Abstract
The COVID-19 pandemic is the third outbreak this century of a zoonotic disease caused by a coronavirus, following the emergence of severe acute respiratory syndrome (SARS) in 20031 and Middle East respiratory syndrome (MERS) in 20122. Treatment options for coronaviruses are limited. Here we show that clofazimine—an anti-leprosy drug with a favourable safety profile3—possesses inhibitory activity against several coronaviruses, and can antagonize the replication of SARS-CoV-2 and MERS-CoV in a range of in vitro systems. We found that this molecule, which has been approved by the US Food and Drug Administration, inhibits cell fusion mediated by the viral spike glycoprotein, as well as activity of the viral helicase. Prophylactic or therapeutic administration of clofazimine in a hamster model of SARS-CoV-2 pathogenesis led to reduced viral loads in the lung and viral shedding in faeces, and also alleviated the inflammation associated with viral infection. Combinations of clofazimine and remdesivir exhibited antiviral synergy in vitro and in vivo, and restricted viral shedding from the upper respiratory tract. Clofazimine, which is orally bioavailable and comparatively cheap to manufacture, is an attractive clinical candidate for the treatment of outpatients and—when combined with remdesivir—in therapy for hospitalized patients with COVID-19, particularly in contexts in which costs are an important factor or specialized medical facilities are limited. Our data provide evidence that clofazimine may have a role in the control of the current pandemic of COVID-19 and—possibly more importantly—in dealing with coronavirus diseases that may emerge in the future.
Similar content being viewed by others
Main
COVID-19 in humans has a broad clinical spectrum that ranges from mild to severe manifestations, with a mortality rate of about 2% worldwide4. The high transmissibility of SARS-CoV-2 has been attributed to a substantial proportion of mild or asymptomatic infections5,6.The genetically diverse coronavirus family—which is currently thought to comprise four genera (Alphacoronavirus, Betacoronavirus, Gammacoronavirus and Deltacoronavirus), infects birds and a variety of mammals (including bats)7. Within the past two decades, there have been three major outbreaks of disease caused by coronaviruses in humans. SARS-CoV was first detected in Guangdong (China) in 2002 and—with the aid of commercial air travel—spread rapidly and globally, and caused more than 8,000 cases with a 10% mortality rate1. In 2012, MERS-CoV may have evolved and spread from bats to humans through an intermediate host (camels); it has caused over 1,700 cases with a mortality rate of almost 40%, and—similar to SARS-CoV—air travel has enabled the global spread of this virus to 27 countries2,8.
There are no widely available specific antiviral therapies for coronaviruses in humans9. Remdesivir has been shown to exhibit pan-coronavirus inhibitory potential10, and has been granted approval by the US Food and Drug Administration (FDA) for the treatment of COVID-19 on the basis of the reduced time taken to recover following treatment with this drug11. However, the therapy is suboptimal (particularly for patients with severe COVID-19), and it can only be administered intravenously to patients in hospital12,13. Thus, the development of additional therapeutic options is urgent, as is the establishment of combinatorial regimens to improve efficacy and reduce the potential for the emergence of drug-resistant variants. For example, the combination of IFNβ-1b, lopinavir–ritonavir and ribavirin has been shown to have beneficial antiviral effects in a randomized clinical trial14.
In efforts to accelerate the development of therapies for COVID-19, a library of known drugs that encompassed approximately 12,000 clinical-stage or FDA-approved small molecules was previously profiled15. This study focused on the antiviral mechanisms of action and in vivo efficacy of clofazimine, which is an FDA-approved molecule that was discovered as an anti-tuberculosis drug and was later used for the treatment of leprosy16. The effective concentration of clofazimine against SARS-CoV-2 in vitro (half-maximal effective concentration (EC50) of 0.31 μM in Vero E6 cells) is clinically achievable with a single dose of 200 mg d−1 (peak serum concentration (Cmax) of 0.86 μM)17. A sufficiently high concentration of clofazimine, as multiples of its 90% effective concentration (EC90) in different cell lines (EC90 of 0.81 to 2.35 μM), is known to accumulate in plasma and lung tissue after several doses18. Here we report the protective effect of clofazimine against infection with SARS-CoV-2 or MERS-CoV in primary human cells and in a hamster model. In addition, clofazimine is an affordable treatment for patients with COVID-19, which may help to relieve the acute healthcare burden created by the pandemic in lower-income countries19.
Clofazimine is a pan-coronavirus inhibitor
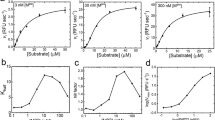
Clofazimine reduced MERS-CoV replication in Vero E6 cells, with an EC50 of 1.48 ± 0.17 μM (Extended Data Fig. 1a). Immunofluorescence staining for MERS-CoV nucleocapsid (N) protein, combined with flow cytometry analysis, revealed the effective suppression of virus infection in human hepatocellular Huh7 cells upon clofazimine treatment (Extended Data Fig. 1b). We then characterized the antiviral activity of clofazimine in two cell lines that are more physiologically relevant to coronavirus diseases: cardiomyocytes derived from human embryonic stem cells and human primary small-airway epithelial cells, which robustly support SARS-CoV-220 and MERS-CoV21 replication, respectively. Clofazimine treatment reduced the SARS-CoV-2 titre up to more than 3 log10, and the MERS-CoV titre by 2 log10 (Fig. 1a). Next, we assessed the antiviral activity of clofazimine in an ex vivo lung culture system and found that clofazimine potently antagonized viral replication in human lung tissues that reflect the primary site of SARS-CoV-2 and MERS-CoV replication (Fig. 1b). To explore whether clofazimine confers cross-protection against other epidemic and seasonal coronaviruses, we performed viral-load reduction assays for SARS-CoV, hCoV-229E and hCoV-OC43 in corresponding cell lines that support virus replication. Viral yields in cell culture supernatants were decreased by about 2 log10 in Vero E6 cells infected with SARS-CoV, by about 4 log10 in human embryonic lung fibroblasts infected with hCoV-229E and by around 3 log10 in monkey BSC1 cells infected with hCoV-OC43 (Fig. 1c). Clofazimine showed negligible cytotoxicity in the matching cell lines as described above for pan-coronavirus inhibitory evaluation (Extended Data Fig. 1c). Overall, clofazimine exhibited broad-spectrum anti-coronavirus efficacy, and antagonized both SARS-CoV-2 and MERS-CoV replication in human primary cell and ex vivo lung models.
a, Clofazimine inhibited SARS-CoV-2 (multiplicity of infection (MOI) of 0.1) replication in human primary cardiomyocytes (left) and MERS-CoV (MOI of 1.0) replication in human primary small-airway epithelial cells (right). Cell lysates were collected for viral load determination; viral genome copies were determined relative to ACTB. Data are mean ± s.d., n = 3 biological replicates. Two-tailed Student’s t-test. b, Ex vivo human lung tissues were infected with SARS-CoV-2 or MERS-CoV-2 followed by treatment with clofazimine (10 μM), remdesivir (10 μM) or DMSO (0.1%). Supernatants were collected for the quantification of viral titre by plaque assay. Data are mean ± s.d., n = 5 biological replicates. Two-tailed Student’s t-test. PFU, plaque-forming unit. c, Antiviral activity of clofazimine against SARS-CoV (MOI of 0.01, 48 h after infection) (left), HCoV-229E (MOI of 0.001, 72 h after infection) (middle) and HCoV-OC43 (MOI of 0.001, 72 h after infection) (right) in Vero E6, human embryonic lung fibroblasts and BSC1 cells, respectively. Viral load in the cell culture supernatant was quantified by qPCR with reverse transcription (RT–qPCR). Data are mean ± s.d., n = 3 biological replicates. One-way ANOVA followed by Sidak’s post-test. All experiments were repeated twice for confirmation. All statistical analyses are compared with the DMSO-treated group (0 μM). ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05.
