Abstract
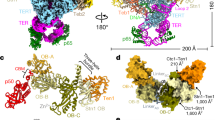
The mammalian DNA polymerase-α–primase (Polα–primase) complex is essential for DNA metabolism, providing the de novo RNA–DNA primer for several DNA replication pathways1,2,3,4 such as lagging-strand synthesis and telomere C-strand fill-in. The physical mechanism underlying how Polα–primase, alone or in partnership with accessory proteins, performs its complicated multistep primer synthesis function is unknown. Here we show that CST, a single-stranded DNA-binding accessory protein complex for Polα–primase, physically organizes the enzyme for efficient primer synthesis. Cryogenic electron microscopy structures of the CST-Polα–primase preinitiation complex (PIC) bound to various types of telomere overhang reveal that template-bound CST partitions the DNA and RNA catalytic centres of Polα–primase into two separate domains and effectively arranges them in RNA–DNA synthesis order. The architecture of the PIC provides a single solution for the multiple structural requirements for the synthesis of RNA–DNA primers by Polα–primase. Several insights into the template-binding specificity of CST, template requirement for assembly of the CST-Polα–primase PIC and activation are also revealed in this study.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The described cryo-EM maps and coordinate files have been deposited in the Electron Microscopy Data Bank and the Protein Data Bank (PDB) under the accession codes EMD-27097 (co-complex–3×TEL-consensus); EMD-27099 (co-complex–3×TEL-local-refined-merged); EMD-27104, PDB ID 8D0B (co-complex–4×TEL-local-refined-merged and consensus maps and model); EMD-27107, PDB ID 8D0K (co-complex–4×TEL–PRIM2C-advanced PIC-local-refined-merged and consensus maps and model); and EMD-27109 (co-complex–6×TEL-local-refined-merged and consensus). The coordinates were assembled and built from individual predicted models from the AlphaFold Protein Structure Database https://alphafold.ebi.ac.uk/.
References
Attali, I., Botchan, M. R. & Berger, J. M. Structural mechanisms for replicating DNA in eukaryotes. Annu. Rev. Biochem. 90, 77–106 (2021).
Baranovskiy, A. G. & Tahirov, T. H. Elaborated action of the human primosome. Genes 8, 62 (2017).
Lim, C. & Cech, T. R. Shaping human telomeres: from shelterin and CST complexes to telomeric chromatin organization. Nat. Rev. Mol. Cell Biol. 22, 283–298 (2021).
Bonnell, E., Pasquier, E. & Wellinger, R. J. Telomere replication: solving multiple end replication problems. Front. Cell Dev. Biol. 9, 668171 (2021).
Pellegrini, L. The Pol alpha-primase complex. Subcell. Biochem. 62, 157–169 (2012).
Braun, K. A., Lao, Y., He, Z., Ingles, C. J. & Wold, M. S. Role of protein-protein interactions in the function of replication protein A (RPA): RPA modulates the activity of DNA polymerase alpha by multiple mechanisms. Biochemistry 36, 8443–8454 (1997).
Casteel, D. E. et al. A DNA polymerase-α·primase cofactor with homology to replication protein A-32 regulates DNA replication in mammalian cells. J. Biol. Chem. 284, 5807–5818 (2009).
Miyake, Y. et al. RPA-like mammalian Ctc1-Stn1-Ten1 complex binds to single-stranded DNA and protects telomeres independently of the Pot1 pathway. Mol. Cell 36, 193–206 (2009).
Surovtseva, Y. V. et al. Conserved telomere maintenance component 1 interacts with STN1 and maintains chromosome ends in higher eukaryotes. Mol. Cell 36, 207–218 (2009).
Lim, C. et al. The structure of human CST reveals a decameric assembly bound to telomeric DNA. Science 368, 1081–1085 (2020).
Chen, L. Y., Redon, S. & Lingner, J. The human CST complex is a terminator of telomerase activity. Nature 488, 540–544 (2012).
Zaug, A. J. et al. CST does not evict elongating telomerase but prevents initiation by ssDNA binding. Nucleic Acids Res. 49, 11653–11665 (2021).
Kelich, J. M., Papaioannou, H. & Skordalakes, E. Pol alpha-primase dependent nuclear localization of the mammalian CST complex. Commun. Biol. 4, 349 (2021).
