Abstract
The cortex organizes sensory information to enable discrimination and generalization1,2,3,4. As systematic representations of chemical odour space have not yet been described in the olfactory cortex, it remains unclear how odour relationships are encoded to place chemically distinct but similar odours, such as lemon and orange, into perceptual categories, such as citrus5,6,7. Here, by combining chemoinformatics and multiphoton imaging in the mouse, we show that both the piriform cortex and its sensory inputs from the olfactory bulb represent chemical odour relationships through correlated patterns of activity. However, cortical odour codes differ from those in the bulb: cortex more strongly clusters together representations for related odours, selectively rewrites pairwise odour relationships, and better matches odour perception. The bulb-to-cortex transformation depends on the associative network originating within the piriform cortex, and can be reshaped by passive odour experience. Thus, cortex actively builds a structured representation of chemical odour space that highlights odour relationships; this representation is similar across individuals but remains plastic, suggesting a means through which the olfactory system can assign related odour cues to common and yet personalized percepts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data will be posted to Github or made available upon reasonable request (www.github.com/dattalab).
Code availability
All code will be posted to Github or made available upon reasonable request (www.github.com/dattalab).
Change history
12 August 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41586-020-2615-z
References
Haberly, L. B. Parallel-distributed processing in olfactory cortex: new insights from morphological and physiological analysis of neuronal circuitry. Chem. Senses 26, 551–576 (2001).
Courtiol, E. & Wilson, D. A. The olfactory mosaic: bringing an olfactory network together for odor perception. Perception 46, 320–332 (2017).
Barnes, D. C., Hofacer, R. D., Zaman, A. R., Rennaker, R. L. & Wilson, D. A. Olfactory perceptual stability and discrimination. Nat. Neurosci. 11, 1378–1380 (2008).
Wilson, D. A. & Sullivan, R. M. Cortical processing of odor objects. Neuron 72, 506–519 (2011).
Haddad, R. et al. A metric for odorant comparison. Nat. Methods 5, 425–429 (2008).
Dravnieks, A. Odor quality: semantically generated multidimensional profiles are stable. Science 218, 799–801 (1982).
Schiffman, S. S. Physicochemical correlates of olfactory quality. Science 185, 112–117 (1974).
Amoore, J. E. Stereochemical theory of olfaction. Nature 198, 271–272 (1963).
Buck, L. & Axel, R. A novel multigene family may encode odorant receptors: a molecular basis for odor recognition. Cell 65, 175–187 (1991).
Malnic, B., Hirono, J., Sato, T. & Buck, L. B. Combinatorial receptor codes for odors. Cell 96, 713–723 (1999).
Mombaerts, P. et al. Visualizing an olfactory sensory map. Cell 87, 675–686 (1996).
Otazu, G. H., Chae, H., Davis, M. B. & Albeanu, D. F. Cortical feedback decorrelates olfactory bulb output in awake mice. Neuron 86, 1461–1477 (2015).
Friedrich, R. W. & Wiechert, M. T. Neuronal circuits and computations: pattern decorrelation in the olfactory bulb. FEBS Lett. 588, 2504–2513 (2014).
Chae, H. et al. Mosaic representations of odors in the input and output layers of the mouse olfactory bulb. Nat. Neurosci. 22, 1306–1317 (2019).
Sosulski, D. L., Bloom, M. L., Cutforth, T., Axel, R. & Datta, S. R. Distinct representations of olfactory information in different cortical centres. Nature 472, 213–216 (2011).
Miyamichi, K. et al. Cortical representations of olfactory input by trans-synaptic tracing. Nature 472, 191–196 (2011).
Davison, I. G. & Ehlers, M. D. Neural circuit mechanisms for pattern detection and feature combination in olfactory cortex. Neuron 70, 82–94 (2011).
Stettler, D. D. & Axel, R. Representations of odor in the piriform cortex. Neuron 63, 854–864 (2009).
Schaffer, E. S. et al. Odor perception on the two sides of the brain: consistency despite randomness. Neuron 98, 736–742.e3 (2018).
Roland, B., Deneux, T., Franks, K. M., Bathellier, B. & Fleischmann, A. Odor identity coding by distributed ensembles of neurons in the mouse olfactory cortex. eLife 6, e26337 (2017).