Effects on SARS-CoV-2 life cycle
We first evaluated the antiviral activity of clofazimine using a time-of-drug addition assay in a single infectious cycle. Treatment with clofazimine during inoculation strongly inhibited SARS-CoV-2 infection, which indicates that clofazimine inhibits viral entry. Clofazimine also blocked SARS-CoV-2 infection at a post-entry step, as evidenced by an observed reduction in viral replication when clofazimine was added at 5 h after infection (Extended Data Fig. 2a). To further evaluate the effect of clofazimine on viral entry, we used vesicular stomatitis virus (VSV)-based spike glycoprotein (S) pseudotyped virions. Clofazimine treatment reduced the infectivity of both SARS-CoV S and SARS-CoV-2 S pseudotyped virions in Vero E6 cells, but did not affect MERS-CoV S pseudotyped virus particles (Fig. 2a). We then examined a series of events to determine the precise step of SARS-CoV-2 entry that is blocked by clofazimine, and excluded possible effects of clofazimine on: (1) the cell-surface expression of the SARS-CoV-2 receptor angiotensin-converting enzyme 2 (ACE2) or the MERS-CoV receptor dipeptidyl peptidase 4 (DPP4) (Extended Data Fig. 2b) and (2) the disruption of binding between ACE2, or another attachment factor (heparan sulfate)22, and SARS-CoV-2 S (Extended Data Fig. 2c). Using a cell membrane fusion assay mediated by SARS-CoV-2 S23,24, we found that clofazimine inhibited cell fusion activity mediated by S. Specifically, we observed that fusion between effector cells that express S and target cells were effectively reduced after clofazimine addition, in a dose-dependent manner(Fig. 2c, Extended Data Fig. 2d). To confirm whether clofazimine also inhibits post-entry steps of viral replication, we evaluated the effect of clofazimine on the production of viral RNA by electroporating in vitro-transcribed viral RNA into Vero E6 cells, which bypasses clofazimine-mediated inhibition on the entry process and directly measures RNA synthesis (Extended Data Fig. 2e). As expected, remdesivir reduced the synthesis of negative-stranded RNA in a dose-dependent manner (blue symbols in Fig. 2c). Viral RNA levels were also reduced by about 1–1.5 log10 in cells treated with clofazimine (red symbols in Fig. 2c). However, no substantial effect was observed on the translation of electroporated GFP mRNA (Extended Data Fig. 2f). We also found that clofazimine had no obvious effect on the activity of main protease and papain-like protease, which are responsible for the cleavage of viral polypeptide (half-maximal inhibitory concentration (IC50) of over 100 μM) (Extended Data Fig. 2g). Because a critical step in the replication and transcription of SARS-CoV-2 requires the formation of a stable complex of the SARS-CoV-2 helicase (nsp13) and holo-RNA-dependent RNA polymerase (RdRp)25, we investigated the effect of clofazimine on this step, and found that clofazimine inhibited the unwinding activity of nsp13 using either a double-stranded DNA or double-stranded RNA substrate (Fig. 2d). However, clofazimine did not block the primer elongation activity that is executed by holo-RdRp (Extended Data Fig. 2h). Collectively, these results demonstrated that clofazimine exerts antiviral activity by targeting several steps in SARS-CoV-2 replication, including interference with S-mediated cell fusion as well as viral helicase activity.
a, VSV-based pseudotyped viral particle assay. Vero E6 cells that were pretreated with indicated compounds were infected with the SARS-CoV-2 S- (left), SARS-CoV S- (middle) or MERS-CoV S- (right) pseudotyped particles. Luciferase signals were quantified at 24 h after infection. Mean ± s.e.m., n = 6 independent experiments. One-way ANOVA followed by Dunnett post-test. MDL28170, a known coronavirus entry inhibitor15, was used as a positive control. RLU, relative light unit. b, Clofazimine inhibits membrane fusion mediated by SARS-CoV-2 S. Vero cells cotransfected with SARS-CoV-2 S and EGFP plasmids were added to the nontransfected Vero cells to induce membrane fusion. Confocal images were acquired at 48 h after transfection. Scale bars, 400 μm. Representative images selected from a pool of images captured in two independent experiments. c, In vitro-transcribed viral RNA replication assay. Remdesivir or clofazimine were added at the indicated doses. Negative-stranded RNA was then quantified at 12 h after electroporation. Error bars are s.e.m., n = 3 independent experiments. Two-tailed Student’s t-test. d, Titration of double-stranded (ds)DNA and dsRNA unwinding activity of the SARS-CoV-2 helicase (nsp13) by clofazimine using an assay based on fluorescence resonance energy transfer. The black curve represents a positive-control inhibitor (ranitidine bismuth citrate) using a DNA-based substrate. Data are mean ± s.d., n = 3 biological replicates. The experiments were repeated twice for confirmation. All statistical analyses are compared with the DMSO-treated or non-treatment group. ***P < 0.001, **P < 0.01, *P < 0.05; NS, not significant.
Transcriptional analysis after treatment
We used RNA sequencing (RNA-seq) to profile transcriptome-wide changes in human Caco-2 cells infected with SARS-CoV-2, in which clofazimine exhibited antiviral potency comparable to that of remdesivir (Extended Data Fig. 3a). We performed transcriptional analysis on Caco-2 cells that were infected or uninfected and that were subsequently treated with clofazimine or dimethyl sulfoxide (DMSO; mock treatment) for 3 or 6 h (Extended Data Fig. 3b). At 3 h after infection, clofazimine treatment caused an overall transcriptome shift towards the mock-treatment control group (Extended Data Fig. 3c), which corroborates the hypothesis that the drug interferes with the early stage of the virus life cycle. At 6 h after infection, there were 607 and 448 genes upregulated and downregulated, respectively, by SARS-CoV-2 infection (Supplementary Tables 1, 2). The RNA level of more than 90% of these genes was reverted by clofazimine treatment, which indicates that clofazimine treatment abrogates the transcriptomic changes that are induced by SARS-CoV-2 infection. This is consistent with a principal components analysis of the dataset, which showed that treatment with clofazimine for 6 h after infection caused a marked shift towards the mock treatment (Extended Data Fig. 3c, d). Clofazimine treatment for 6 h in the absence of infection upregulated the expression of transcription factors that are critical for the immediate–early cellular response, including those of the AP-1, SMAD and MAFF families (Extended Data Fig. 3e). When clofazimine was applied to infected cells for 6 h, we observed an enrichment of upregulated genes that are associated with innate-immunity-related pathways, including MAPK, interleukin and TNF responses (Extended Data Figs. 3f, 4a, b). These results suggest that clofazimine may also rewire the transcriptional landscape to prime the innate-immunity-related pathways.
Prophylactic and therapeutic activities in vivo
Previous pharmacokinetics studies have revealed that, in humans, co-administration of a 200 mg dose of clofazimine with food results in a Cmax of 0.41 mg l−1 (equivalent to 0.86 μM) at 8 h. Here we used a golden Syrian hamster model26 to determine the in vivo antiviral efficacy of clofazimine. Because administration of clofazimine with a high-fat meal provides better bioavailability27, we delivered the drug through the oral route using corn oil as vehicle (Fig. 3a).
a, The prophylactic treatment used oral administration of clofazimine, given at 3, 2 and 1 days before infection, followed by virus challenge at day 0; therapeutic administration of clofazimine was performed at 1, 2, and 3 days after infection. Tissue samples were collected at indicated days after infection. Remdesivir was included as a control in the therapeutic regimen. b, c, Daily body weights of the hamsters under prophylactic (b) or therapeutic (c) treatment. Data are mean ± s.e.m., n = 3 hamsters per group. Two-way ANOVA followed by Sidak’s post-test for prophylactic groups, and two-way ANOVA followed by Tukey’s post-test for therapeutic groups. P value indicates clofazimine versus vehicle group. d, Viral yield in hamster lung tissue, after prophylactic (n = 5 hamsters per group) or therapeutic treatment (n = 11 hamsters (each remdesivir and clofazimine group), n = 13 hamsters (vehicle group)), which were collected at 4 days after infection and titrated by plaque assays. e, Hamster nasal washes collected on 4 days after infection were subjected to live virus titration by plaque assays (n = 5 hamsters per group). f, Hamster faeces were freshly collected at 4 days after infection and subjected to SARS-CoV-2 viral copy detection by RT–qPCR assays (n = 5 hamsters per group). For purposes of statistical analysis, a value of about 10–100 was assigned for any data point below the detection limit (dotted line). g, The IL-6 level in hamster serum was quantified at 4 days after infection. n = 5 hamsters for each prophylaxis group and n = 10 hamsters for each therapeutic group. All data in d–g are mean ± s.d. Two-tailed Student’s t-test for prophylaxis groups and one-way ANOVA followed by Dunnett’s post-test for therapeutic groups, comparing with the vehicle group (black symbols). ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05, NS, not significant.
After clofazimine administration in a prophylactic regimen, we observed a reduction in the loss of body weight after infection with SARS-CoV-2. When clofazimine was given after virus challenge, the time taken for body weight to return to pre-infection levels was shortened (Fig. 3b, c). At 4 days after infection (when viral loads peaked and there were substantial histopathological changes), clofazimine decreased the virus plaque-forming units in lung tissues by about 1–2 log10 (Fig. 3d). We consistently confirmed the suppression of viral genome copies of SARS-CoV-2 in the lungs in hamsters treated with clofazimine (Extended Data Fig. 5a). In addition, we evaluated the capacity of clofazimine to diminish SARS-CoV-2 shedding in nasal wash and faecal samples. At 4 days after infection and after remdesivir or clofazimine therapy, we did not detect any significant (P > 0.05, one-way analysis of variance (ANOVA)) reduction in infectious virus titre in the hamster nasal wash, whereas an average of a twofold reduction was achieved with prophylactic administration of clofazimine (Fig. 3e). Irrespective of whether clofazimine was administered prophylactically or therapeutically, we found substantially lower viral loads in faeces at 4 days after infection compared with the vehicle-treated group. By contrast, remdesivir did not alleviate virus shedding from gastrointestinal tract (Fig. 3f). Overall, we demonstrate in vivo in a hamster model that clofazimine antagonizes SARS-CoV-2 replication in the lung and reduces virus shedding in faeces.
