Abstract
The inflammasome initiates innate defence and inflammatory responses by activating caspase-1 and pyroptotic cell death in myeloid cells1,2. It consists of an innate immune receptor/sensor, pro-caspase-1, and a common adaptor molecule, ASC. Consistent with their pro-inflammatory function, caspase-1, ASC and the inflammasome component NLRP3 exacerbate autoimmunity during experimental autoimmune encephalomyelitis by enhancing the secretion of IL-1β and IL-18 in myeloid cells3,4,5,6. Here we show that the DNA-binding inflammasome receptor AIM27,8,9,10 has a T cell-intrinsic and inflammasome-independent role in the function of T regulatory (Treg) cells. AIM2 is highly expressed by both human and mouse Treg cells, is induced by TGFβ, and its promoter is occupied by transcription factors that are associated with Treg cells such as RUNX1, ETS1, BCL11B and CREB. RNA sequencing, biochemical and metabolic analyses demonstrated that AIM2 attenuates AKT phosphorylation, mTOR and MYC signalling, and glycolysis, but promotes oxidative phosphorylation of lipids in Treg cells. Mechanistically, AIM2 interacts with the RACK1–PP2A phosphatase complex to restrain AKT phosphorylation. Lineage-tracing analysis demonstrates that AIM2 promotes the stability of Treg cells during inflammation. Although AIM2 is generally accepted as an inflammasome effector in myeloid cells, our results demonstrate a T cell-intrinsic role of AIM2 in restraining autoimmunity by reducing AKT–mTOR signalling and altering immune metabolism to enhance the stability of Treg cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The RNA-seq data are available in the Gene Expression Omnibus (GEO) repository at the National Center for Biotechnology Information under accession number GSE133019. The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD018638. All other data supporting the findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
Change history
14 April 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03490-7
References
Guo, H., Callaway, J. B. & Ting, J. P. Inflammasomes: mechanism of action, role in disease, and therapeutics. Nat. Med. 21, 677–687 (2015).
Broz, P. & Dixit, V. M. Inflammasomes: mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 16, 407–420 (2016).
Gris, D. et al. NLRP3 plays a critical role in the development of experimental autoimmune encephalomyelitis by mediating Th1 and Th17 responses. J. Immunol. 185, 974–981 (2010).
Furlan, R. et al. Caspase-1 regulates the inflammatory process leading to autoimmune demyelination. J. Immunol. 163, 2403–2409 (1999).
Shaw, P. J. et al. Cutting edge: critical role for PYCARD/ASC in the development of experimental autoimmune encephalomyelitis. J. Immunol. 184, 4610–4614 (2010).
Martin, B. N. et al. T cell-intrinsic ASC critically promotes TH17-mediated experimental autoimmune encephalomyelitis. Nat. Immunol. 17, 583–592 (2016).
Fernandes-Alnemri, T., Yu, J. W., Datta, P., Wu, J. & Alnemri, E. S. AIM2 activates the inflammasome and cell death in response to cytoplasmic DNA. Nature 458, 509–513 (2009).
Fernandes-Alnemri, T. et al. The AIM2 inflammasome is critical for innate immunity to Francisella tularensis. Nat. Immunol. 11, 385–393 (2010).
Krieg, A. M. AIMing 2 defend against intracellular pathogens. Nat. Immunol. 11, 367–369 (2010).
Rathinam, V. A. et al. The AIM2 inflammasome is essential for host defense against cytosolic bacteria and DNA viruses. Nat. Immunol. 11, 395–402 (2010).
Goverman, J. Autoimmune T cell responses in the central nervous system. Nat. Rev. Immunol. 9, 393–407 (2009).
Reynolds, J. M., Martinez, G. J., Chung, Y. & Dong, C. Toll-like receptor 4 signaling in T cells promotes autoimmune inflammation. Proc. Natl Acad. Sci. USA 109, 13064–13069 (2012).
Heng, T. S. & Painter, M. W. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Papatheodorou, I. et al. Expression Atlas: gene and protein expression across multiple studies and organisms. Nucleic Acids Res. 46, D246–D251 (2018).
Josefowicz, S. Z., Lu, L. F. & Rudensky, A. Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Kitagawa, Y. et al. Guidance of regulatory T cell development by Satb1-dependent super-enhancer establishment. Nat. Immunol. 18, 173–183 (2017).