Wang, F. et al. Human CST has independent functions during telomere duplex replication and C-strand fill-in. Cell Rep. 2, 1096–1103 (2012).
Stewart, J. A. et al. Human CST promotes telomere duplex replication and general replication restart after fork stalling. EMBO J. 31, 3537–3549 (2012).
Mirman, Z. et al. 53BP1–RIF1–shieldin counteracts DSB resection through CST- and Polα-dependent fill-in. Nature 560, 112–116 (2018).
Gu, P. et al. CTC1-STN1 coordinates G- and C-strand synthesis to regulate telomere length. Aging Cell 17, e12783 (2018).
Gu, P. & Chang, S. Functional characterization of human CTC1 mutations reveals novel mechanisms responsible for the pathogenesis of the telomere disease Coats plus. Aging Cell 12, 1100–1109 (2013).
Chen, L. Y., Majerska, J. & Lingner, J. Molecular basis of telomere syndrome caused by CTC1 mutations. Genes Dev. 27, 2099–2108 (2013).
Zhu, W. et al. Mcm10 and And-1/CTF4 recruit DNA polymerase alpha to chromatin for initiation of DNA replication. Genes Dev. 21, 2288–2299 (2007).
Cai, S. W. et al. Cryo-EM structure of the human CST-Polalpha/primase complex in a recruitment state. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-022-00766-y (2022).
Hom, R. A. & Wuttke, D. S. Human CST prefers G-rich but not necessarily telomeric sequences. Biochemistry 56, 4210–4218 (2017).
Bhattacharjee, A., Wang, Y., Diao, J. & Price, C. M. Dynamic DNA binding, junction recognition and G4 melting activity underlie the telomeric and genome-wide roles of human CST. Nucleic Acids Res. 45, 12311–12324 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Baranovskiy, A. G. et al. Mechanism of concerted RNA-DNA primer synthesis by the human primosome. J. Biol. Chem. 291, 10006–10020 (2016).
Kilkenny, M. L., Longo, M. A., Perera, R. L. & Pellegrini, L. Structures of human primase reveal design of nucleotide elongation site and mode of Pol alpha tethering. Proc. Natl Acad. Sci. USA 110, 15961–15966 (2013).
Coloma, J., Johnson, R. E., Prakash, L., Prakash, S. & Aggarwal, A. K. Human DNA polymerase alpha in binary complex with a DNA:DNA template-primer. Sci. Rep. 6, 23784 (2016).
Perera, R. L. et al. Mechanism for priming DNA synthesis by yeast DNA polymerase alpha. Elife 2, e00482 (2013).
Baranovskiy, A. G. et al. Crystal structure of the human primase. J. Biol. Chem. 290, 5635–5646 (2015).
Baranovskiy, A. G. et al. Activity and fidelity of human DNA polymerase alpha depend on primer structure. J. Biol. Chem. 293, 6824–6843 (2018).
Baranovskiy, A. G. et al. Structural basis for inhibition of DNA replication by aphidicolin. Nucleic Acids Res. 42, 14013–14021 (2014).
Baranovskiy, A. G. et al. Insight into the human DNA primase interaction with template-primer. J. Biol. Chem. 291, 4793–4802 (2016).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Lei, M., Podell, E. R. & Cech, T. R. Structure of human POT1 bound to telomeric single-stranded DNA provides a model for chromosome end-protection. Nat. Struct. Mol. Biol. 11, 1223–1229 (2004).
Fan, J. & Pavletich, N. P. Structure and conformational change of a replication protein A heterotrimer bound to ssDNA. Genes Dev. 26, 2337–2347 (2012).
Bhattacharjee, A., Stewart, J., Chaiken, M. & Price, C. M. STN1 OB fold mutation alters DNA binding and affects selective aspects of CST function. PLoS Genet. 12, e1006342 (2016).
He, Y., Song, H., Chan, H. et al. Structure of Tetrahymena telomerase-bound CST with polymerase α-primase. Nature https://doi.org/10.1038/s41586-022-04931-7 (2022).
Goulian, M., Heard, C. J. & Grimm, S. L. Purification and properties of an accessory protein for DNA polymerase alpha/primase. J. Biol. Chem. 265, 13221–13230 (1990).
