Abstract
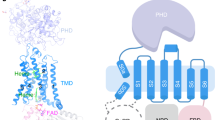
Phagocyte NADPH oxidase, a protein complex with a core made up of NOX2 and p22 subunits, is responsible for transferring electrons from intracellular NADPH to extracellular oxygen1. This process generates superoxide anions that are vital for killing pathogens1. The activation of phagocyte NADPH oxidase requires membrane translocation and the binding of several cytosolic factors2. However, the exact mechanism by which cytosolic factors bind to and activate NOX2 is not well understood. Here we present the structure of the human NOX2–p22 complex activated by fragments of three cytosolic factors: p47, p67 and Rac1. The structure reveals that the p67–Rac1 complex clamps onto the dehydrogenase domain of NOX2 and induces its contraction, which stabilizes the binding of NADPH and results in a reduction of the distance between the NADPH-binding domain and the flavin adenine dinucleotide (FAD)-binding domain. Furthermore, the dehydrogenase domain docks onto the bottom of the transmembrane domain of NOX2, which reduces the distance between FAD and the inner haem. These structural rearrangements might facilitate the efficient transfer of electrons between the redox centres in NOX2 and lead to the activation of phagocyte NADPH oxidase.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Cryo-EM maps and the atomic coordinate of the NOX2 complex in the resting state have been deposited in the Electron Microscopy Data Bank (EMDB) and PDB under the accession codes EMD-38016 and 8X2L. Cryo-EM maps and the atomic coordinate of the NOX2 complex in the activated state have been deposited in the EMDB and PDB under the accession codes EMD-37477 and 8WEJ. The entries 8GZ3, 1E96, 5O0X and 7D3F used in this study were downloaded from the PDB. Source data are provided with this paper.
References
Winterbourn, C. C., Kettle, A. J. & Hampton, M. B. Reactive oxygen species and neutrophil function. Annu. Rev. Biochem. 85, 765–792 (2016).
Lambeth, J. D. & Neish, A. S. Nox enzymes and new thinking on reactive oxygen: a double-edged sword revisited. Annu. Rev. Pathol. 9, 119–145 (2014).
Heyworth, P. G., Cross, A. R. & Curnutte, J. T. Chronic granulomatous disease. Curr. Opin. Immunol. 15, 578–584 (2003).
Diebold, B. A., Smith, S. M., Li, Y. & Lambeth, J. D. NOX2 as a target for drug development: indications, possible complications, and progress. Antioxid. Redox. Signal. 23, 375–405 (2015).
Magnani, F. et al. Crystal structures and atomic model of NADPH oxidase. Proc. Natl Acad. Sci. USA 114, 6764–6769 (2017).
Sun, J. Structures of mouse DUOX1–DUOXA1 provide mechanistic insights into enzyme activation and regulation. Nat. Struct. Mol. Biol. 27, 1086–1093 (2020).
Wu, J. X., Liu, R., Song, K. & Chen, L. Structures of human dual oxidase 1 complex in low-calcium and high-calcium states. Nat. Commun. 12, 155 (2021).
Liu, R. et al. Structure of human phagocyte NADPH oxidase in the resting state. eLife 11, e83743 (2022).
Noreng, S. et al. Structure of the core human NADPH oxidase NOX2. Nat. Commun. 13, 6079 (2022).
Warren, J. J., Ener, M. E., Vlcek, A., Winkler, J. R. & Gray, H. B. Electron hopping through proteins. Coord. Chem. Rev. 256, 2478–2487 (2012).
Winkler, J. R. & Gray, H. B. Long-range electron tunneling. J. Am. Chem. Soc. 136, 2930–2939 (2014).
Sumimoto, H. Structure, regulation and evolution of Nox-family NADPH oxidases that produce reactive oxygen species. FEBS J. 275, 3249–3277 (2008).
Lapouge, K., Smith, S. J., Groemping, Y. & Rittinger, K. Architecture of the p40-p47-p67phox complex in the resting state of the NADPH oxidase. J. Biol. Chem. 277, 10121–10128 (2002).
van de Geer, A. et al. Inherited p40phox deficiency differs from classic chronic granulomatous disease. J. Clin. Invest. 128, 3957–3975 (2018).
Lapouge, K. et al. Structure of the TPR domain of p67phox in complex with Rac·GTP. Mol. Cell 6, 899–907 (2000).
Ogura, K. et al. NMR solution structure of the tandem Src homology 3 domains of p47phox complexed with a p22phox-derived proline-rich peptide. J. Biol. Chem. 281, 3660–3668 (2006).