Lee, W. & Lee, G. R. Transcriptional regulation and development of regulatory T cells. Exp. Mol. Med. 50, e456 (2018).
Luo, C. T. & Li, M. O. Transcriptional control of regulatory T cell development and function. Trends Immunol. 34, 531–539 (2013).
Mottet, C., Uhlig, H. H. & Powrie, F. Cutting edge: cure of colitis by CD4+CD25+ regulatory T cells. J. Immunol. 170, 3939–3943 (2003).
Wilson, J. E. et al. Inflammasome-independent role of AIM2 in suppressing colon tumorigenesis via DNA-PK and Akt. Nat. Med. 21, 906–913 (2015).
Man, S. M. et al. Critical role for the DNA sensor AIM2 in stem cell proliferation and cancer. Cell 162, 45–58 (2015).
Zeng, H. & Chi, H. mTOR signaling in the differentiation and function of regulatory and effector T cells. Curr. Opin. Immunol. 46, 103–111 (2017).
Procaccini, C. et al. The proteomic landscape of human ex vivo regulatory and conventional T cells reveals specific metabolic requirements. Immunity 44, 406–421 (2016).
Fragale, A. et al. IFN regulatory factor-1 negatively regulates CD4+ CD25+ regulatory T cell differentiation by repressing Foxp3 expression. J. Immunol. 181, 1673–1682 (2008).
Overacre-Delgoffe, A. E. et al. Interferon-gamma drives Treg fragility to promote anti-tumor immunity. Cell 169, 1130–1141 (2017).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Merkenschlager, M. & von Boehmer, H. PI3 kinase signalling blocks Foxp3 expression by sequestering Foxo factors. J. Exp. Med. 207, 1347–1350 (2010).
Michalek, R. D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Li, G. et al. EphB3 suppresses non-small-cell lung cancer metastasis via a PP2A/RACK1/Akt signalling complex. Nat. Commun. 3, 667 (2012).
Apostolidis, S. A. et al. Phosphatase PP2A is requisite for the function of regulatory T cells. Nat. Immunol. 17, 556–564 (2016).
Zhou, X. et al. Selective miRNA disruption in Treg cells leads to uncontrolled autoimmunity. J. Exp. Med. 205, 1983–1991 (2008).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Miller, S. D. & Karpus, W. J. Experimental autoimmune encephalomyelitis in the mouse. Curr. Protoc. Immunol. 77, 15.1.1–15.1.18 (2007).
Matsushita, T., Yanaba, K., Bouaziz, J. D., Fujimoto, M. & Tedder, T. F. Regulatory B cells inhibit EAE initiation in mice while other B cells promote disease progression. J. Clin. Invest. 118, 3420–3430 (2008).
Erben, U. et al. A guide to histomorphological evaluation of intestinal inflammation in mouse models. Int. J. Clin. Exp. Pathol. 7, 4557–4576 (2014).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Acknowledgements
The following funding supports are acknowledged: NIH (AI029564, CA156330, DK094779, AI097392, AI123193), the National Multiple Sclerosis Society (CA10068 to J.P.-Y.T. and FG 1968-A-1 to W.C.), NIAID (AI067798 to J.P.-Y.T., H.G. and W.J.B.), the National Multiple Sclerosis Society (RG-1802-30483), and a Yang Family Biomedical Scholars Award to Y.Y.W. We received help from G.D. Sempowski (Duke University, Duke Human Vaccine Institute, Durham, NC, USA, Immune Monitoring Core for AI067798) for the Luminex assay and analysis. We thank B. Sartor (P01-DK094779) for advice regarding the colitis model, N. Fisher and J. Dow (UNC flow cytometry facility supported in part by P30 CA016086 Cancer Center Core Support Grant) for cell sorting, and Dale Cowley (UNC Animal Models Core for P30 CA016086) for generating Aim2fl/fl mice. This work benefitted from a publicly available online database assembled by the ImmGen consortium and The European Bioinformatics Institute (EMBL-EBI) without prior permission. We thank E. Holley-Guthrie for genotyping mice.
