Abstract
The formation of synapses during neuronal development is essential for establishing neural circuits and a nervous system1. Every presynapse builds a core ‘active zone’ structure, where ion channels cluster and synaptic vesicles release their neurotransmitters2. Although the composition of active zones is well characterized2,3, it is unclear how active-zone proteins assemble together and recruit the machinery required for vesicle release during development. Here we find that the core active-zone scaffold proteins SYD-2 (also known as liprin-α) and ELKS-1 undergo phase separation during an early stage of synapse development, and later mature into a solid structure. We directly test the in vivo function of phase separation by using mutant SYD-2 and ELKS-1 proteins that specifically lack this activity. These mutant proteins remain enriched at synapses in Caenorhabditis elegans, but show defects in active-zone assembly and synapse function. The defects are rescued by introducing a phase-separation motif from an unrelated protein. In vitro, we reconstitute the SYD-2 and ELKS-1 liquid-phase scaffold, and find that it is competent to bind and incorporate downstream active-zone components. We find that the fluidity of SYD-2 and ELKS-1 condensates is essential for efficient mixing and incorporation of active-zone components. These data reveal that a developmental liquid phase of scaffold molecules is essential for the assembly of the synaptic active zone, before maturation into a stable final structure.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All reagents are available on request. Source data are provided with this paper.
Change history
18 June 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03340-6
References
Südhof, T. C. Towards an understanding of synapse formation. Neuron 100, 276–293 (2018).
Südhof, T. C. The presynaptic active zone. Neuron 75, 11–25 (2012).
Ghelani, T. & Sigrist, S. J. Coupling the structural and functional assembly of synaptic release sites. Front. Neuroanat. 12, 81 (2018).
Lipton, D. M., Maeder, C. I. & Shen, K. Rapid assembly of presynaptic materials behind the growth cone in dopaminergic neurons is mediated by precise regulation of axonal transport. Cell Rep. 24, 2709–2722 (2018).
Bracha, D., Walls, M. T. & Brangwynne, C. P. Probing and engineering liquid-phase organelles. Nat. Biotechnol. 37, 1435–1445 (2019).
Sala, K. et al. The ERC1 scaffold protein implicated in cell motility drives the assembly of a liquid phase. Sci. Rep. 9, 13530 (2019).
Patel, M. R. et al. Hierarchical assembly of presynaptic components in defined C. elegans synapses. Nat. Neurosci. 9, 1488–1498 (2006).
Dai, Y. et al. SYD-2 Liprin-α organizes presynaptic active zone formation through ELKS. Nat. Neurosci. 9, 1479–1487 (2006).
Zhen, M. & Jin, Y. The liprin protein SYD-2 regulates the differentiation of presynaptic termini in C. elegans. Nature 401, 371–375 (1999).
Patel, M. R. & Shen, K. RSY-1 is a local inhibitor of presynaptic assembly in C. elegans. Science 323, 1500–1503 (2009).
Putnam, A., Cassani, M., Smith, J. & Seydoux, G. A gel phase promotes condensation of liquid P granules in Caenorhabditis elegans embryos. Nat. Struct. Mol. Biol. 26, 220–226 (2019).
Wheeler, J. R., Matheny, T., Jain, S., Abrisch, R. & Parker, R. Distinct stages in stress granule assembly and disassembly. eLife 5, e18413 (2016).
Zeng, M. et al. Phase transition in postsynaptic densities underlies formation of synaptic complexes and synaptic plasticity. Cell 166, 1163–1175 (2016).
Kalla, S. et al. Molecular dynamics of a presynaptic active zone protein studied in Munc13-1-enhanced yellow fluorescent protein knock-in mutant mice. J. Neurosci. 26, 13054–13066 (2006).
Tsuriel, S. et al. Exchange and redistribution dynamics of the cytoskeleton of the active zone molecule bassoon. J. Neurosci. 29, 351–358 (2009).
Deken, S. L. et al. Redundant localization mechanisms of RIM and ELKS in Caenorhabditis elegans. J. Neurosci. 25, 5975–5983 (2005).
Kittelmann, M. et al. Liprin-α/SYD-2 determines the size of dense projections in presynaptic active zones in C. elegans. J. Cell Biol. 203, 849–863 (2013).
Schwartz, M. L. & Jorgensen, E. M. SapTrap, a toolkit for high-throughput CRISPR/Cas9 gene modification in Caenorhabditis elegans. Genetics 202, 1277–1288 (2016).
