Abstract
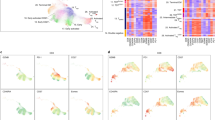
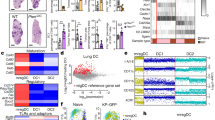
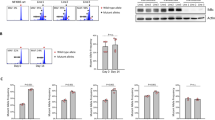
Myeloid cells are known to suppress antitumour immunity1. However, the molecular drivers of immunosuppressive myeloid cell states are not well defined. Here we used single-cell RNA sequencing of human and mouse non-small cell lung cancer (NSCLC) lesions, and found that in both species the type 2 cytokine interleukin-4 (IL-4) was predicted to be the primary driver of the tumour-infiltrating monocyte-derived macrophage phenotype. Using a panel of conditional knockout mice, we found that only deletion of the IL-4 receptor IL-4Rα in early myeloid progenitors in bone marrow reduced tumour burden, whereas deletion of IL-4Rα in downstream mature myeloid cells had no effect. Mechanistically, IL-4 derived from bone marrow basophils and eosinophils acted on granulocyte-monocyte progenitors to transcriptionally programme the development of immunosuppressive tumour-promoting myeloid cells. Consequentially, depletion of basophils profoundly reduced tumour burden and normalized myelopoiesis. We subsequently initiated a clinical trial of the IL-4Rα blocking antibody dupilumab2,3,4,5 given in conjunction with PD-1/PD-L1 checkpoint blockade in patients with relapsed or refractory NSCLC who had progressed on PD-1/PD-L1 blockade alone (ClinicalTrials.gov identifier NCT05013450). Dupilumab supplementation reduced circulating monocytes, expanded tumour-infiltrating CD8 T cells, and in one out of six patients, drove a near-complete clinical response two months after treatment. Our study defines a central role for IL-4 in controlling immunosuppressive myelopoiesis in cancer, identifies a novel combination therapy for immune checkpoint blockade in humans, and highlights cancer as a systemic malady that requires therapeutic strategies beyond the primary disease site.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw RNA-seq data have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE245236. Computed tomography images of patients participating in the clinical trial can be downloaded via Amazon Web Services at the following link: https://himc-project-data.s3.amazonaws.com/lamarche_2023/lamarche_image_data.tar.gz?AWSAccessKeyId=AKIAV3HQ5KORNL3V6W43&Signature=N32aM3ouC5FwYvbMvA7eJJd3V4k%3D&Expires=1698691876. Source data are provided with this paper.
Code availability
Code for generating specific figures will be provided by the corresponding author upon reasonable request.
References
Barry, S. T., Gabrilovich, D. I., Sansom, O. J., Campbell, A. D. & Morton, J. P. Therapeutic targeting of tumour myeloid cells. Nat. Rev. Cancer 23, 216–237 (2023).
Castro, M. et al. Dupilumab efficacy and safety in moderate-to-severe uncontrolled asthma. N. Engl. J. Med. 378, 2486–2496 (2018).
Rabe, K. F. et al. Efficacy and safety of dupilumab in glucocorticoid-dependent severe asthma. N. Engl. J. Med. 378, 2475–2485 (2018).
Simpson, E. L. et al. Two phase 3 trials of dupilumab versus placebo in atopic dermatitis. N. Engl. J. Med. 375, 2335–2348 (2016).
Beck, K. M., Yang, E. J., Sekhon, S., Bhutani, T. & Liao, W. Dupilumab treatment for generalized prurigo nodularis. JAMA Dermatol. 155, 118–120 (2019).
Herbst, R. S., Morgensztern, D. & Boshoff, C. The biology and management of non-small cell lung cancer. Nature 553, 446–454 (2018).
Lavin, Y. et al. Innate immune landscape in early lung adenocarcinoma by paired single-cell analyses. Cell 169, 750–765.e717 (2017).
Leader, A. M. et al. Single-cell analysis of human non-small cell lung cancer lesions refines tumor classification and patient stratification. Cancer Cell 39, 1594–1609.e1512 (2021).
Casanova-Acebes, M. et al. Tissue-resident macrophages provide a pro-tumorigenic niche to early NSCLC cells. Nature 595, 578–584 (2021).
Loyher, P. L. et al. Macrophages of distinct origins contribute to tumor development in the lung. J. Exp. Med. 215, 2536–2553 (2018).
Maier, B. et al. A conserved dendritic-cell regulatory program limits antitumour immunity. Nature 580, 257–262 (2020).