Goulian, M. & Heard, C. J. The mechanism of action of an accessory protein for DNA polymerase alpha/primase. J. Biol. Chem. 265, 13231–13239 (1990).
Zaug, A.J., Goodrich, K.J., Song, J.J. et al. Reconstitution of a telomeric replicon organized by CST. Nature https://doi.org/10.1038/s41586-022-04930-8 (2022).
Zhang, M. et al. Mammalian CST averts replication failure by preventing G-quadruplex accumulation. Nucleic Acids Res. 47, 5243–5259 (2019).
Punjani, A. & Fleet, D. J. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Zerbe, L. K. & Kuchta, R. D. The p58 subunit of human DNA primase is important for primer initiation, elongation, and counting. Biochemistry 41, 4891–4900 (2002).
Scheres, S. H. et al. Disentangling conformational states of macromolecules in 3D-EM through likelihood optimization. Nat. Methods 4, 27–29 (2007).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Agarkar, V. B., Babayeva, N. D., Pavlov, Y. I. & Tahirov, T. H. Crystal structure of the C-terminal domain of human DNA primase large subunit: implications for the mechanism of the primase-polymerase alpha switch. Cell Cycle 10, 926–931 (2011).
Vaithiyalingam, S., Warren, E. M., Eichman, B. F. & Chazin, W. J. Insights into eukaryotic DNA priming from the structure and functional interactions of the 4Fe-4S cluster domain of human DNA primase. Proc. Natl Acad. Sci. USA 107, 13684–13689 (2010).
Sheaff, R. J. & Kuchta, R. D. Mechanism of calf thymus DNA primase: slow initiation, rapid polymerization, and intelligent termination. Biochemistry 32, 3027–3037 (1993).
Yan, J., Holzer, S., Pellegrini, L. & Bell, S. D. An archaeal primase functions as a nanoscale caliper to define primer length. Proc. Natl Acad. Sci. USA 115, 6697–6702 (2018).
Greci, M. D., Dooher, J. D. & Bell, S. D. The combined DNA and RNA synthetic capabilities of archaeal DNA primase facilitate primer hand-off to the replicative DNA polymerase. Nat. Commun. 13, 433 (2022).
Rames, M., Yu, Y. & Ren, G. Optimized negative staining: a high-throughput protocol for examining small and asymmetric protein structure by electron microscopy. J. Vis. Exp. https://doi.org/10.3791/51087 (2014).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Chen, J., Noble, A. J., Kang, J. Y. & Darst, S. A. Eliminating effects of particle adsorption to the air/water interface in single-particle cryo-electron microscopy: bacterial RNA polymerase and CHAPSO. J. Struct. Biol. X 1, 100005 (2019).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Acknowledgements
We thank T. Tahirov and A. Baranovskiy from the University of Nebraska Medical Center for initial discussion and suggestions in setting up our experiments. We thank T. Cech and A. Zaug from the University of Colorado Boulder for help, discussion and suggestions. We also thank our departmental colleagues S. Butcher and J. Kimble for helpful input, discussion and critical reading of the manuscript. We also thank the members of the laboratory of C.L. for helpful discussions and suggestions. This research was, in part, supported by the Cryo-EM Research Center in the Department of Biochemistry at the University of Wisconsin–Madison and the National Cancer Institute’s National Cryo-EM Facility at the Frederick National Laboratory for Cancer Research under contract 75N91019D00024. Support for this research was provided to C.L. by the National Institutes of Health, the National Institute of General Medical Sciences (R00GM131023) and the University of Wisconsin–Madison, Office of the Vice-Chancellor for Research and Graduate Education with funding from the Wisconsin Alumni Research Foundation and the Department of Biochemistry.
Author information
Authors and Affiliations
Contributions
Q.H. and C.L. designed the expression constructs for recombinant protein production, with support from B.L.L., and Q.H. and X.L. performed insect cell culture and baculovirus production for insect cell infection. Q.H. expressed and purified the recombinant human protein complexes. Q.H., B.L.C. and C.L. conducted the negative-stain EM data collection and sample screening. Q.H. and C.L. prepared the cryo-EM samples for data collection. C.L. processed and analysed the cryo-EM datasets with support from Q.H. C.L. and Q.H. performed model building and refinement. C.L. and X.L. performed the enzyme assays, and C.L., X.L. and Q.H. analysed the data. Q.H., X.L., S.A. and B.L.C. performed the electrophoresis mobility shift assays, and Q.H., S.A., X.L. and B.L.C. analysed the data. C.L. directed the project and designed the experiments with input from Q.H., X.L., B.L.C. and S.A. C.L. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Recombinant human CST and pol-α/primase purification and their biochemical characterization assays.