Author information
Authors and Affiliations
Contributions
W.C. and Z.G. contributed equally to this manuscript. W.C., Z.G., Y.Y.W. and J.P.-Y.T. designed experiments and wrote the manuscript. W.C. and Z.G. performed and analysed most of the experiments. H.G., L.C. and G.Z. contributed to the colitis experiments. K.L. and M.D. contributed to the western blots. L.X. and X.C. contributed to IP–MS analysis. S.A.G. contributed to quantitative PCR analysis. X.T. contributed to bioinformatics analysis. E.R. contributed to the metabolism assays. Y.W. and M.A.S. contributed to the scoring and quantitation of spinal cord pathology. S.A.M. contributed to the scoring of colon pathology. W.J.B., L.F. and S.Z. assisted in the mouse experiments. W.J.B. edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Kate Schroder and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Aim2−/− mice show normal T cell homeostasis.
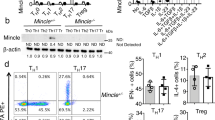
a, b, Flow cytometry of CD4+FOXP3+ Treg cells (a) and IFNγ-, IL-17A-producing CD4+ T cells (b) in the spleen and PLN of wild-type and Aim2−/− mice at day 14 of an EAE course. Representative results (left) and statistical analysis (right) of six experiments are shown. c, Flow cytometry of CD4+FOXP3+ Treg cells in the thymus, spleen and PLN of wild-type and Aim2−/− mice. Representative results (left) and statistical analysis (right) of three experiments are shown. d, Total number of cells isolated from the thymus, spleen and PLN of wild-type and Aim2−/− mice. Experimental design and statistical analysis performed as described in c. e, f, Flow cytometry of naive, effector/memory CD4+ (e) and CD8+ (f) T cells in the spleen and PLN of wild-type and Aim2−/− mice (n = 3 per group) analysed by CD44 and CD62L expression. Experimental design and statistical analysis performed as described in c. g, h, Flow cytometry of IFNγ-, IL-4- or IL-17A-producing CD4+ cells (g) and IFNγ-producing CD8+ T cells (h) in wild-type and Aim2−/− mice (n = 3 per group). Experimental design and statistical analysis performed as described in c. Representative (top) and composite (bottom) data are shown. Data are mean ± s.e.m. P values determined by two-sided t-test. ns, not significant.
Extended Data Fig. 2 Aim2−/− mice show normal Treg cell proliferation, survival and ratio of central or effector Treg cells at steady state or during EAE in vivo, normal CD4+ T cell proliferation in vitro, and normal CD8+ T cell distribution and cytokine production during EAE.
a, Flow cytometry of Ki67 to analyse proliferation of wild-type and Aim2−/− Treg cells in the PLN and spleen at steady state. Representative results (top) and statistical analysis (bottom) of five experiments are shown. b, Apoptosis of wild-type and Aim2−/− Treg cells in the PLN and spleen at steady state was analysed by flow cytometry using annexin V and 7-aminoactinomycin D (7AAD) staining. Representative results (top) and statistical analysis (bottom) of five experiments are shown. c, Flow cytometry of Ki67 to analyse proliferation of wild-type and Aim2−/− Treg cells in the PLN, spleen and spinal cord during EAE. Representative results (top) and statistical analysis (bottom) of six experiments are shown. d, Apoptosis of wild-type and Aim2−/− Treg cells in spinal cord during EAE was analysed by flow cytometry using annexin V and 7AAD staining. Representative results (top) and statistical analysis (bottom) of six experiments are shown. e, Flow cytometry of CD44 and CD62L in wild-type and Aim2−/− Treg cells isolated from spinal during EAE. Representative results (top) and statistical analysis (bottom) of six experiments are shown. f, Flow cytometry of wild-type and Aim2−/− CD4+ T cell proliferation stimulated with different doses of anti-CD3/CD28, determined by CFSE dilution assay. Representative results of two independent experiments. g, Flow cytometry of 2D2 and Aim2-deficient 2D2 (2D2 × Aim2−/− CD4+ T cell proliferation stimulated with bone marrow-derived dendritic cells (BMDCs) pulsed with MOG35–55 peptide) as determined by CFSE dilution assay. Representative of two independent experiments. h, Flow cytometry of wild-type and Aim2−/− CD4+ or CD8+ T cells in the PLN, spleen and spinal cord during EAE. Representative results (left) and statistical analysis (right) of six experiments are shown. i–k, Flow cytometry of IFNγ-, IL-17A-producing (i) or FOXP3+, IFNγ-producing (j) CD8+ T cells in the PLN, spleen and spinal cord during EAE. Representative results (i, j) and statistical analysis (k) of six experiments are shown. Data are mean ± s.e.m. P values determined by two-sided t-test.