Wu, Y. E., Huo, L., Maeder, C. I., Feng, W. & Shen, K. The balance between capture and dissociation of presynaptic proteins controls the spatial distribution of synapses. Neuron 78, 994–1011 (2013).
Chia, P. H., Patel, M. R. & Shen, K. NAB-1 instructs synapse assembly by linking adhesion molecules and F-actin to active zone proteins. Nat. Neurosci. 15, 234–242 (2012).
Collins, K. M. et al. Activity of the C. elegans egg-laying behavior circuit is controlled by competing activation and feedback inhibition. eLife 5, e21126 (2016).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Ryan, V. H. et al. Mechanistic view of hnRNPA2 low-complexity domain structure, interactions, and phase separation altered by mutation and arginine methylation. Mol. Cell 69, 465–479.e7 (2018).
Yamasaki, A. et al. Liquidity is a critical determinant for selective autophagy of protein condensates. Mol. Cell 77, 1163–1175.e9 (2020).
Vernon, R. M. et al. Pi-Pi contacts are an overlooked protein feature relevant to phase separation. eLife 7, e31486 (2018).
Woodruff, J. B., Hyman, A. A. & Boke, E. Organization and function of non-dynamic biomolecular condensates. Trends Biochem. Sci. 43, 81–94 (2018).
Wu, X. et al. RIM and RIM-BP form presynaptic active-zone-like condensates via phase separation. Mol. Cell 73, 971–984.e5 (2019).
Milovanovic, D., Wu, Y., Bian, X. & De Camilli, P. A liquid phase of synapsin and lipid vesicles. Science 361, 604–607 (2018).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Dickinson, D. J., Pani, A. M., Heppert, J. K., Higgins, C. D. & Goldstein, B. Streamlined genome engineering with a self-excising drug selection cassette. Genetics 200, 1035–1049 (2015).
Hubbard, E. J. A. FLP/FRT and Cre/lox recombination technology in C. elegans. Methods 68, 417–424 (2014).
Kato, M., Lin, Y. & McKnight, S. L. Cross-β polymerization and hydrogel formation by low-complexity sequence proteins. Methods 126, 3–11 (2017).
Walther, P. & Ziegler, A. Freeze substitution of high-pressure frozen samples: the visibility of biological membranes is improved when the substitution medium contains water. J. Microsc. 208, 3–10 (2002).
Buser, C. & Walther, P. Freeze-substitution: the addition of water to polar solvents enhances the retention of structure and acts at temperatures around −60 °C. J. Microsc. 230, 268–277 (2008).
Sato, T. A modified method for lead staining of thin sections. J. Electron Microsc. (Tokyo) 17, 158–159 (1968).
Mészáros, B., Erdös, G. & Dosztányi, Z. IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res. 46 (W1), W329–W337 (2018).
Hanson, J., Paliwal, K. & Zhou, Y. Accurate single-sequence prediction of protein intrinsic disorder by an ensemble of deep recurrent and convolutional architectures. J. Chem. Inf. Model. 58, 2369–2376 (2018).
Bolognesi, B. et al. A concentration-dependent liquid phase separation can cause toxicity upon increased protein expression. Cell Rep. 16, 222–231 (2016).
Britton, S. et al. DNA damage triggers SAF-A and RNA biogenesis factors exclusion from chromatin coupled to R-loops removal. Nucleic Acids Res. 42, 9047–9062 (2014).
Monahan, Z. et al. Phosphorylation of the FUS low-complexity domain disrupts phase separation, aggregation, and toxicity. EMBO J. 36, 2951–2967 (2017).
McDonnell, A. V., Jiang, T., Keating, A. E. & Berger, B. Paircoil2: improved prediction of coiled coils from sequence. Bioinformatics 22, 356–358 (2006).
Shen, K. & Bargmann, C. I. The immunoglobulin superfamily protein SYG-1 determines the location of specific synapses in C. elegans. Cell 112, 619–630 (2003).
Schafer, W. F. Genetics of egg-laying in worms. Annu. Rev. Genet. 40, 487–509 (2006).
Acknowledgements
We thank L. Luo, S. Chen and members of the Shen laboratory for critical reading of the manuscript; J. Lee for the GFP–SYD-2 strain; K. M. Collins for HSN strains; P. Calsou and N. L. Fawzi for FUS constructs; E. M. Jorgensen and B. Goldstein for FLPon tools; M. Boxem for a tagBFP2 construct; and L. Feng and Y. Xu for assistance with size-exclusion chromatography. This work was funded by Howard Hughes Medical Institute, of which K.S. is an investigator; by NIH grant R37 NS048392 to K.S.; and by a Helen Hay Whitney postdoctoral fellowship to N.A.M.