Iurilli, G. & Datta, S. R. Population coding in an innately relevant olfactory area. Neuron 93, 1180–1197 (2017).
Babadi, B. & Sompolinsky, H. Sparseness and expansion in sensory representations. Neuron 83, 1213–1226 (2014).
Barak, O., Rigotti, M. & Fusi, S. The sparseness of mixed selectivity neurons controls the generalization-discrimination trade-off. J. Neurosci. 33, 3844–3856 (2013).
Dasgupta, S., Stevens, C. F. & Navlakha, S. A neural algorithm for a fundamental computing problem. Science 358, 793–796 (2017).
Haberly, L. B. & Bower, J. M. Olfactory cortex: model circuit for study of associative memory? Trends Neurosci. 12, 258–264 (1989).
Bekkers, J. M. & Suzuki, N. Neurons and circuits for odor processing in the piriform cortex. Trends Neurosci. 36, 429–438 (2013).
Saito, H., Chi, Q., Zhuang, H., Matsunami, H. & Mainland, J. D. Odor coding by a Mammalian receptor repertoire. Sci. Signal. 2, ra9 (2009).
Cleland, T. A., Morse, A., Yue, E. L. & Linster, C. Behavioral models of odor similarity. Behav. Neurosci. 116, 222–231 (2002).
Bolding, K. A. & Franks, K. M. Recurrent cortical circuits implement concentration-invariant odor coding. Science 361, eaat6904 (2018).
Chapuis, J. & Wilson, D. A. Bidirectional plasticity of cortical pattern recognition and behavioral sensory acuity. Nat. Neurosci. 15, 155–161 (2011).
Shakhawat, A. M., Harley, C. W. & Yuan, Q. Arc visualization of odor objects reveals experience-dependent ensemble sharpening, separation, and merging in anterior piriform cortex in adult rat. J. Neurosci. 34, 10206–10210 (2014).
Sell, C. S. in Chemistry and the Sense of Smell Ch. 5, 237–296 (Wiley, 2014).
Gottfried, J. A., Winston, J. S. & Dolan, R. J. Dissociable codes of odor quality and odorant structure in human piriform cortex. Neuron 49, 467–479 (2006).
Fournel, A., Ferdenzi, C., Sezille, C., Rouby, C. & Bensafi, M. Multidimensional representation of odors in the human olfactory cortex. Hum. Brain Mapp. 37, 2161–2172 (2016).
Diodato, A. et al. Molecular signatures of neural connectivity in the olfactory cortex. Nat. Commun. 7, 12238 (2016).
Boyd, A. M., Kato, H. K., Komiyama, T. & Isaacson, J. S. Broadcasting of cortical activity to the olfactory bulb. Cell Rep. 10, 1032–1039 (2015).
Schoenfeld, T. A. & Macrides, F. Topographic organization of connections between the main olfactory bulb and pars externa of the anterior olfactory nucleus in the hamster. J. Comp. Neurol. 227, 121–135 (1984).
Challis, R. C. et al. Systemic AAV vectors for widespread and targeted gene delivery in rodents. Nat. Protoc. 14, 379–414 (2019).
Bruno, R. M. & Sakmann, B. Cortex is driven by weak but synchronously active thalamocortical synapses. Science 312, 1622–1627 (2006).
Minamisawa, G., Funayama, K., Matsuki, N. & Ikegaya, Y. Intact internal dynamics of the neocortex in acutely paralyzed mice. J. Physiol. Sci. 61, 343–348 (2011).
Simons, D. J. & Carvell, G. E. Thalamocortical response transformation in the rat vibrissa/barrel system. J. Neurophysiol. 61, 311–330 (1989).
Maklad, A., Quinn, T. & Fritzsch, B. Intracranial distribution of the sympathetic system in mice: DiI tracing and immunocytochemical labeling. Anat. Rec. 263, 99–111 (2001).
Doevendans, P. A. J., Daemen, M. J., de Muinck, E. D. & Smits, J. F. Cardiovascular phenotyping in mice. Cardiovasc. Res. 39, 34–49 (1998).
Carey, R. M. & Wachowiak, M. Effect of sniffing on the temporal structure of mitral/tufted cell output from the olfactory bulb. J. Neurosci. 31, 10615–10626 (2011).