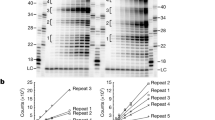
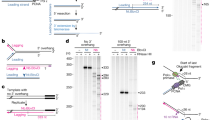
(a) Tandem affinity tags purification of CST and pol-α/primase complexes from baculovirus-infected insect cells. CST heterotrimeric complex was purified by first using Ni-NTA beads to pull down 6xHis-STN1 and 6xHis-TEN1 and then using anti-FLAG beads to pull-down 3xFLAG-CTC1. Pol-α/primase 4-subunit complex was purified by first using Ni-NTA beads to pull down 6xHis-PRIM1, 6xHis-PRIM2, and 6xHis-POLA2 and then using Strep beads to pull-down Strep-POLA1. FT: flowthrough. Trident (GeneTex) high-range prestained protein marker was used in both coomassie-stained denaturing protein gels. The results are reproducible across multiple independent experiments (n > 5). (b) Pull-down assay to characterize direct protein-protein interactions between CST and full-length or truncated (ΔPOLA11-337) pol-α/primase (ΔN-Pol-α/primase). Strep beads were used with pol-α/primase as bait and CST as prey. All input lanes are loaded as 10% of the total sample. Molecular weight markers are annotated for size reference. The pull-down results are reproducible across multiple independent experiments (n = 3). (c) Predicted RNA-DNA primers that are made by pol-α/primase using the repetitive telomeric DNA template in ATP-rich NTP reaction conditions. Primer product sizes are calculated assuming the polymerase reaches the end of the template. (d) Fold-stimulation analysis of direct enzyme assays done at predicted RNA-DNA primer product level (see Fig. 1c). CST does not stimulate pol-α/primase with the 3xTEL template, whether measured at the individual primer product level or total activity (see Fig. 1d). CST can stimulate pol-α/primase using the 6xTEL template. Data are plotted using bars for mean values and error bars for standard deviation as calculated from three independent experiments (n = 3). Data points from each experiment are represented by circle-filled markers in each corresponding bar. The dashed line marked fold-stimulation of one (no stimulation) and values were calculated by dividing the band intensity counts over the corresponding counts from the band and lane without CST added.
Extended Data Fig. 2 Negative-stain electron microscopy 2D classification analysis of CST-pol-α/primase co-complex bound to different telomeric ssDNA template lengths and structures.
(a) Top ten (by distribution) class averages of CST-pol-α/primase preincubated with various lengths of telomeric single-stranded DNA, from 2.5 to 6 repeats of TTAGGG, and structures (ssDNA versus ss/dsDNA). 3xTEL-FB is a DNA template that has a 15 bp hairpin structure at the 5’ end of the GG(TTAGGG)2TTAG overhang. (b) Top ten (by distribution) 2D class averages of each apo-state CST, pol-α/primase, and CST-pol-α/primase co-complex. No new PIC structure was observed in the absence of a template. Instead, individual CST or pol-α/primase 2D class averages were seen in the co-complex analysis. CST-like and pol-α/primase class averages are pointed out by color-coded arrows for visual comparison.
Extended Data Fig. 3 3xTEL, 4xTEL-FB, and 6xTEL co-complexes share the same overall PIC structure.
(a) Comparison of the three co-complexes consensus cryo-EM maps at two different map thresholds. (b) Fourier-shell correlation (FSC) values of the 3xTEL PIC cryo-EM map aligned against the 4xTEL-FB map or 6xTEL map. The FSC values are reported at the level of 0.143.
Extended Data Fig. 4 Pol-α/primase conformation comparison in its apo-state vs. its preinitiation-state.