Author information
Authors and Affiliations
Contributions
N.A.M. and K.S. conceptualized the study. N.A.M. carried out the experiments and analyses. R.D.F. carried out electron-microscopy experiments and analyses. N.A.M. wrote the paper. N.A.M., R.D.F. and K.S. edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Erik Jorgensen, Dragomir Milovanovic and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
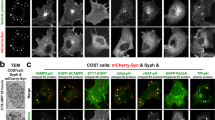
Extended Data Fig. 1 Presynaptic active-zone scaffolds display LLPS characteristics.
a, Predictions of intrinsic disorder in the sequences of presynaptic active-zone components, using the IUPred2a (ref. 37) and Spot-Disorder2 (ref. 38) algorithms. Values above 0.5 indicate intrinsic disorder. b, Predictions of LLPS of active-zone components, using catGRANULE39 and Pi–Pi predictor25 algorithms. Higher values indicate a propensity towards phase separation. c, Expression of GFP- or mCherry-tagged presynaptic active-zone components in COS-7 cells. FUS–GFP is a positive control40,41. Red boxes indicate example puncta that are photobleached in d. Scale bar, 10 μm. d, FRAP analysis of puncta. Data are means ± 95% CI from two independent experiments. Insets show example recovery kinetics of puncta from c. Scale bar, 1 μm. e, Time-lapse analysis of in vivo droplets containing SYD-2 or ELKS-1, showing liquid behaviours of fusion and relaxation. Red arrowheads point to droplets before fusion. Liquid behaviour was observed in multiple cells and in different technical replicates. UNC-10 puncta were dynamic when analysed by FRAP, but never showed liquid behaviour. Scale bars, 1 μm.
Extended Data Fig. 2 Controls for in vitro phase separation assays.
a, In vitro LLPS assay workflow and scoring conditions. b, Size-exclusion chromatography of purified mCherry and mCherry–FUS in 20 mM Tris pH 7.4, 500 mM NaCl, indicating that the protein isolated by the high-salt purification method is soluble. A280, absorption at 280 nm. c, In vitro LLPS assay. Left panels, soluble input materials before dilution to physiological salt concentration. Right panels, comparison between the effects of no crowding and 10% PEG conditions on a known phase-separating motif (FUS)41 and a negative control (mCherry). d, Representative FRAP images (top) and quantified FRAP dynamics (bottom) of FUS and mCherry in vitro condensates formed in 10% PEG. Data are means ± 95% CI from three independent experiments. e, Time-lapse analysis of in vitro condensates containing mCherry and FUS, showing the presence or absence of liquid behaviour. Phase-separation assays were performed in 20 mM Tris pH 7.4, 150 mM NaCl and 10% PEG 3350 where indicated with 10 μM protein. f, g, Phase diagrams for mCh–SYD-2(Nter) (f) and mCh–ELKS-1 (g), showing protein concentrations versus NaCl or PEG concentrations. Each combination was tested three times. Scale bars, 5 μm.
Extended Data Fig. 3 Purified constructs for in vitro phase separation assays.
SDS–PAGE gels, stained by Coomassie blue, of the purified constructs used for in vitro assays in the indicated figures. Many constructs display a secondary band 10 kDa below the major species, due to mCherry cleavage at the fluorophore centre (Y72)33.
Extended Data Fig. 4 Structure–function analysis of SYD-2 phase separation.
a, Diagram showing the structure of SYD-2, including disordered regions and coiled-coils predicted by the indicated algorithms37,38,42. LH1/LH2, liprin-homology domains; SAM, sterile alpha motif domains. b, Purified recombinant mCherry–SYD-2 constructs tested in an in vitro phase-separation assay. mCherry was fused to the N terminus of each sequence. c, Left panels in each column, example time-lapse analyses of in vitro condensates comprising the indicated SYD-2 constructs, showing the presence or absence of liquid fusion behaviour. Right panels, FRAP dynamics of each SYD-2 condensate. Data are means ± 95% CI for at least three condensates from two independent experiments. The y-axes show percentage intensity. Phase-separation assays were performed in 20 mM Tris pH 7.4, 150 mM NaCl with 10 μM protein. FRAP assays were performed on condensates formed in crowding conditions of 10% PEG, except in the case of the blue curves, where there was no crowding agent. Green or blue curves indicate a positive result (scored LLPS+) and black curves a negative result (scored LLPS−). Scale bars, 5 μm.