To ascertain whether clofazimine improves disease prognosis, we determined serum levels of IL-6, which have previously been shown to correlate with respiratory failure and adverse clinical outcomes28. In general, we detected substantially decreased levels of serum IL-6 in both the clofazimine-treated and remdesivir-treated groups (Fig. 3g). It has previously been reported that clofazimine may inhibit lymphocyte function in cell culture29. To investigate this in the context of an immune response induced by SARS-CoV-2, we collected sera from hamsters at 14 days after infection and found that the levels of antibody responses triggered in vehicle-treated and clofazimine-treated groups were similarly high, which indicates that clofazimine does not induce detectable suppression of the humoral immune response of B lymphocytes (Extended Data Fig. 5b). We also performed RNA-seq of hamster lung tissues (Extended Data Fig. 6a). Out of the 73 upregulated genes after prophylactic treatment with clofazimine (Supplementary Table 3), 34 (46.6%) were mapped to biological processes related to the immune response (Extended Data Fig. 6b). Of these 34 genes, 13 were mapped to the ‘leukocyte differentiation’ Gene Ontology category—including two major histocompatibility complex class-II molecules, H2-Aa and H2-Ab1 (which are labelled with an asterisk in Extended Data Fig. 6c). In addition, transcription factors—including Fos, Junb and Egr1—were also upregulated, which is consistent with our transcriptomic analysis of Caco-2 cells treated with clofazimine (Extended Data Fig. 6c). Notably, most of the immune-response-related genes were not changed after administering clofazimine to uninfected hamsters, which indicates that clofazimine priming of the host response is dependent on SARS-CoV-2 infection and is not likely to result in the undue activation of the host immune system (Extended Data Figs. 6d, e).
To determine the severity of lung damage, we performed histological examination of lung tissues that were stained with haematoxylin and eosin. The lungs of mock-treated control hamsters showed severe pathological changes, which were evidenced by large areas of consolidation as well as cell infiltrations in the endothelium of blood vessels, and peribronchiolar regions. By contrast, clofazimine- or remdesivir-treated lungs exhibited an improved morphology and milder infiltrations (Extended Data Fig. 7a, b). Taken together, our results suggest that clofazimine conferred protection against SARS-CoV-2 challenge in the hamster model by reducing virus replication and associated inflammatory dysregulation in the host.
Antiviral synergy with remdesivir
Remdesivir is considered the current standard of care for the treatment of COVID-19. We found that co-application of clofazimine and remdesivir affects SARS-CoV-2 replication in a manner that extends beyond the additive activity predicted by the Bliss independence model (maximal Bliss synergy score of 44.28), which indicates that these two drugs operate in a synergistic antiviral relationship (Extended Data Fig. 8a, b). The addition of 1.25 μM clofazimine (fourfold EC50) in an in vitro assay resulted in a nearly 20-fold decrease in the concentrations of remdesivir required to inhibit viral replication by 90% (Extended Data Fig. 8c) and—importantly—the combination of drugs did not elicit additional cellular cytotoxicity (Extended Data Fig. 8d).
To explore the antiviral synergy of these two drugs in vivo and to recapitulate the scenario in which most patients with COVID-19 will be treated after the onset of disease, we gave SARS-CoV-2-infected hamsters oral clofazimine and intraperitoneal remdesivir together: the first doses were given 24 h after virus challenge. Experimentally, we gave 1.5 mg per kg body weight remdesivir (tenfold lower than the standard dosing) as well as a reduced clofazimine dose (15 mg per kg bodyweight, corresponding to a 100 mg dose in humans) (Fig. 4a). An improvement in weight loss was achieved in the combinatorially treated group at 3 days after infection, compared with the groups treated with vehicle control or low-dose remdesivir alone (Fig. 4b). Monotherapy using low-dose remdesivir caused a marginal reduction of virus titre in the lungs. However, the additional application of clofazimine not only exhibited potent synergy in terms of viral load (Fig. 4c), but also restricted virus replication in the focal bronchiolar epithelial cells from spreading to alveolar areas (Extended Data Fig. 9a). The antiviral synergy also suppressed virus shedding in the nasal wash (Fig. 4d), which was not achievable with therapeutic treatments using remdesivir or clofazimine individually (Fig. 3e). Immunofluorescence staining of hamster nasal turbinate showed that the combinatorial treatment effectively diminished N antigen expression in the epithelium, providing further support for the synergy between the drugs (Extended Data Fig. 9b, c). Taken together, the antiviral synergy between low-dose remdesivir and clofazimine effectively improved viral control, leading to reduced body-weight loss, a suppressed pulmonary virus titre and nasal virus shedding, as well as decreased drug dosages.
a, Experimental design of in vivo study. Oral clofazimine (15 mg per kg body weight) and/or intraperitoneal remdesivir were administrated at the indicated days after infection. Vehicle-treated control hamsters received oral administration of corn oil and an intraperitoneal injection of 2% DMSO in 12% sulfobutyl ether-β-cyclodextrin (SBE-β-CD). The standard dose of remdesivir is 15 mg per kg body weight, a low-dose remdesivir is 1.5 mg per kg; the low dose was used in the combined treatment. b, Daily body weights of the hamsters (n = 5 hamsters per group). Data are mean ± s.d. Two-way ANOVA followed by Tukey’s test. c, Virus in lung tissue was titrated by plaque assay. One-way ANOVA followed by Dunnett’s post-test (n = 5 hamsters per group). d, Virus titre in nasal wash was determined by plaque assays. One-way ANOVA followed by Dunnett’s post-test, compared with the combinatorial group (n = 5 hamsters per group). **P < 0.01, *P < 0.05, NS, not significant.
Discussion
Clofazimine was first used to treat leprosy in 1969 and gained FDA approval in 199630. It is an orally bioavailable drug that is included in the WHO (World Health Organization) Model List of Essential Medicines. It is generally well-tolerated, and is recommended as a WHO group-C drug for the treatment of multidrug-resistant and extensively drug-resistant tuberculosis31.
In SARS-CoV-2 infection, a delayed innate immune response may result in an uncontrolled cytokine storm32,33. The effect of clofazimine on rewiring the transcriptional landscape of the cell towards an antiviral status may be important in the COVID-19 setting, and understanding the contribution of this activity to in vivo disease amelioration can provide insights into its potential to improve viral control through the enhancement of innate immune activities. Elucidation of how clofazimine treatment may balance the regulation of innate and adaptive immune responses will be important to understanding its potential clinical efficacy. Further modification of clofazimine to improve its pharmacokinetics and randomized clinical trials to ascertain its clinical performance are warranted.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized, and investigators were not blinded to allocation during experiments and outcome assessment.
Cells and viruses
The different cell lines and primary cells that were used in this study were chosen according to their high sensitivity to replication of a corresponding coronavirus. Human hepatoma Huh7 (JCRB, 0403) cells, human colon Caco-2 cells (ATCC, HTB-37), monkey Vero E6 cells (ATCC, CRL-1586) and monkey kidney BSC-1 cells (ATCC, CCL-26) were maintained in DMEM culture medium supplemented with 10% heat-inactivated FBS, 50 U ml−1 penicillin and 50 μg ml−1 streptomycin. Human embryonic lung fibroblasts were developed in-house. Human primary small-airway epithelial cells (ATCC, PCS-301-010) were cultured with airway epithelial cell basal medium, according to the manufacturer’s protocol. Ventricular cardiomyocytes were differentiated from the human embryonic stem cell HES2 (ESI) maintained in mTeSR1 medium (STEMCELL Technologies)34. In brief, HES2 cells were dissociated with Accutase (Invitrogen) into single-cell suspensions on day 0. Cells were seeded on low-attachment culture vessels (Corning) and cultured in mTeSR1 medium supplemented with 40 μg ml−1 Matrigel, 1 ng ml−1 BMP4 (Invitrogen) and 10 μM Rho kinase inhibitor (ROCK) (R&D) under hypoxic environment with 5% O2. From day 1 to 3, cells were cultured in StemPro34 SFM (Invitrogen) with 50 μg ml−1 ascorbic acid (AA) (Sigma), 2 mM Gluta-MAX (Invitrogen), 10 ng ml−1 BMP4, and 10 ng ml−1 human recombinant activin-A (Invitrogen). From day 4 to day 7, 5 μM Wnt inhibitor IWR-1 (Tocris) was added. From day 8 to day 14, cells were cultured under normoxia in RPMI 1640 medium (Invitrogen) supplemented with 2 mM Gluta-MAX, 1× B-27 supplement (Invitrogen) and 50 μg ml−1 AA. The cells were then dissociated with Accutase and seeded as a monolayer in desired culture vessels for 3 days before infections. The SARS-CoV-2 HKU-001a strain (GenBank accession number: MT230904) was isolated from the nasopharyngeal aspirate specimen of a patient who was laboratory-confirmed to have COVID-19 in Hong Kong26. The SARS-CoV-2 isolate USA-WA1/2020 was deposited by the Centers for Disease Control and Prevention, and obtained through BEI Resources. The MERS-CoV strain (HCoV-EMC/2012) was a gift from R. Fouchier. The archived clinical strains of SARS-CoV (HCoV-OC43 and HCoV-229E) were obtained from the Department of Microbiology of the University of Hong Kong (HKU)35. All experiments involving live SARS-CoV, SARS-CoV-2 and MERS-CoV followed the approved standard operating procedures of the biosafety level 3 facility at the University of Hong Kong and Sanford Burnham Prebys Medical Discovery Institute, as was previously described15.