Park, M. D. et al. TREM2 macrophages drive NK cell paucity and dysfunction in lung cancer. Nat. Immunol. 24, 792–801 (2023).
Loschko, J. et al. Absence of MHC class II on cDCs results in microbial-dependent intestinal inflammation. J. Exp. Med. 213, 517–534 (2016).
Karasawa, K. et al. Vascular-resident CD169-positive monocytes and macrophages control neutrophil accumulation in the kidney with ischemia-reperfusion injury. J. Am. Soc. Nephrol. 26, 896–906 (2015).
Lee, P. P. et al. A critical role for Dnmt1 and DNA methylation in T cell development, function, and survival. Immunity 15, 763–774 (2001).
Liu, Z. et al. Fate mapping via Ms4a3-expression history traces monocyte-derived cells. Cell 178, 1509–1525 e1519 (2019).
Horton, B. L. et al. Lack of CD8+ T cell effector differentiation during priming mediates checkpoint blockade resistance in non–small cell lung cancer. Sci. Immunol. 6, eabi8800 (2021).
Burger, M. L. et al. Antigen dominance hierarchies shape TCF1+ progenitor CD8 T cell phenotypes in tumors. Cell 184, 4996–5014.e4926 (2021).
Chihara, N. et al. Induction and transcriptional regulation of the co-inhibitory gene module in T cells. Nature 558, 454–459 (2018).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Naluyima, P. et al. Terminal effector CD8 T cells defined by an IKZF2+IL-7R− transcriptional signature express FcγRIIIA, expand in HIV infection, and mediate potent HIV-specific antibody-dependent cellular cytotoxicity. J Immunol 203, 2210–2221 (2019).
Akimova, T., Beier, U. H., Wang, L., Levine, M. H. & Hancock, W. W. Helios expression is a marker of T cell activation and proliferation. PLoS ONE 6, e24226 (2011).
Thome, M. Multifunctional roles for MALT1 in T-cell activation. Nat. Rev. Immunol. 8, 495–500 (2008).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Passegue, E., Wagner, E. F. & Weissman, I. L. JunB deficiency leads to a myeloproliferative disorder arising from hematopoietic stem cells. Cell 119, 431–443 (2004).
Abram, C. L., Roberge, G. L., Hu, Y. & Lowell, C. A. Comparative analysis of the efficiency and specificity of myeloid-Cre deleting strains using ROSA-EYFP reporter mice. J. Immunol. Methods 408, 89–100 (2014).
Mohrs, M., Shinkai, K., Mohrs, K. & Locksley, R. M. Analysis of type 2 immunity in vivo with a bicistronic IL-4 reporter. Immunity 15, 303–311 (2001).
Hanna, R. N. et al. The transcription factor NR4A1 (Nur77) controls bone marrow differentiation and the survival of Ly6C− monocytes. Nat. Immunol. 12, 778–785 (2011).
Kurotaki, D. et al. Transcription factor IRF8 governs enhancer landscape dynamics in mononuclear phagocyte progenitors. Cell Rep. 22, 2628–2641 (2018).
Zhao, F. et al. S100A9 a new marker for monocytic human myeloid-derived suppressor cells. Immunology 136, 176–183 (2012).
Hegde, S., Leader, A. M. & Merad, M. MDSC: markers, development, states, and unaddressed complexity. Immunity 54, 875–884 (2021).
Seita, J. & Weissman, I. L. Hematopoietic stem cell: self-renewal versus differentiation. Wiley Interdiscip. Rev. Syst. Biol. Med. 2, 640–653 (2010).
Yanez, A. et al. Granulocyte-monocyte progenitors and monocyte-dendritic cell progenitors independently produce functionally distinct monocytes. Immunity 47, 890–902 e894 (2017).
Mastio, J. et al. Identification of monocyte-like precursors of granulocytes in cancer as a mechanism for accumulation of PMN-MDSCs. J. Exp. Med. 216, 2150–2169 (2019).
Nelms, K., Keegan, A. D., Zamorano, J., Ryan, J. J. & Paul, W. E. The IL-4 receptor: signaling mechanisms and biologic functions. Annu. Rev. Immunol. 17, 701–738 (1999).
Jenkins, S. J. et al. Local macrophage proliferation, rather than recruitment from the blood, is a signature of TH2 inflammation. Science 332, 1284–1288 (2011).
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015).