Kepecs, A. & Uchida, N. The sniff as a unit of olfactory processing. Chem. Senses 31, 167–179 (2006).
Cheung, M. & Carey, R. A method for generating natural and user-defined sniffing patterns in anesthetized or reduced preparations. Chem. Senses 34, 63–76 (2009).
Moldestad, O., Karlsen, P., Molden, S. & Storm, J. F. Tracheotomy improves experiment success rate in mice during urethane anesthesia and stereotaxic surgery. J. Neurosci. Methods 176, 57–62 (2009).
Ecker, A. S. et al. State dependence of noise correlations in macaque primary visual cortex. Neuron 82, 235–248 (2014).
Goard, M. & Dan, Y. Basal forebrain activation enhances cortical coding of natural scenes. Nat. Neurosci. 12, 1444–1449 (2009).
Ma, L. et al. Distributed representation of chemical features and tunotopic organization of glomeruli in the mouse olfactory bulb. Proc. Natl. Acad. Sci. USA 109, 5481–5486 (2012).
Srinivasan, S. & Stevens, C. A quantitative description of the mouse piriform cortex. Preprint at https://www.bioRxiv.org/content/ 10.1101/099002v1.full (2017).
Hagiwara, A., Pal, S. K., Sato, T. F., Wienisch, M. & Murthy, V. N. Optophysiological analysis of associational circuits in the olfactory cortex. Front. Neural Circuits 6, 18 (2012).
Luna, V. M. & Morozov, A. Input-specific excitation of olfactory cortex microcircuits. Front. Neural Circuits 6, 69 (2012).
Pachitariu, M., Stringer, C., Dipoppa, M. & Schröder, S. Suite2p: beyond 10,000 neurons with standard two-photon microscopy. Preprint at https://www.bioRxiv.org/content/10.1101/061507v2 (2017).
Willmore, B. & Tolhurst, D. J. Characterizing the sparseness of neural codes. Network 12, 255–270 (2001).
McInnes, L., Healy, J. & Melville, J. UMAP: Uniform Manifold Approximation and Projection for Dimension Reduction. Preprint at https://arXiv.org/abs/1802.03426 (2018).
Haxby, J. V. et al. A common, high-dimensional model of the representational space in human ventral temporal cortex. Neuron 72, 404–416 (2011).
Cowley, B. et al. Distance Covariance Analysis. Proc. 20th International Conference on Artificial Intelligence and Statistics 54, 242–251 (2017).
Litwin-Kumar, A., Harris, K. D., Axel, R., Sompolinsky, H. & Abbott, L. F. Optimal degrees of synaptic connectivity. Neuron 93, 1153–1164 (2017).
Abbott, L. F., Rajan, K. & Sompolinsky, H. Interactions between intrinsic and stimulus-evoked activity in recurrent neural networks. Preprint at https://arXiv.org/abs/0912.3832 (2009).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Platt, J. Probabilistic outputs for support vector machines and comparisons to regularized likelihood methods. Adv. Large Margin Classifiers 10, 61–74 (1999).
Kirkpatrick, S., Gelatt, C. D. Jr & Vecchi, M. P. Optimization by simulated annealing. Science 220, 671–680 (1983).
Wilson, D. A. & Linster, C. Neurobiology of a simple memory. J. Neurophysiol. 100, 2–7 (2008).
Acknowledgements
We thank members of the Datta laboratory, J. Pillow, R. Axel, L. Abbott, A. Litvin-Kumar, C. Schoonover, A. Fink and V. Ruta for comments on the manuscript, S. Knemeyer for technical illustrations, and N. Bhagat for laboratory assistance. We thank N. Mathur for assistance with viral delivery of GCaMP6s. We thank A. Giessel for early development of tools for image acquisition and analysis. We thank O. Mazor and P. Gorelik from the Research Instrumentation Core Facility and Ludo Cacheux for engineering support. Core facility support is provided by NIH P30 grants HD18655 and NS072030. S.R.D. is supported by fellowships from the Vallee Foundation, by grants RO11DC016222 and U19 NS112953 from the National Institutes of Health and by the Simons Collaboration on the Global Brain. S.P. is supported by grant NS108410 from the National Institutes of Health. D.C. is supported by a Bertarelli Foundation Fellowship. S.L.P. is supported by grant DA036922 from the National Institutes of Health and the Stuart H.Q. and Victoria Quan Fellowship. G.I. is supported by the Armenise Foundation Career Development Award.