Antiviral evaluation in human ex vivo lung tissues
Human lung tissues for ex vivo studies were obtained from patients who were undergoing surgical operations at Queen Mary Hospital (Hong Kong), as previously described36. The donors and/or parents of the donor gave written consent, as approved by the Institutional Review Board of the HKU/Hospital Authority Hong Kong West Cluster (UW13-364). The freshly obtained lung tissues were processed into small rectangular pieces and were rinsed with advanced DMEM/F12 medium (Gibco) supplemented with 2 mM of HEPES (Gibco), 1× GlutaMAX (Gibco), 100 U ml−1 penicillin and 100 μg ml−1 streptomycin. The specimens were infected with SARS-CoV-2 HKU-001a or MERS-CoV with an inoculum of 1 × 106 PFU ml−1 at 500 μl per well. After 2 h, the inoculum was removed, and the specimens were washed 3 times with PBS. The infected human lung tissues were then cultured in 1 ml of advanced DMEM/F12 medium with 2 mM HEPES (Gibco), 1× GlutaMAX (Gibco), 100 U ml−1 penicillin, 100 μg ml−1 streptomycin, 20 μg ml−1 vancomycin, 20 μg ml−1 ciprofloxacin, 50 μg ml−1 amikacin and 50 μg ml−1 nystatin. Supernatants were collected at 24 h after inoculation for plaque assays.
Antiviral assessment in a hamster model of SARS-CoV-2 infection
Male and female Syrian hamsters, aged 6–10 weeks old, were obtained from the Chinese University of Hong Kong Laboratory Animal Service Centre through the HKU Centre for Comparative Medicine Research. The hamsters were kept in biosafety level 2 housing and given access to standard pellet feed and water ad libitum, as was previously described26. All experimental protocols were approved by the Animal Ethics Committee in the HKU (CULATR) and were performed according to the standard operating procedures of the biosafety level 3 animal facilities (reference code: CULATR 5370-20). Experimentally, each hamster was intranasally inoculated with 105 PFU of SARS-CoV-2 in 100 μl PBS under intraperitoneal ketamine (200 mg per kg) and xylazine (10 mg per kg) anaesthesia.
To simulate the prescribed human dosage (200 mg per day), an equivalent hamster dose of 25 mg per kg per day was converted on the basis of body surface area. Specifically, 25 mg per kg (hamster) × 0.13 (conversion factor) = 3.25 mg per kg (human equivalent dose); and a 60-kg human requires 3.25 mg per kg × 60 kg = 195 mg clofazimine per day. Prophylactic treatment used oral administration of clofazimine given 3, 2 and 1 days before infection (25 mg per kg each time), followed by virus challenge at day 0; therapeutic post-exposure and oral administration of clofazimine (Sigma-Aldrich, C8895) was performed on 1, 2 and 3 days after infection (25 mg per kg each time) with the first dosage given at 24 h after inoculation. Clofazimine was delivered using corn oil (Sigma-Aldrich, C8267) as vehicle. Remdesivir was included as a positive-control drug and dosed at 15 mg per kg via the intraperitoneal route on the basis of its effective dosage in SARS-CoV-infected mice10. Remdesivir (15 mg per kg, MedChemExpress) was prepared as 100 mg ml−1 stock in DMSO and further diluted using 12% SBE-β-CD before intraperitoneal injection. Hamsters receiving pure corn oil (oral) and 2% DMSO in 12% SBE-β-CD (intraperitoneal) were used as the vehicle-control group. Hamsters were killed at 4 days after infection for virological and histopathological analyses. Viral yield in the lung tissue homogenates and/or faeces was detected by plaque assay and/or RT–qPCR. Nasal washes were collected to examine virus shedding from the respiratory tract. Hamsters were treated with isoflurane lightly, and then 200 μl of PBS was injected into one nasal opening while turbid wash was collected from the other one without any blood contamination. The nasal wash was filtered through 0.22-μm filters before being subjected to plaque assay. An ELISA kit was used to determine the amount of IL-6 in the hamster sera at 4 days after infection, according to the manufacturer’s recommendations (ELISAGenie, HMFI0001). Tissue pathology of infected hamsters was examined by haematoxylin and eosin (H&E) staining in accordance with the established protocol37. On day 14 after infection, an enzyme immunoassay was used to determine the antibody titre of hamster sera against SARS-CoV-2 N antigen. In brief, 96-well immune plates (Nunc) were coated with 100 μl per well (0.1 μg per well) of SARS-CoV-2 N in 0.05 M NaHCO3 (pH 9.6) overnight at 4 °C. After blocking, 100 μl of heat-inactivated serum samples was serial-diluted before being added to the wells and incubated at 37 °C for 1 h. The attached antibodies were detected using horseradish-peroxidase-conjugated rabbit anti-hamster IgG antibody (Thermo Fisher Scientific; A18895; 1:2,000). The reaction was developed by adding diluted 3,3′,5,5′-tetramethylbenzidine single solution (TMB) (Invitrogen) and stopped with 0.3 N H2SO4. The optical density was read at 450/620 nm using a microplate reader.
RNA-seq analysis
FASTQ files from RNA-seq were quality-examined by FastQC (v.0.11.7) (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Reads were processed by cutadapt to remove reads with low quality and to trim adapters. For RNA-seq on Caco-2 cells, trimmed reads were aligned to hg38 reference genome and NCBI SARS-CoV-2 reference genome (NC_045512.2) using TopHat38(v.2.1.1); and RNA-seq data from hamster lung tissues were mapped to MesAur1.0 (GCA_000349665.1), downloaded from Ensembl. Reads assigned to each gene were counted by featureCounts39 (v.2.0.1) with human refseq gene sets as references for Caco-2 cells, and gene annotation of golden hamster from Ensembl database for the hamster dataset. Genes without at least 1 read mapped on average in each sample were considered undetectable and were filtered out. Read counts were normalized by the trimmed mean of M-values method and differential expression was calculated using R package edgeR (v.3.28.1) and genewise negative binomial generalized linear models with quasi-likelihood tests method was used for statistical tests. Cut-offs imposed for differential expression analysis were set as a false discovery rate (FDR) of 0.05 and fold change >2 or <0.5. The pathway analysis was performed by R package clusterProfiler40 (v.3.14.3) and Metascape41. Heat maps were plotted using R package pheatmap (v.1.0.12) (http://CRAN.R-project.org/package=pheatmap). Other plots were generated by R package ggplot2 (v.3.3.0) (https://ggplot2.tidyverse.org). PCA analysis was performed by R package factoextra (1.0.7).
Pseudotyping of VSV and pseudotype-based inhibition assay
VSV pseudotyped with the S of MERS-CoV, SARS-CoV or SARS-CoV-2 were generated as previously reported, with some modifications42. In brief, BHK-21/WI-2 cells (Kerafast) overexpressing the S were inoculated with VSV-G pseudotyped ΔG-luciferase VSV (Kerafast). After a 2-h inoculation at 37 °C, the inoculum was removed and cells were refed with DMEM supplemented with 5% FBS and VSV-G antibody (I1, mouse hybridoma supernatant from CRL-2700; ATCC; 1:100). Pseudotyped particles were collected at 24 h after inoculation, then centrifuged at 1,320g to remove cell debris and stored at −80 °C until use.