Extended Data Fig. 3 Aim2 is highly expressed in Treg cells and its promoter is bound by Treg cell-related transcription factors.
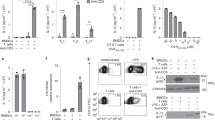
a, b, Mouse Aim2 gene expression in different T cell subsets from publicly available gene microarray (a) and RNA-seq (b) databases (https://www.immgen.org/). c, Human AIM2 gene expression in T cell and macrophage subsets from the Expression Atlas of EMBL-EBI (https://www.ebi.ac.uk/). d, Purity of isolated CD4+CD25+ Treg cells from wild-type and Aim2−/− mice. Flow cytometry of CD4+CD25+FOXP3+ Treg cells shows that more than 97% of isolated Treg cells are FOXP3+ cells. e, Aim2 expression was assessed from isolated CD4+CD25+ Treg cells and CD4+ T cells. Cells were freshly isolated from pooled spleens and lymph nodes and purified by MACS beads. Aim2 mRNA expression was examined by quantitative PCR; n = 4 experiments. f, The mRNA expression of Aim2 in Treg cells and CD4+ T cells stimulated with anti-CD3/CD28 plus IL-2 (500 U ml−1) for 24 h; n = 4 experiments. g, The mRNA expression of Aim2 in Treg cells stimulated with anti-CD3/CD28 plus IL-2 (500 U ml−1) in the absence (−) or presence (+) of TGFβ (1 ng ml−1) for 24 h; n = 4 experiments. h, Aim2 expression was assessed in freshly isolated CD4+CD25+ Treg cells from wild-type and Tgfbr2fl/fl CD4-cre (TGFbRII-KO) mice. Aim2 mRNA expression was examined by quantitative PCR; n = 5 experiments. i, The mRNA expression of Aim2 in naive CD4+ or CD8+ T cells stimulated with anti-CD3/CD28 plus IL-2 (40 U ml−1) in the absence or presence of TGFβ (1 ng ml−1) for 24 h; n = 4 experiments. j, ChIP–seq analysis of RUNX1, ETS1, BCL11B and CREB binding to the Aim2 promoter region in Treg cells and CD4+ Tconv cells (NCBI SRA database number: DRP003376). Data are mean ± s.e.m. **P < 0.01, ***P < 0.001, ****P < 0.0001, two-sided t-test.
Extended Data Fig. 4 AIM2 is essential for Treg cells to suppress T cell-mediated colitis and EAE.
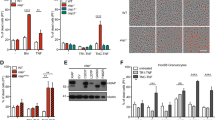
a, Haematoxylin and eosin (H&E) staining of colons from T cell-induced colitis mice transferred with wild-type CD4+CD45RBhi T cells (Tn) alone (n = 5) or in combination with wild-type (n = 6) or Aim2−/− (n = 6) CD4+CD25+ Treg cells, collected 7 weeks after T cell transfer. Scale bars, 1 mm (40×) and 100 μm (100×). b, Statistical analysis of pathology score of colitis mice with biological replicates of each group depicted in a. Tn only: n = 5; Tn + WT Treg: n = 6; Tn + Aim2−/− Treg: n = 6. c, Cytokine levels in the supernatants of colon tissue cultures from mice depicted in a measured by Millipore Luminex assay, collected 7 weeks after T cell transfer. Tn only: n = 4; Tn + WT Treg: n = 5; Tn+Aim2−/− Treg: n = 5. d, Change of body weight of Rag1−/− recipients receiving wild-type naive CD4+CD45RBhi T cells (Tn) alone or in combination with wild-type, Aim2−/− or Asc−/− CD4+CD25+ Treg cells. Rag1−/− recipients of Tn (n = 8), Tn + WT Treg (n = 6), Tn + Aim2−/− Treg (n = 9), Tn + Asc−/− Treg (n = 7); composite of two independent experiments. P value determined by two-way ANOVA. e, Flow cytometry of CD4+FOXP3+ Treg cells in the colons of Rag1−/− recipients of Tn (n = 8), Tn + WT Treg (n = 6), Tn + Aim2−/− Treg (n = 9), Tn + Asc−/− Treg (n = 7), collected 7 weeks after T cell transfer. P value by one-way ANOVA with Tukey’s multiple comparisons test. f, Schema of EAE induction in Rag1−/− mice transferred with 2D2 CD4+ T cells either alone or in combination with wild-type or Aim2−/− CD4+CD25+ Treg cells. Lymphocytes and tissues were obtained 14–15 days after initial T cell transfer for further analysis. g, Flow cytometry shows the distributions of 2D2 CD4+ T cells (Vβ11+) or Treg cells (FOXP3+) before transfer to Rag1−/− recipient mice. h, Mean EAE clinical score of mice depicted in f; n = 5 mice per group. P value by two-way ANOVA and Holm–Sidak post hoc test. Data are representative of three independent experiments. The difference between 2D2 alone and 2D2 with Aim2−/− Treg cells is not significant. i, Flow cytometry of IFNγ+ or IL-17A+ CD4+Vβ11+ T cells in spinal cords from groups depicted in f. Left, representative sample; right, composite data pooled of five mice per group from three independent experiments. j, Flow cytometry of CD4+FOXP3+ Treg cells from spinal cord derived from mice depicted in f. Left, representative sample; right, composite data pooled of five mice per group from three independent experiments. P value determined by one-way ANOVA with Tukey’s multiple comparisons test. k, Flow cytometry of IFNγ+ or IL-17A+CD4+Vβ11− T cells in spinal cord from groups depicted in f. Left, representative sample; right, composite data summarized from five biological replicates. l, CD25−CD44lowCD62Lhi naive CD4+ T cells (Tresp cells) were isolated from wild-type mice and labelled with CFSE. CD4+CD25+ Treg cells were isolated from wild-type or Aim2−/− mice by FACS. Tresp and Treg cells of different genotypes were mixed at indicated ratios and stimulated with anti-CD3 in the presence of irradiated antigen-presenting cells from mixed spleens and lymph nodes. The suppressive activity of Treg cells was assessed by CFSE dilution of Tresp cells. Data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, two-sided t test unless specified.
Extended Data Fig. 5 Aim2fl/fl FGC R26T mice show normal T cell homeostasis, normal Treg cell proliferation and survival at steady state and during EAE.
a, Genotyping of FACS-sorted Treg (CD4+CD25+GFP+) and CD4+ Tconv (CD4+CD25−) cells showing efficient deletion of Aim2 genomic DNA in Treg cells. The image is representative of three independent experiments. b, Immunoblot analysis of AIM2 protein in Treg cells from Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice. The image is representative of three independent experiments. c, Images of Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice and corresponding lymphoid organs of spleen and lymph nodes. d, Flow cytometry of CD4+FOXP3+CD25+Treg cells in the thymus, spleen and PLN of Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice Representative results (top) and statistical analysis (bottom) of four experiments are shown. e, Flow cytometry of CD4+ or CD8+ T cells in the spleen and PLN of Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice. Representative results (left) and statistical analysis (right) of four experiments are shown. f, g, Flow cytometry of naive, effector/memory CD4+ (f) and CD8+ (g) T cells in the spleen and PLN of Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice by assessing CD44 and CD62L expression. Representative results (left for f, top for g) and statistical analysis (right for f, bottom for g) of four experiments are shown. h, i, Flow cytometry of IFNγ-, IL-4- or IL-17A-producing CD4+ cells (h) and IFNγ-producing CD8+ T cells (i) in Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice. Representative results (top) and statistical analysis (bottom) of four experiments are shown. j, Statistical summary of IFNγ-, IL-17A-producing CD4+Tomato− Tconv cells in the PLN (left) and spleen (right) of Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice at day 28 of an EAE course. Composite data summarized from three biological replicates. k, Statistical summary of IFNγ-, IL-17A-producing CD4+Tomato+ Treg cells in the PLN (left) and spleen (right) of Aim2+/+ FGC R26T and Aim2fl/fl FGC R26T mice at day 28 of EAE course. Composite data summarized from three biological replicates. l, Flow cytometry of Ki67 to analyse proliferation of Aim2+/+ FGC and Aim2fl/fl FGC Treg cells in the PLN, spleen and spinal cord during EAE. Left, representative sample; right, composite data summarized from three biological replicates. m, Apoptosis of Aim2+/+ FGC and Aim2fl/fl FGC Treg cells in the spinal cord during EAE was analysed by flow cytometry using annexin V and 7AAD staining. Left, representative sample; right, composite data summarized from three biological replicates. n, Flow cytometry of Ki67 to analyse proliferation of Aim2+/+ FGC and Aim2fl/fl FGC Treg cells in the PLN and spleen at steady state. Left, representative sample; right, composite data of five mice of two independent experiments. o, Apoptosis of Aim2+/+ FGC and Aim2fl/fl FGC Treg cells in the PLN and spleen at steady state was analysed by flow cytometry using annexin V and 7AAD staining. Left, representative sample; right, composite data of five mice of two independent experiments. Data are mean ± s.e.m. P values determined by two-sided t-test unless specified.