Extended Data Fig. 5 Structure–function analysis of ELKS-1 phase separation.
a, Diagram showing the structure of ELKS-1, including disordered regions and coiled-coils predicted by the indicated algorithms37,38,42. b, Purified recombinant mCherry–ELKS-1 constructs tested in an in vitro phase-separation assay. mCherry was fused to the N terminus of each sequence. c, Left panels in each column, example time-lapse analyses of in vitro condensates comprising the indicated ELKS-1 constructs, showing the presence or absence of liquid fusion behaviour. Right panels, FRAP dynamics for each ELKS-1 condensate. Data are means ± 95% CI, for at least three condensates from independent experiments. The y-axes show percentage intensity. Phase-separation assays were performed in 20 mM Tris pH 7.4, 150 mM NaCl with 10 μM protein. FRAP assays were performed on condensates formed in crowding conditions of 10% PEG, except in the case of the blue curves, where there was no crowding agent. Green or blue curves indicate a positive result (scored LLPS+) and black curves a negative result (scored LLPS−). Scale bars, 5 μm.
Extended Data Fig. 6 Phase-separation mutants of SYD-2 and ELKS-1 retain the ability to bind partners.
Shown are the results of yeast two-hybrid assays using the indicated bait and prey constructs. Protein–protein interactions are indicated by growth on −His/−Ade media.
Extended Data Fig. 7 Phase separation of SYD-2 and ELKS-1 is required for the assembly of multiple presynaptic active-zone components.
a–c, Cell-specific endogenous markers of presynaptic active zones in GFP–SYD-2 (a), GFP–ELKS-1 (b) or GFP–SYD-2(IDR∆) plus elks-1(LLPS–) (c) worms. d, Cell-specific endogenous markers of presynaptic active zones in syd-2 and elks-1 complete loss-of-function mutants. e, Quantified intensity of endogenous active-zone markers, as well as GFP–SYD-2 and GFP–ELKS, at HSN synapses in the indicated genotypes. Data are means ± 95% CI from independent animals. Comparisons were made using one-way ANOVA and Dunnett’s test. f, Expression of SNB-1 labelled with yellow fluorescent protein (YFP) from the unc-86 promoter at HSN synapses and cell bodies43. g, Quantification of SNB-1–YFP at HSN synapses and cell bodies. Data are means ± 95% CI from independent animals. Comparisons were made using one-way ANOVA and Dunnett’s test. h, Localization of endogenous GFP–SYD-2 or GFP–SYD-2(IDR∆) in the indicated genetic backgrounds. i, Quantification of endogenous GFP–SYD-2 or GFP–SYD-2(IDR∆) at HSN synapses in the indicated genetic backgrounds. Data are means ± 95% CI from independent animals; comparisons were made using one-way ANOVA and Dunnett’s (versus wild type) or Tukey’s (within genotypes) tests. Data for GFP–SYD-2, GFP–SYD-2(IDR∆) and GFP–SYD-2; elks-1(tm1233) are from Extended Data Fig. 7e. Dotted outlines show synaptic regions. Scale bars, 5 μm.
Extended Data Fig. 8 Phase separation of SYD-2 and ELKS-1 is required for proper synaptic function.
a, Left, the C. elegans egg-laying neuronal circuit44. The hermaphrodite-specific neuron (HSN) extends an axon across the vulval region, making specific synaptic contacts with vm1/2 vulval muscles and inhibitory VC4/5 motor neurons. Right, experimental setup. HSN is activated by channelrhodopsin ChR2 with blue light, while contraction of the vm2 muscle is measured with the Ca2+ sensor GCaMP5 (ref. 21). b, Representative vulval-muscle Ca2+ traces, detected by GCaMP5, in the indicated genotypes. ‘−ATR’ conditions lacked the essential ChR2 cofactor all-trans retinol. c, Quantification of the Ca2+ spike frequency, measured using GCaMP5, in the indicated genotypes. Data are means ± 95% CI from independent animals, with comparisons made using one-way ANOVA and Dunnett’s test.