Pol-α/primase architecture changes from a compact shape to a segregated form upon forming the preinitiation complex (PIC) with a template-bound CST. The APO and PIC structures are depicted as ribbon cartoons. The CST complex and template are hidden in the PIC state to better demonstrate the segregated architecture of the PIC structure – separated into polymerase and primase domains. The lobe boundaries are loosely defined by the dashed lines – black for the polymerase domain and grey for the primase domain. The POLA1 DNA catalytic center is highlighted in orange to illustrate its accessibility upon forming the PIC. POLA1CAT thumb subdomain and PRIM2C domain are not illustrated in the PIC structure because of their flexibility (see main text).
Extended Data Fig. 5 CST and Pol-α/primase interactions.
(a) Overview of TEN1 interactions with primase domain. TEN1 is depicted as surface representation while the primase subunits are drawn as ribbons. (b) TEN1 and STN1N interface sits the PRIM184-101 loop. (c) TEN1 also made contacts with the alpha-helix nine (α9) of the PRIM2N domain. (d) Two hydrogen bonds were identified between TEN1 and POLA2 subunit; TEN1E98 to POLA2R217 and TEN1Q47 to POLA2E277. (e) The loop between residues 547-556 of POLA1 sits on the CTC1OB-G. The POLA1 loop is depicted with heteroatom-coloured sidechains. (f) The template bound by CTC1 forms hydrogen bonds with POLA1 charged residues Q554, N652, and K672. (g) STN1C interacts with POLA1 using negatively charged residues E352 and S355. On the opposite surface, POLA1 uses the positively charged residues H627, H658, and K661. (h, i) The surfaces where POLA1 and STN1C contact has complementary charged residues. In panel h, the POLA1 STN1C-interacting surface, encircled by a white dashed circle, is positively charged. The colour key for the coulombic electrostatic potential is shown at the bottom right of the panel. In panel i, for STN1C, its POLA1-interacting surface, circle by a white dashed circle, is negatively charged. The white dashed circles represent the area of contact for the two protein subunits.
Extended Data Fig. 6 Comparisons of PRIM2N conformation in enzyme apo and preinitiation states and POLA1EXO-CAT domain conformation in the enzyme apo, preinitiation and elongation state.
(a) The PRIM2N structure from our PIC model was compared against the pol-α/primase model in the APO-state (PDB: 5EXR). The PRIM2N small and large subdomains were the same for both enzymatic states but the domain relative position had rotated by about 40°. The structures were aligned using the small subdomain to illustrate the large domain shift. (b) Top-down view of the POLA1EXO-CAT domain at various enzymatic states; apo (APO), preinitiation (PIC), and DNA elongation (ELO). To note, the APO- and ELO-state structures were obtained from either pol-α/primase (PDB: 5EXR) or POLA1 catalytic domain (PDB: 4QCL) alone. The PIC-state was derived from the CST-pol-α/primase co-complex structure. The RNA-DNA primer-template structure of the ELO-state is depicted as a space-filled model. The side-view perspective of each enzyme state is also shown for comparison.
Extended Data Fig. 7 Analysis of PIC-template interactions.
(a) Model of the 15 nt telomeric sequence built into template cryo-EM density (shown as semi-transparent grey coloured volume). The model is depicted as an atomic model that is color-coded T=blue, A=red, and G=green. The modelled sequence is 5’- TAGGGTTAGGGTTAG – 3’ (15mer-TTAG). (b) Model-to-map cross-correlation (CC) analysis reported at per residue level and chain average. The experimental binding affinities of CST to these templates are provided. The template 15mer-TAGG and 15mer-AGGG sequences are 5’- AGGGTTAGGGTTAGG – 3’ and 5’- GGGTTAGGGTTAGGG – 3’ respectively. (c) Independent experiments and quantification for Fig. 2b. See Supplementary Table 1 for template sequences. (d) Overview of protein-DNA interactions involved in template binding. (e, f) CTC1OB-F interactions with the template at the binding site 1. (g) CTC1OB-G residues binding to the template at binding site two are shown. (h) POLA1 residues binding to the template at binding site 2 are shown. (i) STN1 W89 base stacks with A14 of the template. This is supported by T13-T14 base stacking. (j) Other protein-DNA interactions between STN1N and template at binding site 3. STN1 R139 hydrogen bonds with T13 phosphate backbone and STN1 Y141 base stacks with G15 aromatic base. (k) The 15 nt DNA template model is depicted as a deep pink atomic model. Sites 1 and 2 are positively charged while site 3 is more neutral. Site 1 DNA-binding residues are mostly made from CTC1OB-F, site 2 residues are from CTC1OB-G and POLA1, and site 3 residues are from the STN1N domain. The electrostatic potential colour key is in units of kcal/(mol·e) at 298 K.
