Abstract
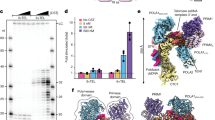
Telomerase adds G-rich telomeric repeats to the 3′ ends of telomeres1, counteracting telomere shortening caused by loss of telomeric 3′ overhangs during leading-strand DNA synthesis (‘the end-replication problem’2). Here we report a second end-replication problem that originates from the incomplete duplication of the C-rich telomeric repeat strand (C-strand) by lagging-strand DNA synthesis. This problem is resolved by fill-in synthesis mediated by polymerase α-primase bound to Ctc1–Stn1–Ten1 (CST–Polα-primase). In vitro, priming for lagging-strand DNA replication does not occur on the 3′ overhang and lagging-strand synthesis stops in a zone of approximately 150 nucleotides (nt) more than 26 nt from the end of the template. Consistent with the in vitro data, lagging-end telomeres of cells lacking CST–Polα-primase lost 50–60 nt of telomeric CCCTAA repeats per population doubling. The C-strands of leading-end telomeres shortened by around 100 nt per population doubling, reflecting the generation of 3′ overhangs through resection. The measured overall C-strand shortening in the absence of CST–Polα-primase fill-in is consistent with the combined effects of incomplete lagging-strand synthesis and 5′ resection at the leading ends. We conclude that canonical DNA replication creates two telomere end-replication problems that require telomerase to maintain the G-rich strand and CST–Polα-primase to maintain the C-strand.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Greider, C. W. & Blackburn, E. H. Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell 43, 405–413 (1985).
Lingner, J., Cooper, J. P. & Cech, T. R. Telomerase and DNA end replication: no longer a lagging strand problem. Science 269, 1533–1534 (1995).
Griffith, J. D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
Doksani, Y., Wu, J. Y., de Lange, T. & Zhuang, X. Super-resolution fluorescence imaging of telomeres reveals TRF2-dependent T-loop formation. Cell 155, 345–356 (2013).
Watson, J. D. Origin of concatemeric T7 DNA. Nat. New Biol. 239, 197–201 (1972).
Olovnikov, A. M. A theory of marginotomy. The incomplete copying of template margin in enzymic synthesis of polynucleotides and biological significance of the phenomenon. J. Theor. Biol. 41, 181–190 (1973).
Greider, C. W. & Blackburn, E. H. Telomeres, telomerase and cancer. Sci. Am. 274, 92–97 (1996).
Zhao, Y. et al. Telomere extension occurs at most chromosome ends and is uncoupled from fill-in in human cancer cells. Cell 138, 463–475 (2009).
Hirai, Y., Masutomi, K. & Ishikawa, F. Kinetics of DNA replication and telomerase reaction at a single-seeded telomere in human cells. Genes Cells 17, 186–204 (2012).
Wu, P., Takai, H. & de Lange, T. Telomeric 3′ overhangs derive from resection by Exo1 and Apollo and fill-in by POT1b-associated CST. Cell 150, 39–52 (2012).
Wang, F. et al. Human CST has independent functions during telomere duplex replication and C-strand fill-in. Cell Rep. 2, 1096–1103 (2012).
Dai, X. et al. Molecular steps of G-overhang generation at human telomeres and its function in chromosome end protection. EMBO J. 29, 2788–2801 (2010).
Stewart, J. A., Wang, Y., Ackerson, S. M. & Schuck, P. L. Emerging roles of CST in maintaining genome stability and human disease. Front. Biosci. 23, 1564–1586 (2018).
Chow, T. T., Zhao, Y., Mak, S. S., Shay, J. W. & Wright, W. E. Early and late steps in telomere overhang processing in normal human cells: the position of the final RNA primer drives telomere shortening. Genes Dev. 26, 1167–1178 (2012).
Yeeles, J. T., Deegan, T. D., Janska, A., Early, A. & Diffley, J. F. Regulated eukaryotic DNA replication origin firing with purified proteins. Nature 519, 431–435 (2015).
Yeeles, J. T. P., Janska, A., Early, A. & Diffley, J. F. X. How the eukaryotic replisome achieves rapid and efficient DNA replication. Mol. Cell 65, 105–116 (2017).
