling for each biological study.
Supplementary Data 2
Set of true positives and true negatives for evaluating the MS-DIAL annotation pipeline.
Source data
Source Data Fig. 2
Statistical Source Data for Fig. 2a–h
Source Data Extended Data Fig. 4
Statistical Source Data for Extended Data Fig. 4
Source Data Extended Data Fig. 5
Statistical Source Data for Extended Data Fig. 5
Rights and permissions
About this article
Cite this article
Tsugawa, H., Ikeda, K., Takahashi, M. et al. A lipidome atlas in MS-DIAL 4. Nat Biotechnol 38, 1159–1163 (2020). https://doi.org/10.1038/s41587-020-0531-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0531-2
This article is cited by
-
A lipidome landscape of aging in mice
Nature Aging (2024)
-
Modeling dry eye with an air–liquid interface in corneal epithelium-on-a-chip
Scientific Reports (2024)
-
Two mitochondrial DNA polymorphisms modulate cardiolipin binding and lead to synthetic lethality
Nature Communications (2024)
-
Unbiased serum metabolomic analysis in cats with naturally occurring chronic enteropathies before and after medical intervention
Scientific Reports (2024)
-
Single-cell and spatial RNA sequencing reveal the spatiotemporal trajectories of fruit senescence
Nature Communications (2024)