Author information
Authors and Affiliations
Contributions
S.L.P. and S.R.D. conceived and designed the experiments. S.L.P. performed imaging experiments and K.D. performed behavioural experiments. G.I. assisted with analysis and performed single-unit recordings to validate the TeLC experiment. D.B. generated reagents and performed infections and histology for the bulbar afferent experiments, and helped to modify and test reagents for the TeLC experiment. K.F. provided reagents and guidance for the TeLC experiment. S.L.P., G.I. and D.C. analysed the data. S.P. provided guidance about population analysis. S.L.P., G.I. and S.R.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Vijay Balasubramanian and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Volumetric population imaging of PCx L2 and L3 during wakefulness using rationally designed odour sets.
a, Left, cartoon of the volumetric multi-photon imaging approach used to characterize odour responses in PCx in wakeful, semi-paralysed mice (Methods). Right, approximate position of an imaging volume (green dotted line) in a typical experiment superimposed on a Nissl-stained coronal section through PCx. Scanning volumes were oriented to acquire similarly sized cortical populations in L2 and L3 (red dotted lines), despite decreased neuron density in L3 (Methods). Imaging was performed in the most anterior portion of the posterior PCx. b, Sample fields of view for a single imaging session. PCx L2 is depicted on top; PCx L3 on bottom. Segmentation masks associated with each layer are shown on the right. c, Global, clustered and tiled odour sets superimposed on the collection of odours constituting odour space as defined by principal components analysis (Methods). Global odours are indicated by black dots; tiled and clustered odour sets via the indicated colour code. d, Plot of the amount of molecular variance contributed by each additional principal component for each odour set in descriptor space; this analysis reveals that each odour set tiles odour space at a distinct level of resolution. e, Molecular structures and associated photoionization detector (PID) signals of the odours comprising the global, clustered and tiled odour sets. These PID traces are shown to illustrate the controlled kinetics of the olfactometer only; because detector reports depend on the ability of an odour to be photo-ionized, the relative amplitudes of the traces between odours are not meaningful. For example, heavy aliphatics elicit a minimal PID response because their photo-ionization energies lie outside the range of the detector; however, odours with low or absent PID traces still induced cortical activity in 5–20% of the imaged population, consistent with effective odour delivery. Five odours are shared between the global and clustered odour sets. These are indicated by bold lettering (and in c, as black circles with coloured edges). Colour code as in c.
Extended Data Fig. 2 Odour responses in PCx are substantially altered by anaesthesia.
a, Left, EEG power spectral density plot from an individual subject depicting differences in cortical state between ketamine–medetomidine anaesthesia and wakefulness (Methods). Under anaesthesia, the EEG signal is enriched in the delta band (0.5–4 Hz) at the expense of high frequency (40–100 Hz) gamma oscillations; by contrast, gamma activity increases and delta activity decreases during wakefulness. Right, summary of differences in EEG power content expressed as delta/gamma ratio during anaesthesia and wakefulness averaged from four subjects. Error bars indicate s.e.m. b, Comparison of the fraction of responsive neurons (obtained from the population of neurons that respond to at least one odour during the wakefulness) (Methods) to the tiled odour set in the same field of view (obtained from PCx L2 and PCx L3) during the awake state and under anaesthesia. Responses are defined according to auROC analysis (Methods). Each dot represents a single odour (L2: 504 neurons, L3: 418 neurons). c, Top, black trace represents heart rate (average over 10 s, non-overlapping windows) recorded from an awake mouse in the home cage. Blue traces are example raw heart rate (HR) signal indicating the range of heart rate fluctuations observed during the awake state. The high variability in heart rates (which span approximately 350 to 650 beats per min) reflects ongoing behaviour in the awake mouse. Bottom, as in the top panel, but for heart rate recorded during wakefulness and after induction of ketamine–medetomidine anaesthesia (Methods). Grey arrow indicates time of induction. Grey and red rectangles and associated inset traces are 20-s segments of real-time heart-rate signal. During wakefulness, fluctuations in heart rate remain within a physiologically normal range of 300–500 beats per min, without any detectible episodes of tachycardia (Methods). Periodic large-amplitude dips in the recorded heart rate during wakefulness reflect moments when pharmacological agents are being administered, which briefly interrupts the heart rate monitor.