Extended Data Fig. 8 CST-pol-α/primase PIC structure is incompatible with CST decamer formation at early oligomerization states.
(a) Dimension comparison of the PIC model to the CST decamer model (ten copies of CST; PDB: 6W6W). (b) The PIC primase domain is not sterically hindered by the CST inverted dimer and thus could exist. However, its polymerase domain directly clashes with CST dimer formation. (c) The PIC primase and polymerase domains clash with CST tetramer formation. As in panel b, the polymerase domain clashes with the dimer formation, while the primase domain is now in the way of the fourth CST in the CST tetramer model. Subunits are coloured as in Fig. 1 for all panels.
Extended Data Fig. 9 Polymerase domain swivels on top of the template-bound CST.
(a) Three-dimensional (3D) variable analysis (3DVA) principal component analysis revealed a mode that showed the polymerase domain rotating about on the top of the template-bound CST. The first and last frames of the PCA mode are shown here as pink and navy blue cryo-EM maps. The cryo-EM maps are filtered to a resolution of 8 Å. The polymerase domain swivelling motion is seemly controlled by the STN1C domain and rotation angle restrained by the STN1 linker. (b) The corresponding components in A depicted on the PIC atomic model. CTC1 is represented as surface and coloured light grey. The rest of the PIC are drawn as ribbons. The color scheme follows Fig. 1.
Extended Data Fig. 10 Cryo-EM heterogeneity analysis reveals template-bound human primase structure at an advanced preinitiation state.
(a) The 4xTEL-FB PIC structure was masked with a PRIM1-PRIM2C for particle subtraction and 3D classification to isolate a subset of particles that allowed higher-resolution 3D reconstruction of the PRIM2C domain. An eight classes 3D classification step without image alignment was used to sieve out a conformation (~13.8%) that has an extra EM density in front of the PRIM1 subunit. This subtracted particles subset (32,550 particles) was reverted to their original images before subjecting to 3D refinement to obtain a 4.3 Å cryo-EM map. The resolved PRIM2C domain density is circled in green dashed lines. (b) Two crystal structure variants of the PRIM2C domain are docked into the cryo-EM density that was resolved in Fig. 5 and panel a. The all-helical variant (PDB: 3Q36) is a better fit than the β-sheet-containing version (PDB: 3L9Q), as can be seen in the region pointed out by the black arrow. (c) PRIM1 and PRIM2C have minimal contacts except for two loop regions; PRIM1192-200 and PRIM2376-385. (d) PRIM1 and PRIM2 interactions with the template region into the primase crevice. (e) The five additional template residues were built based on the resolved primase density. The last three nucleotides are annotated. (f) The PRIM1 and PRIM2C domains form a crevice that the template sits in for RNA primer synthesis. The crevice has the PRIM1 RNA catalytic center facing the template. (g) The 3’ end of the extended 20 nt template model in the PIC structure connects seamlessly with the template model in the previously solved RNA-DNA primer-template-bound PRIM2C crystal structure (PDB: 5F0Q).
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Tables 1 and 2.
Supplementary Video 1
3DVA analysis of the polymerase domain swivelling on top of a template-bound CST in a PIC structure.
Supplementary Video 2
3DVA analysis of the PRIM2(C) domain sampling the space in front of PRIM1 and POLA1 and the template.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, Q., Lin, X., Chavez, B.L. et al. Structures of the human CST-Polα–primase complex bound to telomere templates. Nature 608, 826–832 (2022). https://doi.org/10.1038/s41586-022-05040-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05040-1
This article is cited by
-
A mechanistic model of primer synthesis from catalytic structures of DNA polymerase α–primase
Nature Structural & Molecular Biology (2024)
-
CST–polymerase α-primase solves a second telomere end-replication problem
Nature (2024)
-
Human primase hangs on the primer–template and Polα to facilitate primer termination
Nature Structural & Molecular Biology (2023)
-
DNA-binding mechanism and evolution of replication protein A
Nature Communications (2023)
-
Molecular choreography of primer synthesis by the eukaryotic Pol α-primase
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.