To determine the effect of the compounds on viral entry, Vero E6 cells were treated with clofazimine at a concentration of 2.5 μM for 1 h before inoculation with respective pseudotyped VSV. After a 2-h inoculation in the presence of the compounds, the inoculum was removed, and cells were refed with fresh medium for further culture. The activity of firefly luciferase was measured using bright-Glo luciferase assay (Promega) for quantitative determination at 16 h after transduction.
The effect of clofazimine on SARS-CoV-2 viral replication
The full-length SARS-CoV-2 viral RNA transcripts were in vitro-synthesized from an infectious clone of SARS-CoV-2 (provided by P.-Y. Shi) according to a recently published protocol43. Ten μg of total RNA transcripts and 5 μg SARS-CoV-2 N gene transcript were mixed with Vero E6 cells stably expressing SARS-CoV-2 N and then added into a 0.2-cm cuvette for nucleofection with the 4D-Nucleofector Core Unit (Lonza) using pulse code V-001. Immediately after electroporation, 1,000 μl of prewarmed medium was added to the cuvette and cells were subsequently aliquoted into 384-well plates. Two hours after seeding, compounds at different concentrations were added into each well. At 12 h after electroporation, intracellular and viral RNA was purified from the treated cells with TurboCapture 384 mRNA Kit (Qiagen) in accordance with the manufacturer’s instructions. The purified RNA was subjected to first-strand cDNA synthesis using the high-capacity cDNA reverse transcription kit (Applied Biosystems) with the following primer (TagRdRp-F: 5′-CGGTCATGGTGGC GAATAACCCTGTGGG TTTTACACTTAA-3′). Real-time PCR analysis was performed using TaqPath 1-step RT-qPCR Master Mix (Applied Biosystems). The following primers and probe were used for negative-stranded RNA detection: Tag-F: 5′-CGGTCATGGTGGCGAATAACCCTGT-3′, ORF1ab-R: 5′-ACGATTGTGCATCAGCTGA-3′, ORF1ab-P: 5′-6FAM-CCGTCTGCGGTA TGTGGAAAGGTTATGG -BHQ1-3′). In parallel, 5 μg of the in vitro-transcribed GFP mRNA (StemMACS eGFP mRNA, 130-101-114) was electroporated into Vero E6 cells. At 2 h after seeding, cells were treated either with clofazimine or remdesivir, and then cultured for a further 24 h. The GFP signal was measured by flow cytometry analysis using FlowJo (v.10.0.7).
Detection of S binding against ACE2 or heparin
Binding between the purified SARS-CoV-2 S and the cellular entry factors ACE2 or heparin were detected by ELISA, as was previously described22. High-binding microtitre plates were coated with heparin–BSA (100 ng per well) or recombinant ACE2 (200 ng per well) overnight at 4 °C. The plates were then blocked for 3 h at 37 °C with TSM buffer (20 mM Tris buffer, pH 7.4, containing 150 mM NaCl, 2 mM MgCl2, 2 mM CaCl2, 0.05% Tween-20 and 1% BSA). Next, 10 nM biotinylated SARS-CoV-2 S in a dilution of clofazimine (10–0.1 μM) in TSM buffer was added to the plates in triplicate. Bound biotinylated protein was detected by adding Avidin-HRP (405103, BioLegend) diluted 1:2,000 in TSM buffer. Finally, the plates were developed with TMB turbo substrate for 5–15 min. The reaction was quenched using 1 M sulfuric acid and the absorbance was measured at 450 nm.
S-mediated membrane fusion assay
SARS-CoV-2 S-mediated cell–cell fusion assay was performed as was previously established, with some modifications22. Vero cells were cotransfected with 1 μg SARS-CoV-2 S plasmid and 0.4 μg EGFP plasmid for 8 h. In another 24-well plate, Vero cells (that is, target cells expressing ACE2) were washed with PBS before being cocultured with the effector cells (that is, Vero cells cotransfected with S and EGFP), in the presence of the indicated concentration of clofazimine. After another 48 h, S-mediated membrane fusion, as reflected by the green florescence area, was evaluated using confocal imaging. Vero cells transfected with EGFP, without S transfection, were included as a negative control.
Time-of-addition assay
A time-of-drug-addition assay was performed to investigate which stage of the SARS-CoV-2 life cycle it is that clofazimine interferes with, as previously described15. In brief, Vero E6 cells were seeded in 96-well plates (4 × 104 cells per well). The cells were infected by SARS-CoV-2 USA-WA1/2020 at an MOI of 1.5 and then incubated for additional 1 h. The viral inoculum was then removed, and the cells were washed twice with PBS. At 1 h after inoculation (that is, after entry), clofazimine at a concentration of 5 μM was added to the infected cells at time points indicated, followed by incubation at 37 °C in 5% CO2 until 10 h after inoculation (that is, one virus life cycle). Cells were fixed at 10 h after inoculation for the quantification of the percentage of infected cells using an immunofluorescence assay targeting SARS-CoV-2 N.
In vitro primer elongation assay using RdRp core complex
Expression plasmids for SARS-CoV-2 nsp7, nsp8 and nsp12 were provided by the laboratory of S. Darst and E. Campbell. The expression and purification of nsp7, nsp8 and nsp12 were performed as previously described25. The transcription scaffold was prepared by annealing a 2:1:3 molar ratio of template strand RNA (5′-CUAUCCCCAUGUGAU UUUAAUAGCUUCUUAGGAGAAUGACGUAGCAUGCUACGCG), 32P labelled primer RNA (5′-CGCGUAGCAUGCUACGUCAUUCUCCUAAGAAGCUA) and nontemplate DNA (5′-ATCACATGGGGATAG) at 95 °C for 5 min, and a slow cooldown to room temperature in elongation buffer (20 mM Tris pH 7.5, 40 mM KCl, 5 mM MgCl2 and 5 mM DTT). The prepared scaffold was incubated with different concentrations of clofazimine or SL-11128 for 2 h at room temperature. SL-11128 has previously been reported to be effective against SARS-CoV-215. The RdRp core complex was prepared by mixing a 1:3 molar ratio of nsp12 to nsp7 and nsp 8 and incubating for 20 min in ice. The scaffold was added to the RdRp complex and incubated for 20 min at 30 °C. To start the reaction, an equal volume of rNTP was added to the RdRp–scaffold complex. The final concentrations for assays were: 20 nM scaffold, 250 nM RdRp, 5–40 μM of clofazimine or SL-11128 and 10 μM rNTP. The reaction was quenched by mixing with loading buffer (90% formamide, 50 mM EDTA, 0.05% xylene cyanol and 0.05% bromophenol blue) at different time points. All samples were denatured at 95 °C for 20 min and analysed by 8% TBE and urea denaturing PAGE.
In vitro helicase unwinding assay using nsp13
Helicase inhibition assays were performed as previously described37. Recombinant SARS-CoV-2 nsp13 was expressed in Escherichia coli BL21(DE3) cells and purified using the Ni2+-loaded HiTrap Chelating System (GE Healthcare) according to the manufacturer’s instructions. The DNA oligomers FL-Cy3 oligonucleotide (5′-TTTTTTTTTTTTTTTTTTTTC GAGCACCGCTGCGGCTGCACC(Cy3)-3′) and RL-BHQ oligonucleotide (5′-(BHQ2)GGTGCAGCCGCAGCGGTGCTCG-3′) were purchased from Metabion. RNA oligonucleotides, including RNA_31/18-mer-Cy3 (5′-CGCAGUCUUCUCCUGGUGCUCGAAC AGUGAC(Cy3)-3′) and RNA_31/18-mer_BHQ (5′-(BHQ2)GUCACUGUUCGAGCACCA-3′), were synthesized from IDT. Both oligonucleotides were annealed in the buffer of 20 mM Tris-HCl pH 8.0 and 150 mM NaCl. Assays were performed in buffer composed of 20 mM Tris-HCl buffer, pH 7.4, 150 mM NaCl, 0.1 mg ml−1 BSA, 5 mM MgCl2, 5 mM TCEP, 5% glycerol and 10 nM helicase, followed by the addition of 0.5 μl of 100 mM ATP and 1.5 μl of oligonucleotide mixture to make the final concentration of FL-Cy3:RL-BHQ oligonucleotide and FL-Cy3:RL oligonucleotide 5 nM and 10 nM, respectively. Fluorescence (λex = 550 nm, λem = 620 nm) was measured using SpectraMax iD3 Multi-Mode microplate reader to determine the extent of DNA–RNA duplex unwinding.