Extended Data Fig. 6 Enhanced glycolytic, IFN and MYC target signatures are found in Aim2−/−Treg cells isolated in vivo.
a, Glycolytic activity of wild-type (n = 7 biological replicates per group) and Aim2−/− (n = 6 biological replicates per group) Treg cells untreated (UNT) or treated (TRE) with anti-CD3/CD28 antibodies plus IL-2 (500 U ml−1) for 24 h. Statistics of glycolysis (ECAR rate after glucose addition) and glycolytic capacity (maximal ECAR after subtracting the ECAR rate after exposure to 2-deoxy-d-glucose (2-DG) calculated from Fig. 3a. Data are mean ± s.e.m. **P < 0.01, ***P < 0.001, two-sided t-test. b, Heat map of IFNα response signature of RNA-seq data from Aim2−/− compared to wild-type Treg cells stimulated with anti-CD3/CD28 plus IL-2 (500 U ml−1) at indicated time points (0 or 24 h). c, Heat map of IFNγ response signature as described in b. d, Heat map of MYC target profiles as described in b.
Extended Data Fig. 7 Enhanced gene signature found in TGFβ-induced Aim2−/− Treg cells using RNA-seq analysis.
a, Flow cytometry of IFNγ+ or IL-17A+ CD4+ cells from wild-type and Aim2−/− mice after four days of differentiation under TH1, pTH17 and cTH17 conditions, respectively, as described in Fig. 4c. Data are representative of four independent experiments. b, Summary of top pathways positively enriched in anti-CD3/CD28 activated Aim2−/− CD4+ T cells in the presence of TGFβ (2 ng ml−1) and IL-2 (40 U ml−1) for 24 h, by GSEA analysis of the RNA-seq dataset. c, d, Enrichment of IFNγ (c) and IFNα response pathways (d) by GSEA (left) and heat map (right) of pathway-related genes in Aim2−/− versus wild-type CD4+ T cells stimulated with anti-CD3/CD28 in the presence of TGFβ (2 ng ml−1) and IL-2 (40 U ml−1) for 24 h.
Extended Data Fig. 8 RNA-seq analysis reveals enhanced mTOR, MYC and glycolytic signatures in TGFβ-induced Aim2−/− Treg cells.
a, Heat map of PI3K–AKT–mTORC-related gene expression in wild-type and Aim2−/− CD4+ T cells stimulated with anti-CD3/CD28 in the presence of TGFβ (2 ng ml−1) and IL-2 (40 U ml−1) for 24 h. b, Heat map of mTORC1 signalling-related gene expression, with samples described in a. c, Heat map of MYC target-related gene expression, with samples described in a. d, Heat map of glycolysis-related gene expression, with samples described in a.
Extended Data Fig. 9 AKT–mTOR signalling in wild-type and Aim2−/− Treg, CD4+ and CD8+ T cells.
a, b, Immunoblot analysis of p-AKT(S473), p-FOXO1, p-FOXO3A, p-S6, p-4E-BP1, MYC and β-actin in wild-type and Aim2−/− CD4+ T cells (a) or CD8+ T cells (b) stimulated with anti-CD3/CD28 plus IL-2 (40 U ml−1) for 24 h. c–e, Immunoblot analysis of p-AKT(S473), p-FOXO1, p-FOXO3A, p-S6, p-4E-BP1, MYC and β-actin in wild-type and Aim2−/− Treg cells (c) stimulated with anti-CD3/CD28 plus IL-2 (500 U ml−1), or CD4+ (d) and CD8+ (e) T cells stimulated with anti-CD3/CD28 plus IL-2 (40 U ml−1) for indicated time points. Left, representative results; right, quantification for statistics by densitometric analysis using Image Lab software; n = 4 experiments (a–c); n = 3 experiments (d, e). *P < 0.05, **P < 0.01, analysed by two-sided paired t-test.