Soudet, J., Jolivet, P. & Teixeira, M. T. Elucidation of the DNA end-replication problem in Saccharomyces cerevisiae. Mol. Cell 53, 954–964 (2014).
Aria, V. & Yeeles, J. T. P. Mechanism of bidirectional leading-strand synthesis establishment at eukaryotic DNA replication origins. Mol. Cell 73, 199–211 (2019).
Jones, M. L., Aria, V., Baris, Y. & Yeeles, J. T. P. How Pol α-primase is targeted to replisomes to prime eukaryotic DNA replication. Mol. Cell 83, 2911–2924.e16 (2023).
Lim, C. J. & Cech, T. R. Shaping human telomeres: from shelterin and CST complexes to telomeric chromatin organization. Nat. Rev. Mol. Cell Biol. 22, 283–298 (2021).
Cai, S. W. & de Lange, T. CST–Polα/Primase: the second telomere maintenance machine. Genes Dev. 37, 555–569 (2023).
Goulian, M., Heard, C. J. & Grimm, S. L. Purification and properties of an accessory protein for DNA polymerase alpha/primase. J. Biol. Chem. 265, 13221–13230 (1990).
Goulian, M. & Heard, C. J. The mechanism of action of an accessory protein for DNA polymerase alpha/primase. J. Biol. Chem. 265, 13231–13239 (1990).
Lim, C. J. et al. The structure of human CST reveals a decameric assembly bound to telomeric DNA. Science 368, 1081–1085 (2020).
Lue, N. F. Evolving linear chromosomes and telomeres: a C-strand-centric view. Trends Biochem. Sci 43, 314–326 (2018).
Gao, H., Cervantes, R. B., Mandell, E. K., Otero, J. H. & Lundblad, V. RPA-like proteins mediate yeast telomere function. Nat. Struct. Mol. Biol. 14, 208–214 (2007).
Miyake, Y. et al. RPA-like mammalian Ctc1–Stn1–Ten1 complex binds to single-stranded DNA and protects telomeres independently of the Pot1 pathway. Mol. Cell 36, 193–206 (2009).
Surovtseva, Y. V. et al. Conserved telomere maintenance component 1 interacts with STN1 and maintains chromosome ends in higher eukaryotes. Mol. Cell 36, 207–218 (2009).
Cai, S. W. et al. Cryo-EM structure of the human CST–Polα/primase complex in a recruitment state. Nat. Struct. Mol. Biol. 29, 813–819 (2022).
He, Q. et al. Structures of the human CST–Polα-primase complex bound to telomere templates. Nature 608, 826–832 (2022).
Zaug, A. J., Goodrich, K. J., Song, J. J., Sullivan, A. E. & Cech, T. R. Reconstitution of a telomeric replicon organized by CST. Nature 608, 819–825 (2022).
Feng, X., Hsu, S. J., Kasbek, C., Chaiken, M. & Price, C. M. CTC1-mediated C-strand fill-in is an essential step in telomere length maintenance. Nucleic Acids Res. 45, 4281–4293 (2017).
Takai, H. et al. A POT1 mutation implicates defective telomere end fill-in and telomere truncations in Coats plus. Genes Dev. 30, 812–826 (2016).
McElligott, R. & Wellinger, R. J. The terminal DNA structure of mammalian chromosomes. EMBO J. 16, 3705–3714 (1997).
Damm, K. et al. A highly selective telomerase inhibitor limiting human cancer cell proliferation. EMBO J. 20, 6958–6968 (2001).
Larrivée, M., LeBel, C. & Wellinger, R. J. The generation of proper constitutive G-tails on yeast telomeres is dependent on the MRX complex. Genes Dev. 18, 1391–1396 (2004).
Baird, D. M., Rowson, J., Wynford-Thomas, D. & Kipling, D. Extensive allelic variation and ultrashort telomeres in senescent human cells. Nat. Genet. 33, 203–207 (2003).
Tesmer, V. M., Brenner, K. A. & Nandakumar, J. Human POT1 protects the telomeric ds-ss DNA junction by capping the 5′ end of the chromosome. Science 381, 771–778 (2023).