Illustrations
Hamster illustrations in Fig. 3a and Extended Data Fig. 6a were created with BioRender software (https://biorender.com/).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The complete sequences of the SARS-CoV-2 HKU-001a and SARS-CoV-2 USA-WA1/2020 strains that we used were obtained through GenBank (accession numbers MT230904 (HKU-001a), MT246667 and MN908947 (USA-WA1/2020)). The raw RNA-seq data discussed in this Article have been deposited in NCBI’s Gene Expression Omnibus and are accessible through GEO series accession number GSE162899. The hg38 reference genome was downloaded from UCSC database (https://hgdownload.soe.ucsc.edu/goldenPath/hg38/bigZips/). The refseq gene annotation was retrieved from the UCSC Table Browser (https://genome.ucsc.edu/cgi-bin/hgTables). The NCBI SARS-CoV-2 reference genome (NC_045512.2) was downloaded from the NCBI database (https://www.ncbi.nlm.nih.gov/nuccore/1798174254). The MesAur1.0 genome assembly (GCA_000349665.1) and annotation were obtained from the Ensembl database (https://asia.ensembl.org/Mesocricetus_auratus/Info/Index). Any other relevant data are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Peiris, J. S. et al. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 361, 1319–1325 (2003).
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. & Fouchier, R. A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 367, 1814–1820 (2012).
Barry, V. C. et al. A new series of phenazines (rimino-compounds) with high antituberculosis activity. Nature 179, 1013–1015 (1957).
World Health Organization. Coronavirus Disease (COVID-19) Situation Reports. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports (2020).
Chan, J. F. et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet 395, 514–523 (2020).
Hung, I. F. et al. SARS-CoV-2 shedding and seroconversion among passengers quarantined after disembarking a cruise ship: a case series. Lancet Infect. Dis. 20, 1051–1060 (2020).
Chan, J. F., To, K. K., Tse, H., Jin, D. Y. & Yuen, K. Y. Interspecies transmission and emergence of novel viruses: lessons from bats and birds. Trends Microbiol. 21, 544–555 (2013).
Chan, J. F. et al. Middle East respiratory syndrome coronavirus: another zoonotic betacoronavirus causing SARS-like disease. Clin. Microbiol. Rev. 28, 465–522 (2015).
Zumla, A., Chan, J. F., Azhar, E. I., Hui, D. S. & Yuen, K. Y. Coronaviruses – drug discovery and therapeutic options. Nat. Rev. Drug Discov. 15, 327–347 (2016).
Sheahan, T. P. et al. Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci. Transl. Med. 9, eaal3653 (2017).
Beigel, J. H., Tomashek, K. M. & Dodd, L. E. Remdesivir for the treatment of Covid-19 – preliminary report. Reply. N. Engl. J. Med. 383, 994 (2020).
Wang, Y. et al. Remdesivir in adults with severe COVID-19: a randomised, double-blind, placebo-controlled, multicentre trial. Lancet 395, 1569–1578 (2020).
Goldman, J. D. et al. Remdesivir for 5 or 10 days in patients with severe Covid-19. N. Engl. J. Med. 383, 1827–1837 (2020).
Hung, I. F. et al. Triple combination of interferon beta-1b, lopinavir-ritonavir, and ribavirin in the treatment of patients admitted to hospital with COVID-19: an open-label, randomised, phase 2 trial. Lancet 395, 1695–1704 (2020).
Riva, L. et al. Discovery of SARS-CoV-2 antiviral drugs through large-scale compound repurposing. Nature 586, 113–119 (2020).
Gopal, M., Padayatchi, N., Metcalfe, J. Z. & O’Donnell, M. R. Systematic review of clofazimine for the treatment of drug-resistant tuberculosis. Int. J. Tuberc. Lung Dis. 17, 1001–1007 (2013).
Schaad-Lanyi, Z., Dieterle, W., Dubois, J. P., Theobald, W. & Vischer, W. Pharmacokinetics of clofazimine in healthy volunteers. Int. J. Lepr. Other Mycobact. Dis. 55, 9–15 (1987).
Mansfield, R. E. Tissue concentrations of clofazimine (B663) in man. Am. J. Trop. Med. Hyg. 23, 1116–1119 (1974).
Kissler, S. M., Tedijanto, C., Goldstein, E., Grad, Y. H. & Lipsitch, M. Projecting the transmission dynamics of SARS-CoV-2 through the postpandemic period. Science 368, 860–868 (2020).
Sharma, A. et al. Human iPSC-derived cardiomyocytes are susceptible to SARS-CoV-2 infection. Cell Rep. Med. 1, 100052 (2020).
Yuan, S. et al. SREBP-dependent lipidomic reprogramming as a broad-spectrum antiviral target. Nat. Commun. 10, 120 (2019).
Clausen, T. M. et al. SARS-CoV-2 infection depends on cellular heparan sulfate and ACE2. Cell 183, 1043–1057 (2020).
Lu, L. et al. Structure-based discovery of Middle East respiratory syndrome coronavirus fusion inhibitor. Nat. Commun. 5, 3067 (2014).
Xia, S. et al. A pan-coronavirus fusion inhibitor targeting the HR1 domain of human coronavirus spike. Sci. Adv. 5, eaav4580 (2019).
Chen, J. et al. Structural basis for helicase-polymerase coupling in the SARS-CoV-2 replication–transcription complex. Cell 182, 1560–1573 (2020).
Chan, J. F.-W. et al. Simulation of the clinical and pathological manifestations of coronavirus disease 2019 (COVID-19) in golden Syrian hamster model: implications for disease pathogenesis and transmissibility. Clin. Infect. Dis. 71, 2428–2446 (2020).
Nix, D. E. et al. Pharmacokinetics and relative bioavailability of clofazimine in relation to food, orange juice and antacid. Tuberculosis 84, 365–373 (2004).
Herold, T. et al. Elevated levels of IL-6 and CRP predict the need for mechanical ventilation in COVID-19. J. Allergy Clin. Immunol. 146, 128–136 (2020).
Cholo, M. C., Steel, H. C., Fourie, P. B., Germishuizen, W. A. & Anderson, R. Clofazimine: current status and future prospects. J. Antimicrob. Chemother. 67, 290–298 (2012).
Hwang, T. J. et al. Safety and availability of clofazimine in the treatment of multidrug and extensively drug-resistant tuberculosis: analysis of published guidance and meta-analysis of cohort studies. BMJ Open 4, e004143 (2014).
Falzon, D. et al. World Health Organization treatment guidelines for drug-resistant tuberculosis, 2016 update. Eur. Respir. J. 49, 1602308 (2017).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Weng, Z. et al. A simple, cost-effective but highly efficient system for deriving ventricular cardiomyocytes from human pluripotent stem cells. Stem Cells Dev. 23, 1704–1716 (2014).
Chan, J. F. et al. Development and evaluation of novel real-time reverse transcription–PCR assays with locked nucleic acid probes targeting leader sequences of human-pathogenic coronaviruses. J. Clin. Microbiol. 53, 2722–2726 (2015).
Chu, H. et al. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin. Infect. Dis. 71, 1400–1409 (2020).
Yuan, S. et al. Metallodrug ranitidine bismuth citrate suppresses SARS-CoV-2 replication and relieves virus-associated pneumonia in Syrian hamsters. Nat. Microbiol. 5, 1439–1448 (2020).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e8 (2020).
Xie, X. et al. An infectious cDNA clone of SARS-CoV-2. Cell Host Microbe 27, 841–848.e3 (2020).
Ianevski, A., Giri, A. K. & Aittokallio, T. SynergyFinder 2.0: visual analytics of multi-drug combination synergies. Nucleic Acids Res. 48, W488–W493 (2020).
Acknowledgements
The authors acknowledge the assistance of the University of Hong Kong Li Ka Shing Faculty of Medicine Centre for PanorOmic Sciences. We thank S. Darst and E. Campbell for providing us plasmids of nsp7, nsp8 and nsp12; and K. Godula and R. Porell the for heparin–BSA conjugate. This study was partly supported by funding to the University of Hong Kong: the Health and Medical Research Fund (grant no. COVID190121 and CID-HKU1-11), the Food and Health Bureau, The Government of the Hong Kong Special Administrative Region; the Innovation and Technology Fund, Innovation and Technology Commission, the Government of the Hong Kong Special Administrative Region; Sanming Project of Medicine in Shenzhen, China (SZSM201911014); and the High Level-Hospital Program, Health Commission of Guangdong Province, China; and donations from R. Y.-H. Yu and C. K.-W. Yu, the Shaw Foundation Hong Kong, M. S.-K. Tong, M. M.-Y. Tam Mak, Lee Wan Keung Charity Foundation Limited, Hong Kong Sanatorium & Hospital, the M. Hui, Hui Hoy and Chow Sin Lan Charity Fund Limited, the Chan Yin Chuen Memorial Charitable Foundation, M. M.-W. Lee, the Hong Kong Hainan Commercial Association South China Microbiology Research Fund, the Jessie & George Ho Charitable Foundation, Perfect Shape Medical Limited, Kai Chong Tong, Norman & Cecilia Yip Foundation, L. K.-M. Tse, Foo Oi Foundation Limited, B. H.-C. Lee, P. C. So and the Lo Ying Shek Chi Wai Foundation. This work was also supported by the grants to the Sanford Burnham Prebys Medical Discovery Institute: DoD: W81XWH-20-1-0270; DHIPC: U19 AI118610; NCI Cancer Center grant P30 CA030199-40; Fluomics/NOSI: U19 AI135972, as well as generous philanthropic donations from D. Ruch, and S. and J. Blair. The funding sources had no role in the study design, data collection, analysis, interpretation or writing of the report.