Kami, K., Takeya, R., Sumimoto, H. & Kohda, D. Diverse recognition of non-PxxP peptide ligands by the SH3 domains from p67phox, Grb2 and Pex13p. EMBO J. 21, 4268–4276 (2002).
Wilson, M. I., Gill, D. J., Perisic, O., Quinn, M. T. & Williams, R. L. PB1 domain-mediated heterodimerization in NADPH oxidase and signaling complexes of atypical protein kinase C with Par6 and p62. Mol. Cell 12, 39–50 (2003).
Han, C. H., Freeman, J. L., Lee, T., Motalebi, S. A. & Lambeth, J. D. Regulation of the neutrophil respiratory burst oxidase. Identification of an activation domain in p67phox. J. Biol. Chem. 273, 16663–16668 (1998).
Dahan, I., Smith, S. M. & Pick, E. A Cys-Gly-Cys triad in the dehydrogenase region of Nox2 plays a key role in the interaction with p67phox. J. Leukoc. Biol. 98, 859–874 (2015).
Mizrahi, A., Berdichevsky, Y., Casey, P. J. & Pick, E. A prenylated p47phox-p67phox-Rac1 chimera is a quintessential NADPH oxidase activator: membrane association and functional capacity. J. Biol. Chem. 285, 25485–25499 (2010).
Nisimoto, Y., Motalebi, S., Han, C. H. & Lambeth, J. D. The p67phox activation domain regulates electron flow from NADPH to flavin in flavocytochrome b 558. J. Biol. Chem. 274, 22999–23005 (1999).
Roos, D. et al. Hematologically important mutations: the autosomal forms of chronic granulomatous disease (third update). Blood Cells Mol. Dis. 92, 102596 (2021).
Koker, M. Y. et al. Clinical, functional, and genetic characterization of chronic granulomatous disease in 89 Turkish patients. J. Allergy Clin. Immunol. 132, 1156–1163 (2013).
Stasia, M. J. et al. Molecular and functional characterization of a new X-linked chronic granulomatous disease variant (X91+) case with a double missense mutation in the cytosolic gp91phox C-terminal tail. Biochim. Biophys. Acta 1586, 316–330 (2002).
Rae, J. et al. X-linked chronic granulomatous disease: mutations in the CYBB gene encoding the gp91-phox component of respiratory-burst oxidase. Am. J. Hum. Genet. 62, 1320–1331 (1998).
Milburn, M. V. et al. Molecular switch for signal transduction: structural differences between active and inactive forms of protooncogenic ras proteins. Science 247, 939–945 (1990).
Boog, B. et al. Identification and functional characterization of two novel mutations in the alpha-helical loop (residues 484-503) of CYBB/gp91phox resulting in the rare X91+ variant of chronic granulomatous disease. Hum. Mutat. 33, 471–475 (2012).
Zhen, L., Yu, L. & Dinauer, M. C. Probing the role of the carboxyl terminus of the gp91phox subunit of neutrophil flavocytochrome b558 using site-directed mutagenesis. J. Biol. Chem. 273, 6575–6581 (1998).
Punjani, A. & Fleet, D. J. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Wu, X. et al. Mechanistic insights on heme-to-heme transmembrane electron transfer within NADPH oxydases from atomistic simulations. Front. Chem. 9, 650651 (2021).
Hayward, S. & Lee, R. A. Improvements in the analysis of domain motions in proteins from conformational change: DynDom version 1.50. J. Mol. Graph. Model. 21, 181–183 (2002).
Veevers, R. & Hayward, S. Methodological improvements for the analysis of domain movements in large biomolecular complexes. Biophys. Physicobiol. 16, 328–336 (2019).
Deng, Z. et al. A productive NADP+ binding mode of ferredoxin–NADP+ reductase revealed by protein engineering and crystallographic studies. Nat. Struct. Biol. 6, 847–853 (1999).
Kean, K. M. et al. High-resolution studies of hydride transfer in the ferredoxin:NADP+ reductase superfamily. FEBS J. 284, 3302–3319 (2017).
Lans, I. et al. Theoretical study of the mechanism of the hydride transfer between ferredoxin-NADP+ reductase and NADP+: the role of Tyr303. J. Am. Chem. Soc. 134, 20544–20553 (2012).
Freeman, J. L. & Lambeth, J. D. NADPH oxidase activity is independent of p47phox in vitro. J. Biol. Chem. 271, 22578–22582 (1996).
Koshkin, V., Lotan, O. & Pick, E. The cytosolic component p47phox is not a sine qua non participant in the activation of NADPH oxidase but is required for optimal superoxide production. J. Biol. Chem. 271, 30326–30329 (1996).