Extended Data Fig. 3 PCx L3 neurons exhibit denser, broader and more reliable odour responses than neurons in PCx L2.
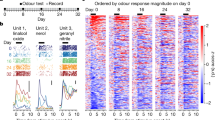
a, Examples of odour-evoked excitation and suppression in PCx. Each panel corresponds to a single cell-odour pair. Grey lines represent individual trials. Coloured overlays represent trial-mean activity. Shaded grey rectangles delimit the odour presentation period. b, Trial-averaged population response raster depicting odour-evoked activity in response to 22 odours (global odour set) across L2 and L3. Responses are ∆F/F0 with redder colours indicating excitatory transients and bluer colours indicating odour-evoked suppression. x axis is time; double vertical bars delimit 2-s odour presentation periods. c, Response types observed in L2 and L3 (clustered odour set). Individual panels correspond to clusters identified using a Gaussian mixture model (Methods). Grey traces correspond to trial-averaged cell-odour pairs. Coloured overlays represent mean response time course associated with each cluster. Right, fraction of all cell-odour pairs exhibiting excitation or suppression. d, Response amplitudes of cell-odour pairs obtained from PCx L3 depicted on a trial-by-trial basis. Each row represents the response of a given neuron to 10 consecutive presentations of the same odour. Neurons are sorted hierarchically using average linkage and correlation distance. Despite the presence of some habituation in response to several presentations of the same odorant across the experiment, habituation does not appear uniform across the neural population nor does it appear to dominate neural responses to odours. Different groups of neurons were identified with maximal responses to an odour peaking at different times across the experiment; see examples depicted on the right. Each row of traces corresponds to a single cell-odour pair. e, At the population level, odour responses do not uniformly habituate across the experiment. Top, cartoon depiction of procedure for determining change in response amplitude over the course of the experiment for a single cell odour pair. Middle and bottom, pooled data for all cell–odour pairs, sorted by layer. Red lines correspond to distribution means (clustered odour set). f, Lifetime sparseness distributions (used to quantify tuning breadth) (Methods) in L2 and L3 across all experiments (1 = perfectly odour selective, 0 = completely non-selective, *P < 0.01, permutation test on layer label). Distributions are built using all responsive neurons (significant response to at least one odour by auROC analysis) pooled by layer across all experiments (here and throughout, global: n = 3 mice, L2 = 854 neurons, L3 = 616 neurons; clustered: n = 3 mice, L2 = 867 neurons, L3 = 488 neurons; tiled: n = 3 mice, L2 = 427 neurons, L3 = 334 neurons). g, Population sparseness distributions (used to quantify response density) (Methods) in L2 and L3 (1 = few neurons active overall, 0 = all neurons active overall to an equal level). *P < 0.01, permutation test on layer label. h, Probability density distributions of coefficient of variation for all significant cell-odour pairs identified with auROC analysis. *P < 0.01, permutation test on layer label. i, Probability density distributions of ensemble correlations (that is, pairwise correlations between odour-evoked ensembles) between trial-averaged population odour responses in L2 (left) and L3 (middle). Dashed control curves indicate the distribution of ensemble correlations after shuffling odour labels independently across neurons. Ensemble correlations were determined independently for each mouse, and subsequently pooled. *P < 0.01, permutation test on odour label. L3 exhibits greater correlations at the population level than L2 (right). *P < 0.01, permutation test on layer label.