Extended Data Fig. 10 AIM2 interacts with the RACK1–PP2A–AKT complex and is critical to regulate AKT–mTOR signalling for Treg cell generation.
a, Flow cytometry analysis of FOXP3 in wild-type and Aim2−/− CD4+ T cells stimulated by anti-CD3/CD28 plus IL-2 (40 U ml−1) and TGFβ (2 ng ml−1) and then cultured with DMSO (n = 4 biological replicates/group), rapamycin (1 nM) (n = 7 biological replicates/group), or pp242 (0.5 μM) (n = 7 biological replicates/group) for 96 h. Results are representative of three independent experiments. P value by one-way ANOVA with Tukey’s multiple comparisons test. b, Schema of IP–MS approach to identify AIM2-interacting proteins in TGFβ-induced Treg cells. Wild-type and Aim2−/− naive CD4+ T cells were activated with anti-CD3/CD28 in the presence of TGFβ (2 ng ml−1) and IL-2 (40 U ml−1) for 24 h and protein lysates from each group were collected for further IP–MS analysis. c, Interaction of AIM2 and RACK1 detected by immunoprecipitation using anti-RACK1 antibody or anti-IgG as control in TGFβ-induced Treg (iTreg) cells and CD4+ T cells, and immunoblotted with different antibodies, including anti-PP2Aca, anti-AKT, anti-RACK1 and anti-AIM2. Arrow points to the AIM2 protein. Results are representative of three independent experiments. d, Wild-type and Aim2−/− CD4+ T cells were stimulated with anti-CD3 and CD28 plus IL-2 (40 U ml−1) and TGFβ (2 ng ml−1) for 24 h and transduced either with MIT-PP2A and MIG-RACK1, or with MIT and MIG vector controls. The cells were collected 3 days after virus transduction. The populations expressing PP2A (Thy1.1+), RACK1 (GFP+) and both (Thy1.1+GFP+) were identified by flow cytometry. e, Flow cytometry of p-AKT of wild-type and Aim2−/− Treg cells that overexpressed PP2A (Thy1.1+) or RACK1 (GFP+) compared to corresponding vector controls. Representative FACS plots (top) and statistical analysis (bottom) of six experiments are shown. P value by multiple unpaired t-test with Holm–Sidak method. f, Model for AIM2 function shows that AIM2 facilitates the interaction between RACK1 and PP2A phosphatase, causing dephosphorylation of AKT to restrain the activity of the mTOR pathway, therein promoting Foxp3 expression and Treg cell stability. Data are mean ± s.e.m. *P < 0.05, ****P < 0.0001.
Supplementary information
Supplementary Figures
This file contains the uncropped gels (Supplementary Fig. 1) and gating strategies for flow cytometry analysis (Supplementary Fig. 2).
Source data
Rights and permissions
About this article
Cite this article
Chou, WC., Guo, Z., Guo, H. et al. AIM2 in regulatory T cells restrains autoimmune diseases. Nature 591, 300–305 (2021). https://doi.org/10.1038/s41586-021-03231-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03231-w
This article is cited by
-
Inter-relationships of galectin-3 and NLR family pyrin domain containing 3 inflammasomes with oral lichen planus: a preliminary cross-sectional in vitro study
BMC Oral Health (2024)
-
Cytosolic DNA sensors in neurodegenerative diseases: from physiological defenders to pathological culprits
EMBO Molecular Medicine (2024)
-
Transcriptomic analysis reveals molecular characterization and immune landscape of PANoptosis-related genes in atherosclerosis
Inflammation Research (2024)
-
Inflammation and DNA damage: cause, effect or both
Nature Reviews Rheumatology (2023)
-
Intricacies of TGF-β signaling in Treg and Th17 cell biology
Cellular & Molecular Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.