Takemoto, D., Tanaka, A. & Scott, B. NADPH oxidases in fungi: diverse roles of reactive oxygen species in fungal cellular differentiation. Fungal Genet. Biol. 44, 1065–1076 (2007).
Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133–138 (2010).
Guo, W., Wang, M. & Chen, L. A co-expression vector for baculovirus-mediated protein expression in mammalian cells. Biochem. Biophys. Res. Commun. 594, 69–73 (2022).
Pedelacq, J. D., Cabantous, S., Tran, T., Terwilliger, T. C. & Waldo, G. S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Scheich, C., Kummel, D., Soumailakakis, D., Heinemann, U. & Bussow, K. Vectors for co-expression of an unrestricted number of proteins. Nucleic Acids Res. 35, e43 (2007).
Li, N. et al. Structure of a pancreatic ATP-sensitive potassium channel. Cell 168, 101–110 (2017).
Yamauchi, A. et al. Location of the epitope for 7D5, a monoclonal antibody raised against human flavocytochrome b558, to the extracellular peptide portion of primate gp91phox. Microbiol. Immunol. 45, 249–257 (2001).
Kim, J. et al. Structure and drug resistance of the Plasmodium falciparum transporter PfCRT. Nature 576, 315–320 (2019).
Pleiner, T., Bates, M. & Görlich, D. A toolbox of anti-mouse and anti-rabbit IgG secondary nanobodies. J. Cell Biol. 217, 1143–1154 (2018).
Guan, C. et al. Structural insights into the inhibition mechanism of human sterol O-acyltransferase 1 by a competitive inhibitor. Nat. Commun. 11, 2478 (2020).
Zhou, M., Diwu, Z., Panchuk-Voloshina, N. & Haugland, R. P. A stable nonfluorescent derivative of resorufin for the fluorometric determination of trace hydrogen peroxide: applications in detecting the activity of phagocyte NADPH oxidase and other oxidases. Anal. Biochem. 253, 162–168 (1997).
Jesaitis, A. J., Riesselman, M., Taylor, R. M. & Brumfield, S. in NADPH Oxidases (eds. Knaus, U. & Leto, T.) 39–59 (Humana Press, 2019).
Patel, A., Toso, D., Litvak, A. & Nogales, E. Efficient graphene oxide coating improves cryo-EM sample preparation and data collection from tilted grids. Preprint at bioRxiv https://doi.org/10.1101/2021.03.08.434344 (2021).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Wang, N. et al. Structural basis of human monocarboxylate transporter 1 inhibition by anti-cancer drug candidates. Cell 184, 370–383 (2021).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Acknowledgements
We thank previous lab member J.-X. Wu for her early work on NOX2; all of the members of the L.C. laboratory for their help; E. Pick for suggestions; J. Han for providing the cDNA for p47, p67 and Rac1; and the National Center for Protein Sciences at Peking University for assistance with negative-stain electron microscopy. Cryo-EM data collection was supported by the Electron Microscopy Laboratory and the Cryo-EM Platform at Peking University; Beijing National Laboratory for Condensed Matter Physics, the Institute of Physics, Chinese Academy of Sciences; and the Beijing branch of Songshan Lake Laboratory for Materials Science with the assistance of X. Li, Z. Guo, C. Qin, X. Pei, X. Hui, G. Wang and D. Sun. Part of the structural computation was also performed on the Computing Platform of the Center for Life Sciences and the High-Performance Computing Platform at Peking University. We thank H. Deng and M. Han in Proteinomics Facility at Technology Center for Protein Sciences, Tsinghua University, for antibody sequencing analysis of 7D5. The work was supported by grants from the Ministry of Science and Technology of China (National Key R&D Program of China, 2022YFA1303000 to L.C.), the National Natural Science Foundation of China (32225027 to L.C.) and the Center For Life Sciences (CLS to L.C.).
Author information
Authors and Affiliations
Contributions
L.C. initiated the project and wrote the manuscript draft. K.S., R.L., X.L. and Y.S. screened various combinations of activators and NOX2–p22 complex. X.L. purified the protein, prepared the cryo-EM sample and collected the cryo-EM data. X.L. and L.C. processed the data and built the model. X.L., R.L. and Y.S. purified the protein sample for the activity assay and measured the activity. All authors contributed to manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks David E. Heppner, Andrea Mattevi, Chuangye Yan and Ming Zhou for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Protein expression and purification.