Kwok, I. et al. Combinatorial single-cell analyses of granulocyte-monocyte progenitor heterogeneity reveals an early uni-potent neutrophil progenitor. Immunity 53, 303–318.e305 (2020).
Olsson, A. et al. Single-cell analysis of mixed-lineage states leading to a binary cell fate choice. Nature 537, 698–702 (2016).
Anderson, K. G. et al. Intravascular staining for discrimination of vascular and tissue leukocytes. Nat. Protoc. 9, 209–222 (2014).
Tsutsui, H. et al. The basophil-specific protease mMCP-8 provokes an inflammatory response in the skin with microvascular hyperpermeability and leukocyte infiltration. J. Biol. Chem. 292, 1061–1067 (2017).
Cohen, M. et al. Lung single-cell signaling interaction map reveals basophil role in macrophage imprinting. Cell 175, 1031–1044.e1018 (2018).
Obata, K. et al. Basophils are essential initiators of a novel type of chronic allergic inflammation. Blood 110, 913–920 (2007).
Schultze, J. L., Mass, E. & Schlitzer, A. Emerging principles in myelopoiesis at homeostasis and during infection and inflammation. Immunity 50, 288–301 (2019).
Veglia, F., Sanseviero, E. & Gabrilovich, D. I. Myeloid-derived suppressor cells in the era of increasing myeloid cell diversity. Nat. Rev. Immunol. 21, 485–498 (2021).
Pellefigues, C. et al. Diverse innate stimuli activate basophils through pathways involving Syk and IkappaB kinases. Proc. Natl Acad. Sci. USA 118, e2019524118 (2021).
Gandhi, L. et al. Pembrolizumab plus chemotherapy in metastatic non-small-cell lung cancer. N. Engl. J. Med. 378, 2078–2092 (2018).
Spigel, D. R. et al. Five-year survival outcomes from the PACIFIC trial: durvalumab after chemoradiotherapy in stage III non-small-cell lung cancer. J. Clin. Oncol. 40, 1301–1311 (2022).
Herbst, R. S. et al. Five year survival update from KEYNOTE-010: pembrolizumab versus docetaxel for previously treated, programmed death-ligand 1-positive advanced NSCLC. J. Thorac. Oncol. 16, 1718–1732 (2021).
Choi, H. et al. Transcriptome analysis of individual stromal cell populations identifies stroma–tumor crosstalk in mouse lung cancer model. Cell Rep. 10, 1187–1201 (2015).
Patil, N. S. et al. Intratumoral plasma cells predict outcomes to PD-L1 blockade in non-small cell lung cancer. Cancer Cell 40, 289–300.e284 (2022).
Petitprez, F. et al. B cells are associated with survival and immunotherapy response in sarcoma. Nature 577, 556–560 (2020).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Anderson, N. R., Minutolo, N. G., Gill, S. & Klichinsky, M. Macrophage-based approaches for cancer immunotherapy. Cancer Res. 81, 1201–1208 (2020).
Alam, A. et al. Fungal mycobiome drives IL-33 secretion and type 2 immunity in pancreatic cancer. Cancer Cell 40, 153–167.e111 (2022).
DeNardo, D. G. et al. CD4+ T cells regulate pulmonary metastasis of mammary carcinomas by enhancing protumor properties of macrophages. Cancer Cell 16, 91–102 (2009).
Remark, R. et al. In-depth tissue profiling using multiplexed immunohistochemical consecutive staining on single slide. Sci. Immunol. 1, aaf6925 (2016).
Wang, F. et al. A basophil–neuronal axis promotes itch. Cell 184, 422–440.e417 (2021).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Acknowledgements
We thank members of the Merad laboratory for thoughtful feedback on the manuscript, members of the Center for Comparative Medicine and Surgery at the Icahn School of Medicine at Mount Sinai for animal husbandry, members of the Human Immune Monitoring Center at the Icahn School of Medicine at Mount Sinai for patient sample processing and scRNA-seq assistance, and all patients who participated in this study and their families. M.M. was partially supported by National Institutes of Health (NIH) grants CA257195, CA254104 and CA154947. T.U.M. and the clinical trial presented in Fig. 4 were partially supported by the Feldman Foundation. N.M.L. was supported by the Cancer Research Institute/Bristol Myers Squibb Irvington Postdoctoral Research Fellowship to Promote Racial Diversity (award no. CRI3931). S.H. was supported by the National Cancer Institute predoctoral-to-postdoctoral fellowship K00 CA223043. R.M. was supported by the 2021 AACR-AstraZeneca Immuno-oncology Research Fellowship, Grant Number 21-40-12-MATT. O.C.O. was supported by the Cancer Research Institute Irvington Postdoctoral Research Fellowship (award no. CRI3617). B.D.B. was partially supported by NIH grant U01CA282114. S.G. was partially supported by NIH grants CA224319, DK124165, CA263705 and CA196521.