Hockemeyer, D., Daniels, J. P., Takai, H. & de Lange, T. Recent expansion of the telomeric complex in rodents: two distinct POT1 proteins protect mouse telomeres. Cell 126, 63–77 (2006).
Anderson, B. H. et al. Mutations in CTC1, encoding conserved telomere maintenance component 1, cause Coats plus. Nat. Genet. 44, 338–342 (2012).
Guilliam, T. A. & Yeeles, J. T. P. Reconstitution of translesion synthesis reveals a mechanism of eukaryotic DNA replication restart. Nat. Struct. Mol. Biol. 27, 450–460 (2020).
Komura, J. & Riggs, A. D. Terminal transferase-dependent PCR: a versatile and sensitive method for in vivo footprinting and detection of DNA adducts. Nucleic Acids Res. 26, 1807–1811 (1998).
Hanish, J. P., Yanowitz, J. L. & de Lange, T. Stringent sequence requirements for the formation of human telomeres. Proc. Natl Acad. Sci. USA 91, 8861–8865 (1994).
Smogorzewska, A., Karlseder, J., Holtgreve-Grez, H., Jauch, A. & de Lange, T. DNA ligase IV-dependent NHEJ of deprotected mammalian telomeres in G1 and G2. Curr. Biol. 12, 1635–1644 (2002).
Harley, C. B., Futcher, A. B. & Greider, C. W. Telomeres shorten during ageing of human fibroblasts. Nature 345, 458–460 (1990).
Acknowledgements
The authors thank C. Price for providing the CTC1fl/fl HCT116 cell line; members of the de Lange laboratory for comments on this work; V. Risca and A. Mansisidor for advice on the PCR method; J. C. Zinder for the CTC1 antibody; and S. Cai for purified human RPA. This work is supported by grants from the NCI (R35CA210036), the NIA (RO1AG016642), and the BCRF to T.d.L.; a grant from the NCI to H.T. (5R50CA243771-02); and a grant to J.T.P.Y. from the Medical Research Council (no. MC_UP_1201/12).
Author information
Authors and Affiliations
Contributions
H.T. and T.d.L. conceived and designed the study. V.A. and J.T.P.Y. designed and performed the in vitro replication experiments. All in vivo experiments were executed by H.T. with help from P.B. in the early stages of the project. T.d.L. and J.T.P.Y. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Joachim Lingner and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Changing views of the end-replication problem.
a, The first version of the end-replication problem as perceived by Watson5 and Olovnikov6 after it was shown that Okazaki fragments carry RNA primers at their 5′ ends. In this end-replication problem, the 5′ ended strand of the lagging-strand DNA synthesis product is shorter than the parental strand. b, Telomerase was suggested to solve the end-replication problem by extending the 3′ ends before DNA replication7. c, When it had become clear that telomeres ended in a 3′ overhang, it was argued that the end-replication problem involved the inability of leading-strand DNA synthesis to recreate this part of the telomeric DNA2. Therefore, the end-replication problem involved the shortening of the G-rich strand at leading-end telomeres. Telomerase can solve the leading-end replication problem by acting after DNA replication (and 5′ end resection) as shown in Fig. 1a. If the last Okazaki fragments start on the 3′ overhang (as shown), no sequence loss will occur at lagging-end telomeres. The end-replication problem was therefore “no longer a lagging strand problem” (quote from ref. 2).
Extended Data Fig. 2 Analysis of replisome-mediated DNA synthesis at the end of linear DNA templates.
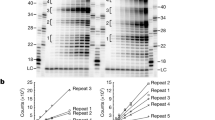
a, Schematic of the replication templates used for in vitro budding yeast DNA replication and the anticipated reaction products of origin-dependent replication. b and c, Analysis of the data presented in Fig. 1d (b) and 1f (c). Product length was determined using a standard curve derived from the molecular weight standards. Replication product intensity was normalised by dividing each data point by the amplitude of the most intense product for a given condition. d, Schematic illustrating the terminal TTAGGG repeats that are used to initiate Okazaki fragment synthesis during in vitro DNA replication. e, Replication reaction performed and analyzed as in Fig. 1f but with the concentration of Polα-primase increased from 20 nM to 80 nM. To enable clear visualization of the Nb.BbvCI products, the Nt.BbvCI product bands are saturated. f, Comparison of Nb.BbvCI digested replication products in the absence of RNase HII at 20 nM (Fig. 1d) and 80 nM (e) Polα-primase. Data were processed as described for (b) and (c).