a, The amino acid sequence of p22-GFP-p67. The colours of each fragment are the same as in Fig. 1a. b, The amino acid sequence of GFPnb-p47-Rac1. c, The amino acid sequence of trimera. d, Size-exclusion chromatography profile of the activated NOX2 complex in nanodisc. Fractions between dashes were used for cryo-EM sample preparation. e, Coomassie brilliant blue-stained SDS–PAGE gel of purified protein complex composed of NOX2, p22-GFP-p67, GFPnb-p47-Rac1, 7D5, and TP1170 in nanodiscs. The experiments were repeated independently twice with similar results. For gel source data, see Supplementary Fig. 1. f, Coomassie brilliant blue-stained SDS–PAGE gel of purified complex between NOX2 and p22-GFP–7D5–TP1170 protein in nanodiscs and trimera mutants. The experiments were repeated independently twice with similar results. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Cryo-EM data collection and image processing.
a, Representative raw micrograph (11,146 in total) of the NOX2 complex in nanodiscs. b, 2D class averages of the NOX2 sample. c, Workflow of the cryo-EM data processing. The processing of the resting NOX2 is shown with a pink background and the processing of the activated NOX2 is shown with a light blue background.
Extended Data Fig. 3 Cryo-EM reconstructions.
a, Gold-standard Fourier shell correlation (GSFSC) curves of consensus refinement of NOX2 in resting state. b, Angular distributions of the consensus refinement of NOX2 in resting state. c, Local resolution distribution of the resting NOX2 complex after consensus refinement. d, Cut-open view of c. e, A 90°-rotated cut-open view of d. f, Local resolution distribution of the resting NOX2 complex after focused refinement. g, Cut-open view of f. h, A 90°-rotated cut-open view of g. i, Structural comparison of the resting NOX2 between the previously determined structure (green, PDB ID: 8GZ3) and the structure determined in this study (pink, PDB ID:8X2L). j, Cryo-EM density map around the FAD- and NADPH-binding site in the resting NOX2 contoured at 9.49σ. FAD is coloured in pink. The empty NADPH-binding site is circled by dash lines. k, Gold-standard Fourier shell correlation curves of consensus refinement of NOX2 in the activated state. l, Angular distributions of the consensus refinement of NOX2 in the activated state. m, Local resolution distribution of the activated NOX2 complex after consensus refinement. n, Cut-open view of m. o, A 90°-rotated cut-open view of n. p, Local resolution distribution of the activated NOX2 complex after focused refinement. q, Cut-open view of p. r, A 90°-rotated cut-open view of q.
Extended Data Fig. 4 Representative densities of NOX2 in the activated state.
a, Unsharpened consensus map of the activated NOX2 complex, contoured at 1.0σ. Regions corresponding to NOX2, p22, p67(1–219), Rac1 and p47(154–281) are coloured as in Fig. 1a. The tails of p22 and Rac1 that insert into the lipid bilayer are denoted by arrows. The zoomed-in window shows the density of the consensus map in a semi-transparent surface with the model of p47 SH3 (154–281) together with the p22 tail fitted in. b, The densities of outer haem, inner haem, FAD-Mg2+, GTP-Mg2+, NADPH, p47(149–158), p67(192–219), NOX2(2–7), NOX2(375–380), NOX2(409–412), NOX2(484–506), NOX2(536–541) and NOX2(567–570) are contoured at 11.77σ, 11.77σ, 16.42σ, 15.34σ, 4.16σ, 6.10σ, 7.56σ, 11.65σ, 5.83σ, 11.65σ, 11.65σ, 11.65σ and 8.12σ, respectively. Mg2+ ions are shown as cyan spheres.
Extended Data Fig. 5 Structural comparisons.
a, Structural comparison of the NOX2 TMD between the resting state (pink, PDB ID: 8X2L) and the activated state (light blue). Conformational changes are indicated by red arrows. The RMSD value is labelled below. b, Structural comparison between p67–Rac1 in the activated NOX2 complex (coloured) and that in the crystal structure (grey, PDB ID:1E96). p67 was used for structural alignment. c, Structural comparison between Rac1 in the activated NOX2 complex (yellow) and that in the crystal structure (grey, PDB ID: IE96). d, Structural comparison between NOX2 DH in the activated state (light blue) and csNOX5 DH (cyan, PDB ID: 5O0X). FBD was used for structural alignment. e, Structural comparison between NOX2 DH in the activated state (light blue) and DUOX1 DH in the high-calcium state (grey, PDB ID: 7D3F). FBD was used for structural alignment.