Author information
Authors and Affiliations
Contributions
N.M.L., M.M. and T.U.M. conceptualized and obtained funding for the project. N.M.L., M.M. and T.U.M. designed experiments. N.M.L., S.H., M.D.P., J.L.B., B.B.M., L.T., I.R.-T, M. Belabed, R.M., A.M.R., R.Z., O.C.O. and E.N. performed experiments. N.M.L., M.D.P., S.H. and P.H. analysed experiments. D.B.D., N.C.R., J.E.G., R.V., N.H. and T.U.M. provided clinical care to patients in the clinical trial. C.H., T.C. and N.V. managed clinical specimens. M. Buckup, I.F., V.R., S.G., E.G.-K., K.M., H.K., F.G., Z.L., E.P., S.K.-S., B.D.B., F.R.H. and B.S.K. provided intellectual input, essential reagents and computational tools. N.M.L. wrote the manuscript. M.M. and T.U.M. edited the manuscript. All authors provided feedback on the manuscript draft.
Corresponding author
Ethics declarations
Competing interests
M.M. and T.U.M. have submitted a patent related to the clinical trial described in this study. M.M. serves on the scientific advisory board and holds stock from Compugen Inc., Myeloid Therapeutics Inc., Morphic Therapeutic Inc., Asher Bio Inc., Dren Bio Inc., Oncoresponse Inc., Owkin Inc., Larkspur Inc. and DEM BIO, Inc. M.M. serves on the scientific advisory board of Innate Pharma Inc., OSE Inc. and Genenta Inc. M.M. receives funding for contracted research from Regeneron Inc. and Boerhinger Ingelheim Inc. T.U.M. has served on Advisory and/or Data Safety Monitoring Boards for Rockefeller University, Regeneron Pharmaceuticals, Abbvie, Bristol Myers Squibb, Boehringer Ingelheim, Atara, AstraZeneca, Genentech, Celldex, Chimeric, Glenmark, Simcere, Surface, G1 Therapeutics, NGMbio, DBV Technologies, Arcus and Astellas, and has research grants from Regeneron, Bristol-Myers Squibb, Merck and Boehringer Ingelheim. B.S.K. is a consultant for Regeneron and Sanofi. D.B.D. sits on the advisory boards of Astrazeneca, Mirati Therapeutics, Summit Therapeutics, G1 Therapeutics and Sanofi, and is a consultant for Sonata Therapeutics. S.G. reports other research funding from Boehringer Ingelheim, Bristol-Myers Squibb, Celgene, Genentech, Regeneron and Takeda not related to this study.
Peer review
Peer review information
Nature thanks Benjamin Izar, Elvira Mass and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 A role for myeloid IL-4Rα expression in NSCLC development.
(a-b) Tumor burden in KP-lung tumor (a) and B16 melanoma metastasis (b) bearing mice treated with isotype or αIL-4 antibodies. Images at 20X. n = 4 mice per group in panel a; n = 5 mice per group in panel b. Representative of two independent experiments. (c) KP lung tumor burden in Il4ra∆DC, Il4ra∆RTM, and Il4ra∆T mice normalized to WT littermate controls (n = 8-24 mice per group). n = 24, 22, 16, 9, 8, 11 mice per group. Pooled from three independent experiments. (d) Number of viable KP cells at indicated seeding values after 5 days of culture in complete media with indicated concentrations of IL-4. Two technical replicates per timepoint. Representative of two independent experiments. (e) IL-4Rα expression in indicated myeloid populations of tumor-bearing Il4ra+/+ and Il4ra∆Ms4a3 mice, normalized to mean MFI of Il4ra+/+ group. n = 15 Il4ra+/+ mice and 9 Il4ra∆Ms4a3 mice for neutrophils; 15 Il4ra+/+ mice and 10 Il4ra∆Ms4a3 mice for monocytes, RTMs, and DCs; 10 Il4ra+/+ mice and 7 Il4ra∆Ms4a3 mice for mo-macs, and 10 Il4ra+/+ mice and 8 Il4ra∆Ms4a3 mice for GMPs. Pooled from three independent experiments. (f) KP tumor burden in Ms4a3-Cre heterozygous mice bearing no floxed allele (n = 11) compared to age-matched WT controls (n = 9). Scale bar=2 mm. One experiment. (g) Number of circulating blood neutrophils in naïve and KP tumor-bearing Il4ra+/+ and Il4ra∆Ms4a3 mice. n = 3, 3, 15, 17 mice per group. Pooled from 3 independent experiments. (h) Number of lung myeloid populations in naïve and KP tumor-bearing Il4ra+/+ and Il4ra∆Ms4a3 mice. n = 3, 3, 7, 7 mice per group. Representative of two independent experiments. Unpaired two-tailed Student’s t-test for all statistical analyses shown. Data are mean ± s.d.