Extended Data Fig. 3 Analysis of lagging-strand priming within TTAGGG repeats.
a, Schematic illustrating the TTAGGG repeat containing templates and the anticipated products of Nt./Nb.BbvCI digestion. By linearizing the plasmids with different restriction enzymes, the TTAGGG repeats are located either at the end of the template, or 121 bp from the end of the template. b, Denaturing 5% polyacrylamide/urea gel analysis of a replication reaction on the templates shown in (a) analyzed as described in Fig. 1d. To enable clear visualization of the Nb.BbvCI products, the Nt.BbvCI product bands are saturated. c, Comparison of Nb.BbvCI digested replication products in the absence of RNase HII from (b). The asterix marks the presence of an RNase HII insensitive replication product. The position of the template strand TTAGGG repeats are illustrated below the traces. d, Comparison of Polα-primase in the yeast and human replisomes (from reference19). For clarity only Polα-primase (pink), the CMG helicase (grey) and DNA (black) are shown. The shortest distance between the point of strand separation and the primase active site are illustrated (dashed white lines).
Extended Data Fig. 4 CsCl-gradient separation of leading- and lagging-end telomeres from CTC1F/F cells lacking telomerase activity.
a, Schematic of the experimental approach to separate telomeres replicated by leading- and lagging-strand DNA synthesis. b, TRAP assay on extracts from CTC1F/F cells from which hTERT is deleted (in bulk) using CRISPR/Cas9 (sgTERT). Cells treated with a Luciferase sgRNA are used as the control. Cell equivalents are indicated above the lanes. 250H: Heat-inactivated extract (250 cells). Positive control sample (pos cntrl) provided by the manufacturer. I: internal PCR control. c, Examples of slot blots to detect telomeric DNA in fractions from CsCl gradients. Pooled fractions used for in gel 3′ overhang assays are indicated at the top.
Extended Data Fig. 5 Validation of the PCR overhang assay (Method I).
a, Schematic of the telomere model substrate generated by ligating a telomeric 3′ overhang to a HindIII/BglII fragment of pSXneo1.6T2AG3. b, PCR overhang assay on the model telomeric substrate before and after E. coli 3′ exonuclease ExoI treatment to remove the 3′ overhang. c, PCR overhang assays performed with synthetic telomeric oligos containing 5 to 20 TTAGGG repeats. Tel10 -Lig: Tel10 oligo processed in parallel but without ligation to the adaptor. d, Relative signal intensity of 6 independent PCR overhang assays as in (c) were plotted. The signal intensity of Tel5 oligo in each experiment was set at 1. e, In-gel hybridization assay showing that telomerase expression does not strongly alter 3′ overhang lengths when CTC1-profient cells and that the ExoI digestion used in Fig. 2e worked. Genomic DNAs from CTC1F/F cells with or without sgTERT and BIBR1532 treatment were treated with E. coli ExoI as indicated and digested with MboI/AluI for the in-gel overhang assay shown. Left: TelC probe detecting the overhang signals. Right: same gel probed with TelC after in situ denaturation to detect the total telomeric DNA for normalization.
Extended Data Fig. 6 Overhang measurement with ExoI digestion (Method II).