Author information
Authors and Affiliations
Contributions
S.Y., X.Y., X.M., Z.-W.Y., L.R., N.M., L.P., Y.P., R.A.L., C.C.-Y.C., P.-M.L., J.C., R.L., Y.-Y.Z., Y.-D.T. and W.X. designed and/or performed experiments. S.Y., X.Y., X.M., Z.-W.Y., J.F.-W.C., L.P., Y.D., K.-H.S. and H. Chu analysed data. S.Y., X.Y., Z.-W.Y., C.-K.Y., K.-H.K., K.-Y.S, W.-K.A., R.W., Y.-Y.Z., Y.-D.T., C.-Y.L., T.M.C., J.P., J.O. and L.S. generated critical reagents. K.-Y.Y., S.K.C., R.S., S.Y., X.Y., X.M. and L.P. wrote the manuscript. S.Y., J.F.-W.C., C.C.-S.C., V.K.-M.P. and K.T. provided the hamster data. A.C.-Y.L, A.J.Z., S.Y. and J.F.-W.C. performed histopathological evaluation. S.K.C., R.S., I.F.-N.H., R.A.L., H. Chen, H.S., D.W., X.-H.C., J.D.E. and D.-Y.J. provided conceptual advice and troubleshooting. K.-Y.Y., S.K.C. and R.S. oversaw the conception and supervised the study. K.-Y.Y. and S.K.C. provided the grant support.
Corresponding authors
Ethics declarations
Competing interests
J.F.-W.C. has received travel grants from Pfizer Corporation Hong Kong and Astellas Pharma Hong Kong Corporation Limited, and was an invited speaker for Gilead Sciences Hong Kong Limited and Luminex Corporation. S.K.C. is an inventor on a patent application on repurposed antiviral compounds for SARS-CoV-2 owned by Scripps Research and Sanford Burnham Prebys. US Patent Application Serial No. 63/010630, entitled ‘Methods and Compositions for Antiviral Treatment’ relates to aspects of this work and was filed on 15 April 2020. The corresponding authors had full access to all the data in the study and had final responsibility for the decision to submit for publication. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Publishe r’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Anti-MERS-CoV activity and cytotoxicity measurement of clofazimine in matching cells.
a, Clofazimine inhibited MERS-CoV replication in a dose-dependent manner. EC50 was achieved by plaque reduction assay and plotted using logistic nonlinear regression model (GraphPad Prism 7). Data are mean ± s.d., n = 3 biologically independent samples. b, MERS-CoV-infected Huh7 cells (MOI of 0.01) were treated with clofazimine (5 μM) or were not treated. Top, immunofluorescence staining of MERS-CoV-N antigen (green), and Huh7 cell nucleus (blue). Scale bars, 20 μm. Representative images selected from a pool images captured in two independent experiments. Bottom, MERS-CoV-N-positive cells quantified by flow cytometry. The experiments were performed twice and representative quantifications are shown. c, The cell viability was determined using CellTiter-Glo assays and in the absence of virus infection. The drug-incubation time in the cytotoxicity assay was consistent with that in the antiviral assay: for example, at 24 h after treatment for Huh7 cells, primary human small-airway epithelial cells (HSAEpC) and human embryonic stem-cell-derived cardiomyocytes (CM); at 48 h after treatment for Vero E6 cells; and at 72 h after treatment for BSC1 and human embryonic lung fibroblasts (HELF). Data are mean ± s.d., n = 3 biological replicates. The experiment was repeated twice for confirmation.
Extended Data Fig. 2 Exploration of possible effects of clofazimine on virus entry and replication.
a, Time-of-addition assay. SARS-CoV-2-infected VeroE6 cells were incubated with clofazimine and at the time points indicated. Infection at 10 h after infection was quantified by immunostaining for N. Data are normalized to the DMSO-treated cells, and are mean ± s.d., n = 6 independent experiments. One-way ANOVA followed by Tukey’s post hoc test. b, Clofazimine has no effect on ACE2 and DPP4 expression. Caco-2 cells were treated with clofazimine for 16 h before collection for western blotting analysis. The expression of ACE2 and DPP4 were determined using anti-ACE2 antibody (Abcam; ab108252; 1:1,000) and anti-DPP4 antibody (Cell Signaling; 67138T; 1:1,000), respectively. For gel source data, see Supplementary Fig. 1. The experiment was repeated twice for confirmation. c, Clofazimine has no inhibition on the binding between either ACE2 or heparin and SARS-CoV-2 S, which are two critical cellular components for viral attachment and infection. Clofazimine was titrated as the indicated concentrations. Dashed line represents binding without inhibitor (that is, 0 μM). Data are mean ± s.d., n = 3 biologically independent samples. d, Quantification on the basis of GFP positive area using ImageJ software (corresponding to Fig. 2b). Error bars are s.e.m., n = 5 randomly selected images. One-way ANOVA followed by Dunnett’s post test. e, Scheme of in vitro-transcribed viral RNA replication assay. Vero E6 cells were electroporated with in vitro-transcribed viral RNA. At 2 h after seeding, the cells were treated with the drug. Negative-stranded RNA was then quantified at 12 h after electroporation. f, GFP mRNA was used as a negative control and its translation, determined at 24 h after electroporation, was not affected by remdesivir or clofazimine. Error bars are s.e.m., n = 3 independent experiments. n.s., not significant (P > 0.05, when compared with the 0-μM group by two-tailed Student’s t-test). g, Clofazimine shows a marginal effect against main protease (Mpro) and papain-like protease (PLpro) activity. The activity of purified SARS-CoV-2 Mpro and SARS-CoV-2 PLpro enzymes was measured after adding their respective peptide substrates. Enzyme activity in the absence (100% activity) and presence of clofazimine were calculated. Data are mean ± s.d., n = 3 independent experiments. h, Clofazimine shows no inhibition on the polymerase activity of nsp7–nsp8–nsp12–RdRp complex. The scaffold used in this in vitro transcription inhibition assay is listed in the top panel. Inhibition effect of clofazimine or SL-11128 on the RdRp core complex was analysed by a primer elongation assay. SL-11128, a SARS-CoV-2 inhibitor15, shows some inhibition effect over time (0, 15 s, 1 min, 5 min and 20 min), whereas that of clofazimine was minor (ranging from around 5 to 40 μM). All of the reactions were performed at 30 °C. All the above experiments were repeated twice for confirmation.
Extended Data Fig. 3 Transcriptional analysis of clofazimine treatment.
a, Caco-2 cells were infected with SARS-CoV-2 at an MOI of 0.1 before being treated with clofazimine, remdesivir and DMSO (0.1%) at the indicated concentrations. Cell culture supernatant was collected at 48 h after infection, and was subjected to viral load determination. Two-tailed Student’s t-test, compared with the DMSO group. *P < 0.05, ** P < 0.01. Data are mean ± s.d., n = 3 independent experiments. b, Timeline of the transcriptomic study (MOI = 4). h, hours after drug treatment; hpi, hours post virus infection; CFZ, clofazimine. c, Principal components analysis of RNA-seq dataset after ‘reads per kilobase of transcript per million reads mapped’ normalization on each gene expression level. Each dot represents one sample. The percentage labelled on the x or y axis represents the proportion of variance explained with each principal component (PC). d, Patterns of transcription levels across all samples. The genes that were significantly and differentially expressed (fold change >2 or <0.5, FDR < 0.05) between 6 h after infection and mock are shown. Conditions include 3 h and 6 h after infection for Caco-2 cells (MOI = 4) with or without clofazimine treatment. Genes were clustered using the K-means method. e, Heat map of 197 transcription factors regulated by clofazimine treatment without infection (left), and known interactions among these transcription factors (right). f, Heat map of the genes enriched in MAPK signalling, TNF signalling, interleukin signalling or cytokine–cytokine receptor interaction. These genes are upregulated (fold change >2, FDR < 0.05) by either 6 h after treatment with clofazimine (without infection) or at 6 h after infection with clofazimine treatment, compared to mock infection.