Extended Data Fig. 2 Transcriptional and histological analyses of lung tumors in Il4ra+/+ and Il4ra∆Ms4a3 mice.
(a) Heatmap showing myeloid scRNA-seq clusters (y axis) along with cluster-defining genes (x axis). (b) Gene expression in indicated lung immune clusters of tumor-bearing Il4ra+/+ and Il4ra∆Ms4a3 mice (n = 3 mice per group). One experiment. (c) Average number of CD4 T cells per mm2 of tumor in KP lesions of Il4ra+/+ (n = 13) and Il4ra∆Ms4a3 (n = 10) mice. Representative of three independent experiments. (d) Heatmap showing scRNA-seq clusters from sorted T cells (left) and fine-clustered CD4 (middle) and CD8 (right) T cells (y axis) along with cluster-defining genes (x axis). One experiment. (e) Proportion of the “Exhausted CD8” and “Effector CD8” clusters from panel d among all lung T cells in Il4ra+/+ and Il4ra∆Ms4a3 mice (n = 3 mice per group). One experiment. Data are mean (b), median (c), or mean ± s.d (e). Mann-Whitney test (c) or unpaired two-tailed Student’s t-test (e).
Extended Data Fig. 3 IL-4Rα expression in myeloid populations of tumor-bearing Il4ra∆Cx3cr1 and Il4ra∆S100a8 mice compared to littermate controls.
Representative flow cytometry histograms showing IL-4Rα protein expression in indicated immune populations from tumor-bearing Il4ra∆Cx3cr1 and Il4ra∆S100a8 mice along with WT littermate controls, normalized to mean MFI of WT group. n = 14 Il4ra+/+ and 10 Il4ra∆Cx3cr1 mice; 10 Il4ra+/+ and 10 Il4ra∆S100a8 mice for neutrophils; 11 Il4ra+/+ and 10 Il4ra∆S100a8 mice for monocytes; mo-macs, and RTMs; 10 Il4ra+/+ and 9 Il4ra∆S100a8 mice for GMPs. Pooled from three independent experiments. Grey histogram = Isotype control. Unpaired two-tailed Student’s t-test. Data are mean ± s.d.
Extended Data Fig. 4 No evidence for Th2-biased immune microenvironment in NSCLC.
(a) Expression of IL4 in matched NSCLC tumors and adjacent normal tissue from TCGA database. (b) Expression of indicated genes (y axis) across T cell clusters (x axis) from Leader et al human NSCLC scRNA-seq dataset. (c) IL-4 production after PMA/I stimulation (left) and IL4-eGFP expression in CD4 T cells (middle) and total live cells (right) from lungs of naïve and tumor-bearing mice. n = 5 mice per group for IL-4POS lung CD4 T cells, 10 naïve and 10 tumor-bearing mice for IL4-eGFPPOS CD4 T cells and total IL4-eGFPPOS cells. Representative of two independent experiments. (d) IL-4 production by lung-seeding KP cells in vivo after PMA/I stimulation. One experiment. Unpaired two-tailed Student’s t-test. Data are mean ± s.d.
Extended Data Fig. 5 IL-4 controls myelopoiesis.