a, Top: Telomere model substrate with or without overhang (as in Extended Data Fig. 5a) with predicted fragment lengths indicated. Bottom: Telomere model substrate with and without overhang was treated with or without ExoI, separated by alkaline agarose gel electrophoresis and detected by in-gel hybridization with the TelC probe. Sizes of the detected fragments were calculated based on the molecular weight marker. Shortening of the model telomere substrate G-strand by ExoI is shown below the gel. b, Example of determination of G-strand shortening by ExoI treatment. DNAs from cells with or without depletion of telomerase (sgTERT+BIBR) were treated with ExoI as indicated and subsequently digested with MboI and AluI. DNAs were separated on an alkaline agarose gel and the gel was dried and hybridized with TelC. Changes in G-strand length after ExoI treatment are indicated below the lanes. The MW markers are generated by restriction digestion of pSXneo1.6T2AG3 (see Extended Data Fig. 5a and Methods) and fragments containing TTAGGG repeats are detected with TelC. c, Graph of migration (pixel position) versus the MW of the telomeric marker in lane 3 and lane 12 of the gel in (b) generated by scanning of the gel using Fiji software (ImageJ, ver2.0). The equation of the curve is used to convert migration (pixel #) to telomere lengths. d, Measured lengths of the telomere markers in lane 1 and 2 was calculated using the position of the fragment (pixel #) in the gel and the equation generated from standard curve in (c) and compared to the expected length. e, Overlay of the scan profiles of G-strand telomeric signal from DNA treated with or without ExoI shown in (b). Line graphs shift to the right (lower MW) in DNAs treated with ExoI compared to mock-treated DNAs. e, Graph showing average 3′ overhang lengths and SDs determined by Method II on 6 biological replicates. ns, not significant based on unpaired two-sided t-test.
Extended Data Fig. 7 Calculation of C-strand shortening at lagging-end telomeres.
a and b, Calculation of the C-strand loss at lagging-end telomeres in cells lacking CTC1 and telomerase activity. The schematic in (a) shows the changes at leading- and lagging-end telomeres over three rounds of replication assuming 97 nt (average of 110 and 84 nt) resection of the leading-end products as determined in Fig. 2i. Sequence loss at the lagging-end C-strand is calculated to be 54 nt based on the 3′ overhang length of 201 nt (average of 228 and 173) for lagging-end telomeres after 5 divisions without CTC1 (Fig. 2i) and applying the equation in (b). Note that the 97 nt loss at the leading-end telomeres is constant, whereas the loss at the lagging-end telomeres increases with each division. c, Predicted overhang length changes in 5 successive rounds of replication after CTC1 deletion from telomerase-deficient cells as depicted in (a). Note that the value for the lagging-end overhangs is predicted to plateau (at ~205 nt) while the leading-end overhangs are constant at 97 nt. Therefore, the average overhang length is predicted to plateau at ~150 nt which represents a doubling of the length (starting length 74 nt before CTC1 deletion) consistent with the two-fold increase in the relative overhang signal observed in Fig. 3c.
Extended Data Fig. 8 C-strand shortening in HCT116 cells lacking CTC1.
a–c, Measurement of C-strand shortening CTC1F/F cells with or without 4-OHT treatment based on alkaline gel analysis of the telomeric G- and C-strands as in Fig. 4b–d.
Extended Data Fig. 9 Schematic illustrating the dynamics of the C- and G-strand in CTC1-deficient cells expressing telomerase.
a, Schematic showing the changes of G- and C-strand telomere length over three rounds of replication in telomerase-proficient cells lacking CTC1. G-strand sequences added by telomerases are depicted with a blue line. The numbers given are as follows (based on Fig. 2i,j). X: nucleotides added by telomerase; 74 nt: initial 3′ overhang length; 54 nt: loss of sequences due to incomplete lagging-strand synthesis; 97 nt: loss of sequences due to resection of the C-strand at leading-end telomeres. b, Equation describing changes in average G-strand length after each round of replication (n) relative to initial G-strand telomere length. c, Increase in average G-strand length with cell divisions calculated using the equation in (b) and assuming that telomerase adds an average of 240 nt (X) to each end. d, Modeled changes in the length of the G-strand in Ctc1-deficient cells based on the initial G-strand length of 3.24 kb. To determine X, the value for telomerase extension was increased stepwise by 10 nt, plotted and the plot was compared to the actual G-strand changes obtained in Fig. 4d. When X is 240 nt, the model plot is a close fit with the in vivo rate of G-strand elongation in Fig. 4d.
Supplementary information
Supplementary Figure 1
Uncropped images of blots for Figs. 1–4 and Extended Data Figs. 3–6 and 8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Takai, H., Aria, V., Borges, P. et al. CST–polymerase α-primase solves a second telomere end-replication problem. Nature 627, 664–670 (2024). https://doi.org/10.1038/s41586-024-07137-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07137-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.