Extended Data Fig. 6 Sequence alignment.
a, Multiple sequence alignment (MSA) of DH domains of Homo sapiens NOX2 (hsNOX2, UniProt ID: P04839), Homo sapiens NOX1 (hsNOX1, UniProt ID: Q9Y5S8), Homo sapiens NOX3 (hsNOX3, UniProt ID: Q9HBY0), Danio rerio NOX2 (drNOX2, UniProt ID: Q7T2A7), Epichloe festucae NOXA (efNOXA, UniProt ID: Q2PEP0) and Epichloe festucae NOXB (efNOXB, UniProt ID: Q2PEN8). The sequences are downloaded from UniProt and aligned using ClustalX. The result of MSA is further processed in BioEdit. α-helices and β-sheets are shown as cylinders and wide arrows respectively. The conserved NADPH-interacting arginines (R356 in hsNOX2) and the C-terminal phenylalanines (F570 in hsNOX2) are boxed in blue and red, respectively. b, MSA of Homo sapiens p67 (hsP67, UniProt ID: P19878), Homo sapiens NOXA1 (hsNOXA1, UniProt ID: Q86UR1), Danio rerio p67 (drP67, UniProt ID: A2VCY2) and Epichloe festucae NOXR (efNOXR, UniProt ID: A0JC82). Conserved residues at the interfaces are highlighted in bright orange. The residues of the activation domain are presented in a red box. c, MSA of Homo sapiens Rac1 (hsRac1, UniProt ID: P63000), Homo sapiens Rac2 (hsRac2, UniProt ID: P15153), Danio rerio Rac1 (drRac1, UniProt ID: Q7ZSZ9) and Epichloe festucae RacA (efRacA, UniProt ID: A0JC80). Conserved residues at the interfaces are highlighted in bright orange.
Extended Data Fig. 7 The NADPH-binding site and F570 region.
a, The NADPH-binding site in activated NOX2. The structure of NOX2 in the activated state is shown in surface representation coloured the same as in Fig. 1. NADPH is shown as spheres. b, The activity of the NOX2 mutant R356A versus the concentration of NADPH. Data were fitted to the Michaelis–Menten equation. Each data point is shown as an open circle. The 95% confidence interval values are denoted in brackets. Data are mean ± s.d.; n = 3 technical replicates. The experiment was performed independently twice with similar results. c, The activity of the NOX2 mutants on crude cell membrane. DPI, diphenyleneiodonium, an inhibitor of NADPH oxidase. Each data point is shown as an open circle. Data are shown as mean ± standard deviations, n = 3 technical replicates. The experiment was performed independently twice with similar results. d, The activity of the NOX2 mutant F570G versus the concentration of NADPH. Data were fitted to the Michaelis–Menten equation. Each data point is shown as an open circle. The 95% confidence interval values are denoted in brackets. Data are mean ± s.d.; n = 3 technical replicates. The experiment was performed independently twice with similar results. e–h, The local cryo-EM density map of the FAD- and NADPH-binding site in four 3D classes generated from 3DVA of the activated NOX2 data (class 1, e; class 2, f; class 3, g; and class 4, h). FAD and NADPH are coloured in pink and green, respectively. The NADPH-binding site is outlined with dashes.
Extended Data Fig. 8 Working model for NOX2 activation.
The model for phagocyte NADPH oxidase activation. NOX2, p22 and the cytosolic factors are coloured the same as in Fig. 1a. FAD, NADPH, GTP and GDP are presented as spheres. The side chain of F215 between two haems is shown as a hexagon. The side chain of F570 at the C terminus is shown as a hexagon in dashes, indicating its high structural dynamics. The electron transfer pathway is indicated with cyan. In the resting state, the DH domain of NOX2 is in the undocked conformation and NADPH binding is not stable. In the activated state, the binding of cytosolic factors promotes the docking of the DH domain onto the bottom of the TMD and also induces the contraction of the DH domain, stabilizing the binding of NADPH and bringing NADPH close to FAD. The transient displacement of the F570 side chain allows the tight packing of NADPH with FAD, enabling the efficient hydride transfer between them and the subsequent electron transfer process in NOX2.
Supplementary information
Supplementary Figure 1
Uncropped SDS -PAGE gels. Cropped regions shown in Extended Data Fig. 1e, f are indicated with dashed lines.
Supplementary Video 1
Cryo-EM maps of human NOX2 in the activated state. The unsharpened consensus map coloured the same as Fig. 1a is initially shown and then the composite map is shown.
Supplementary Video 2
Structural changes of NOX2 during activation. The morph of NOX2 structure between the resting state and the activated state.
Supplementary Video 3
Structural changes of the DH domain of NOX2 during activation. The morph of the DH domain of NOX2 between the resting state and the activated state.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, X., Shi, Y., Liu, R. et al. Structure of human phagocyte NADPH oxidase in the activated state. Nature 627, 189–195 (2024). https://doi.org/10.1038/s41586-024-07056-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07056-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.