Extended Data Fig. 4 Transcriptional comparison of clofazimine-treated cells with or without SARS-CoV-2 infection.
a, Top-enriched pathways of significantly upregulated genes (FDR < 0.05, fold change > 2) compared between clofazimine treatment alone at 6 h versus mock-infected cells (6 h CFZ versus mock) and at 6 h after infection under clofazimine treatment versus mock (6 h CFZ versus mock). Pathway analysis was performed by Metascape. *Innate-immunity-related pathways. b, Network of enriched terms represented as pie charts. Pies are colour-coded on the basis of the identities of the gene lists. ‘6 h. clofazimine vs mock’, upregulated genes by clofazimine treatment without infection at 6 h, compared with mock; ‘6 hpi. clofazimine vs mock’, upregulated genes by clofazimine treatment at 6 h after infection, and compared with mock.
Extended Data Fig. 5 Prophylactic and therapeutic administration of clofazimine reduced viral load in the lungs without compromising humoral response in hamsters.
Experimental design is shown in Fig. 3a. a, Viral yield in lung tissue from hamsters, after prophylactic (n = 5 hamsters per group) or therapeutic treatment (n = 11 hamsters (each remdesivir and clofazimine group) and n = 13 hamsters (vehicle group)), collected at 4 days after infection and titrated by RT–qPCR assays. Data are mean ± s.d. Two-tailed Student’s t-test for prophylaxis groups and one-way ANOVA followed by Dunnett’s post test for therapeutic groups, comparing with the vehicle group (black symbols). ***P < 0.001, **P < 0.01. b, Hamsters exhibited a normal humoral immune response after SARS-CoV-2 infection and clofazimine treatment. The sera were serially diluted before adding to the N-coated ELISA plate (n = 3 hamsters per group).
Extended Data Fig. 6 Transcriptional analysis of hamster lung tissues with clofazimine administration.
a, Experimental design. Tissue samples were collected at the indicated time points. b, Gene Ontology Biological Process analysis results for upregulated genes, comparing prophylactic clofazimine administration with its corresponding vehicle controls. c, RNA expression (reads per kilobase of transcript per million mapped reads) of the 13 genes enriched in the ‘leukocyte differentiation’ category of the Gene Ontology Biological Process analysis. These genes are upregulated (fold change > 1.5, P value < 0.01) in the prophylactic clofazimine group versus vehicle controls. *MHC class-II molecules. ↑Transcription factors upregulated by clofazimine in both Caco-2 cells and hamster lung tissues. Data are mean ± s.e.m., n = 3 hamsters per group. d, Heat map of immune-response-related genes in lungs of uninfected or infected hamsters administered prophylactic clofazimine or vehicle controls. e, Gene Ontology Biological Process analysis results for upregulated genes, comparing clofazimine- and vehicle-treated hamster lungs without virus infection.
Extended Data Fig. 7 Histological analysis lung pathology in each group.
a, Representative images of H&E-stained lung tissue section from hamsters treated as indicated. Numbered circled areas are shown in magnified images to the right, illustrating the severity of (1) bronchiolar and/or peribronchiolar cell death; (2) alveoli destruction and/or alveolar infiltration; and (3) blood vessel and perivascular infiltration. Black arrows indicate sites of infiltration. These representative images were selected from a pool of 15 images captured in 3 randomly selected hamsters per group. Scale bars, 200 μm. b, Semiquantitative histology scores were given to each lung tissue by grading the severity of damage in bronchioles, alveoli and blood vessels and accumulating the total scores. For bronchioles: 0 = normal structure; 1 = mild peribronchiolar infiltration; 2 = peribronchiolar infiltration plus epithelial cell death; and 3 = score 2 plus intrabronchiolar wall infiltration and epithelium desquamation. For alveoli: 0 = normal structure; 1 = alveolar wall thickening and congestion; 2 = focal alveolar space infiltration or exudation; and 3 = diffuse alveolar space infiltration or exudation or haemorrhage. For blood vessels: 0 = normal structure; 1 = mild perivascular oedema or infiltration; 2 = vessel wall infiltration; and 3 = severe endothelium infiltration. Data are mean ± s.d., 3 randomly selected slides for each group. Unpaired two-tailed Student’s t-test between the two prophylactic groups. One-way ANOVA followed by Dunnett’s post test for the therapeutic groups. **P < 0.01, ***P < 0.001, compared to the vehicle-control group. The histological score of mock infection was set as zero.
Extended Data Fig. 8 Clofazimine exhibits antiviral synergy with remdesivir in vitro.
a, Remdesivir at the indicated doses was combined with clofazimine at the indicated doses or a negative control (DMSO), and antiviral dose–response relationships were determined. Vero E6 cells were pre-treated for 16 h with increasing concentrations of the indicated compound and then infected with SARS-CoV-2 at a MOI of 0.01. Thirty hours after infection, the infected cells were analysed by immunofluorescence imaging. For each condition, the percentage of infection was calculated as the ratio of the number of infected cells stained for SARS-CoV-2 N to the number of cells stained with DAPI. Data are normalized to mean values for DMSO-treated wells, and are mean ± s.e.m. (n = 4 biological repeats over 2 independent experiments). b, Topographical two-dimensional map of synergy scores determined in synergy finder44. The colour gradient indicates the synergy score (red, highest score). x axis, remdesivir up to 10 μM, y axis, clofazimine up to 10 μM. c, Dose–response analysis of remdesivir alone (black) and in combination with 0.15625 μM (blue) or 0.625 μM (red) clofazimine. The observed compound activities are represented by solid lines, and the predicted additive combinatorial activities are indicated by dashed lines. The dotted black line denotes 90% inhibition of infection. Data are normalized to mean values for DMSO-treated wells, and are mean ± s.e.m. (n = 4 biological repeats over 2 independent experiments). d, Counting of cell numbers in each drug combination, as indicated. Mean ± s.d., n = 4 biologically independent samples. The experiments were repeated twice for confirmation.
Extended Data Fig. 9 Clofazimine exhibits antiviral synergy with remdesivir in hamsters.
Experiments were performed as shown in Fig. 4b. a, Representative image of infected cells by immunofluorescence staining in lung at 4 days after infection. SARS-CoV-2 N expression (green) is shown in diffuse alveolar areas (thick white arrow) and in the focal bronchiolar epithelial cells (thin white arrow) of the vehicle-treated hamster lungs, whereas standard and low-dose remdesivir (Rem) groups as well as the clofazimine group (Clo) exhibit reduced N expression. Combinatorial therapy restricts the virus replication within the entry gate of lung infection (that is, bronchiolar epithelial cells). These representative images were selected from a pool of 15 images captured in 3 randomly selected hamsters per group. b, Representative image of infected cells by immunofluorescence staining in nasal turbinate at 4 days after infection. SARS-CoV-2 N (NP) (green) and cell nuclei (blue) were stained. Scale bars, 200 μm. These representative images were selected from a pool of 15 images captured in 5 hamsters per group. c, N-positive cells per 50× field per the nasal turbinate section of a hamster. One-way ANOVA followed by Dunnett’s post test and compared with vehicle (n = 5 hamsters per group). **P < 0.01, n.s., not significant.
Supplementary information
Supplementary Information
This file includes Supplementary Figures 1 and 2.
Supplementary Table 1
List of up-regulated genes comparing SARS-CoV-2 infected Caco-2 cells at 6 hpi versus un-infected cells. FDR<0.05, fold change>2 or <0.5 compared with mock.
Supplementary Table 2
List of down-regulated genes comparing SARS-CoV-2 infected Caco-2 cells at 6 hpi versus un-infected cells. FDR<0.05, fold change>2 or <0.5 compared with mock.
Supplementary Table 3
List of up-regulated genes in hamster lung tissues comparing prophylactic administration of Clofazimine versus vehicle control group at 4 dpi. Fold change >1.5, p<0.05 in prophylactic clofazimine administration group compared with vehicle controls at 4 dpi.
Rights and permissions
About this article
Cite this article
Yuan, S., Yin, X., Meng, X. et al. Clofazimine broadly inhibits coronaviruses including SARS-CoV-2. Nature 593, 418–423 (2021). https://doi.org/10.1038/s41586-021-03431-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03431-4
This article is cited by
-
Modulation of alveolar macrophage and mitochondrial fitness by medicinal plant-derived nanovesicles to mitigate acute lung injury and viral pneumonia
Journal of Nanobiotechnology (2024)
-
PMI-controlled mannose metabolism and glycosylation determines tissue tolerance and virus fitness
Nature Communications (2024)
-
COVID-19 drug discovery and treatment options
Nature Reviews Microbiology (2024)
-
Molecular docking as a tool for the discovery of novel insight about the role of acid sphingomyelinase inhibitors in SARS- CoV-2 infectivity
BMC Public Health (2024)
-
Identification of the SARS-unique domain of SARS-CoV-2 as an antiviral target
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.