(a) Top pathways enriched among DEGs from indicated lung monocyte populations in tumor-bearing Il4ra+/+ and Il4ra∆Ms4a3 mice. (b) Flow cytometry plots showing CD45NEGGFPPOS cells in lung and BM of mice bearing GFP-expressing KP tumors. (c) Number of GMPs per femur of naïve mice or KP tumor-bearing mice treated with isotype or αIL-4 antibodies. n = 5, 7, 7 mice per group. Representative of two independent experiments. (d) Number of Ly6cNEG GMP, GP, and cMoP progenitors in BM of KP tumor-bearing Il4ra+/+ and Il4ra∆Ms4a3 mice. n = 14 Il4ra+/+ and 12 Il4ra∆Ms4a3 mice for Ly6cNEG GMPs and GPs and 6 Il4ra+/+ and 7 Il4ra∆Ms4a3 mice for cMoPs. Pooled from two independent experiments. (e) Methocult differentiation assay of Il4ra+/+ and Il4ra∆Ms4a3 BM. n = 4 technical replicates per condition. Representative of two independent experiments. (f) Representative gating strategy for BM progenitor populations in vehicle and IL-4c treated mice. (g) Total BM cellularity from legs of WT mice treated with vehicle or IL-4c (n = 10 mice per group). Pooled from two independent experiments. (h) Number of GMPs per leg of mice treated with vehicle or IgG1κ antibodies. n = 5 mice per group. One experiment (i) Number of GMP per leg of Il4ra+/+ (WT) or Il4raΔMs4a3 (KO) mice treated with vehicle or IL-4c. n = 3, 3, 4 mice per group. One experiment. (j) Expression of macrophage polarization markers in BMDMs differentiated under indicated conditions. Representative of two independent experiments. (k) scRNA-seq of BM myeloid cells and myeloid progenitors from naïve, IL-4c treated, and KP tumor-bearing mice. Heatmap shows indicated clusters (y axis) and cluster-defining genes (x axis). (l) Total number of genes up or downregulated in each condition relative to naïve in indicated BM scRNA-seq clusters. One experiment. Fisher’s Exact Test (a), Unpaired two-tailed Student’s t-test (c,d,g,h), or One-way ANOVA with post hoc Tukey’s test (i). Data are mean ± s.d. Lin = CD3e, B220, Ter-119, CD11b, Ly6g, NK1.1.
Extended Data Fig. 6 Dynamics of Type 2 granulocytes in BM.
(a) Flow cytometry strategy for BM populations analyzed in Fig. 3a. (b) Percentage of IL4-eGFPPOS cells among indicated BM populations in WT KP tumor-bearing mice. n = 5 mice. Representative of two independent experiments. (c) Number of BM eosinophils and basophils in naïve and tumor-bearing WT mice. n = 5 mice per group. Representative of two independent experiments. (d) Number of CD45-IV negative eosinophils and basophils in lungs of tumor-bearing WT mice. n = 5 mice per group. Representative of two independent experiments. (e) Representative microscopy of BM of tumor-bearing mice showing colocalization of basophils (green) with hematopoietic progenitors (red) (representative of 3 mice). (f) Basophil depletion efficiency in indicated organs after using αFCER1A (left) or αCD200R3 (right) depletion strategies n = 5 isotype and 5 αFCER1A-treated mice; 7 isotype and 5 αCD200R3-treated mice for BM; 6 isotype and 3 αCD200R3-treated mice for blood. Unpaired two-tailed Student’s t-test. Data are mean ± s.d.
Extended Data Fig. 7 Gating Strategies.
(a) Flow cytometry gating strategy for indicated mouse immune populations. Note “basophil alternative gating” using CD200R3 was used in experiments where mice were treated with αFCER1A antibodies. (b) CyTOF gating strategy for human whole blood.
Supplementary information
Supplementary Table 1
Pathway analysis of DEGs between tumour-infiltrating mo-macs and normal lung RTMs in mice and humans.
Supplementary Table 2
Average UMI expression in myeloid and progenitor clusters from bone marrow scRNA-seq, segregated by treatment.
Supplementary Table 3
Clinical metadata for treatment-naive patients with NSCLC profiled by plasma Olink analysis.
Supplementary Table 4
Olink values for plasma from treatment-naive patients with NSCLC and healthy donors.
Supplementary Table 5
Clinical metadata from clinical trial patients.
Supplementary Table 6
Olink values for clinical trial patient plasma.
Supplementary Table 7
Human blood CyTOF panel antibodies.
Supplementary Table 8
Immunohistochemistry antibodies and staining conditions.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
LaMarche, N.M., Hegde, S., Park, M.D. et al. An IL-4 signalling axis in bone marrow drives pro-tumorigenic myelopoiesis. Nature 625, 166–174 (2024). https://doi.org/10.1038/s41586-023-06797-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06797-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.