Extended Data Fig. 9 Presynaptic assembly phenotypes of syd-2(IDR∆) mutants are rescued by certain exogenous phase-separation motifs only.
a, Diagram showing the replacement constructs tested. b, Comparison of amino-acid complexity between SYD-2/ELKS-1 and FUS/hnRNPA2 LLPS motifs. c, Left, time-lapse analysis of in vitro condensates comprising rescue constructs, showing liquid behaviour. Right, FRAP dynamics of in vitro condensates comprising rescue constructs. Data are means ± 95% CI across two independent experiments. SAM domains were omitted in these constructs. Assays were performed in 20 mM Tris pH 7.4, 150 mM NaCl with 10 μM protein. d, e, Cell-specific endogenous markers of presynaptic active zones (UNC-10, d) or synaptic vesicles (RAB-3, e) in the indicated GFP–SYD-2 alleles. Dotted boxes show synaptic regions. f, Quantified intensity of UNC-10 and RAB-3 at HSN synapses in the indicated genotypes. Data are means ± 95% CI from independent animals, with comparisons made using one-way ANOVA and Dunnett’s (versus wild type) or Tukey’s (IDR∆ versus IDR∆ + FUS) tests. Data for wild type, syd-2(IDR∆) and syd-2(IDR∆ + FUS) are from Fig. 3. g, Quantification of egg stages laid by animals of the indicated genotypes. The syd-2(IDR∆ + hnRNPA2 construct rescues egg laying only in the presence of mScarlet-I–FLPon–RAB-3, for unknown reasons. Data are means ± 95% CI from independent animals. Data for wild type, syd-2(IDR∆) and syd-2(IDR∆ + FUS) are from Fig. 3. Scale bars, 5 μm.
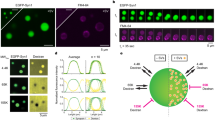
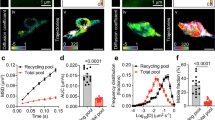
Extended Data Fig. 10 Loss of liquidity of SYD-2 and ELKS-1 impairs the scaffolding of GIT-1.
a, Endogenous GFP–ELKS-1(LLPS–) at early (1.5-fold embryo) nerve-ring synapses. Insets show FRAP images of bleached synapses. Main scale bar, 5 μm; inset scale bars, 1 μm. b, Quantification of nerve-ring GFP–ELKS-1 FRAP dynamics. Data are means ± 95% CI across two independent experiments. Wild-type data are from Fig. 1. c, FRAP dynamics of in vitro condensates comprising SYD-2 and ELKS-1 fragments. Data are means ± 95% CI from two independent experiments. Phase-separation assays were performed in 20 mM Tris pH 7.4, 150 mM NaCl and 10% PEG 3350, with 10 μM protein. Data for the ‘<5 min’ curve are from Fig. 2. d, Diagram showing the in vitro scaffolding experiment. e, f, Left, fluorescence images of the indicated condensates. Right, linescans performed through the centre of each condensate. Scale bars, 5 μm. g, Incorporation index (Iinc) for GIT-1–Dylight633. Data are means ± 95% CI from three independent experiments, with comparisons made using one-way ANOVA and Dunnett’s test.
Extended Data Fig. 11 Timing of assembly of the presynaptic active zone in early nerve-ring synapses.
Localization of endogenous GFP-tagged presynaptic active-zone components at sequential stages of embryo development. Each panel shows a maximum z projection of a separate embryo. Embryos are outlined in blue. A zy resliced projection through the nerve-ring region is shown to the right of each image. Red asterisks and arrowheads indicate visible localization to the nerve ring. Most puncta visible throughout the embryo are background autofluorescence, present even in untagged embryos (bottom). The timing of the emergence of fluorescence is stereotypical and was replicated in a second experiment. Scale bars, 10 μm.
Supplementary information
Supplementary Table
Supplementary Table 1. C. elegans strains used in this study.
Supplementary Table
Supplementary Table 2. Plasmids used in this study.
Source data
Rights and permissions
About this article
Cite this article
McDonald, N.A., Fetter, R.D. & Shen, K. Assembly of synaptic active zones requires phase separation of scaffold molecules. Nature 588, 454–458 (2020). https://doi.org/10.1038/s41586-020-2942-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2942-0
This article is cited by
-
Liprin-α proteins are master regulators of human presynapse assembly
Nature Neuroscience (2024)
-
Synaptic vesicle proteins and ATG9A self-organize in distinct vesicle phases within synapsin condensates
Nature Communications (2023)
-
Protein condensation diseases: therapeutic opportunities
Nature Communications (2022)
-
Phase separation and zinc-induced transition modulate synaptic distribution and association of autism-linked CTTNBP2 and SHANK3
Nature Communications (2022)
-
A brief guideline for studies of phase-separated biomolecular condensates
Nature Chemical Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.