Extended Data Fig. 4 Cortical odour representations are stable from trial to trial and not chemotopically organized.
a, Left, pairwise odour chemical correlation matrices for the global, clustered and tiled odour sets. Rows and columns are sorted according to the chemical similarity between odours as assessed by hierarchical clustering (Methods). Middle and right, Pairwise correlation distances of single-trial, population representations for odours in the global, clustered, and tiled odour experiments in PCx L2 and L3 (and boutons for the tiled odour set). Rows and columns are sorted according to the chemical similarity between odours as on the left. Chemical colour code (x and y axis labels of matrices, indicating functional group associated with each group of molecules) is shown in the legend. R values indicate Pearson’s correlation to odour chemistry. b, Top, structured odour relationships persist from trial to trial over the course of the experiment. Blue line represents the similarity of two correlation distance matrices built from population responses obtained on consecutive trials. Grey dashed line indicates mean across all trial-pair comparisons (10 trials, 9 trial pairs; clustered odour set, L3). Bottom, chemistry-based odour relationships correspond to matched cortical relationships obtained on a trial-by-trial basis. Dashed grey line represents the similarity of chemical and neural activity distances on a trial-by-trial basis. c, Correspondence between odour structure in PCx L3 (clustered odour set) and odour chemistry using three different distance metrics (correlation distances, Euclidean distances and cosine distances). Distance matrices calculated from population activity are obtained using instantaneous ∆F/F0 over 130 ms increments (F0: baseline fluorescence averaged over a 1-s sliding window). Vertical lines delimit the 2-s odour presentation. d, Odour chemical relationships emerge within a few hundred milliseconds after odour onset and persist for several seconds after odour offset (see Extended Data Fig. 1e for associated PID traces). e, Example PCx L2 and L3 FOVs from a single mouse with each responsive neuron coloured according to its preferred odour in the clustered odour set. Neurons preferring odours belonging to different classes (legend) appear spatially intermingled in both L2 and L3. f, Contour plots of pairwise signal correlations, plotted with respect to distance in L2 and L3 for the clustered and tiled experiments. Darker colours indicate increased density (see margin distributions). Pearson’s r is overlaid and indicates no spatial organization of odour representations in PCx.
Extended Data Fig. 5 Lasso optimization identifies parsimonious sets of chemical descriptors that predict neural odour relationships.
a, Left, descriptors identified through training on one odour set also improve Pearson’s correlation (r) between corresponding chemical and neural distances for held-out sets of odours. C, clustered; G, global; T, tiled. A value of 1 in the matrix corresponds to no improvement from baseline Pearson’s r value after optimization. Baseline chemical-neural correlation is 0.22 for global; 0.48 for clustered; 0.37 for tiled (see Supplementary Table 1 for optimal descriptor sets). Right, reduction in mean-squared error (MSE) between chemical and neural odour pair distances for held-out odour sets (indicated below the x axis) after training on a single odour set (indicated above). Note that the five odours in common between the global and clustered odour sets (names in bold in Extended Data Fig. 2e) were discarded when evaluating performance on held-out data. The chemical features learned from the tiled odour set improved chemical–neural Pearson’s correlations in the clustered odour experiment but not the global odour experiment, consistent with the odours belonging to the tiled set covering only a limited region of chemical odour space (left). However, despite the limited chemical overlap between the tiled and global odour sets, training on the tiled odour set still improved the correspondence between odour chemistry and neural responses for the global odour set as assessed by a reduction in the mean-squared error (right). b, Identifying a subset of chemical descriptors (from the original superset used to define odour space) using Lasso optimization on odour distances improves the correspondence to cortical activity (Methods, Supplementary Table 1). Training data were derived from the bouton dataset, and testing was performed for bouton responses to held-out odours within the tiled odour set, and also to cortical responses of the tiled odour set. Data are mean ± s.e.m. over cross-validation folds. c, The same procedure as in b was performed on a limited subset of 15 semantically relevant descriptors that comprise the ‘molecular properties’ block of the Dragon database; these descriptors include metrics that reflect molecular properties associated with functional groups (for example, donor or acceptor atom surface area), molecular weight (for example, van der Waals molecular volume) or a combination of both, such as ‘hydrophilic factor’, and reflect the main axes of diversity in the tiled odour set. Most descriptors enriched in the olfactory bulb covary with molecular weight (red descriptors). Most descriptors enriched in PCx reflect the combined presence of a charged atom and variable number of carbon atoms along the aliphatic series of the tiled odour set (blue descriptors). Note that these descriptors differ from those identified when querying the entire Dragon set using Lasso optimization (Supplementary Table 1), as this limited set of targeted descriptors (selected because their semantic meaning is transparent) may not afford optimal predictions over neural data.
Extended Data Fig. 6 Functional imaging of OB axons in PCx via axonally targeted GCaMP6s.
a, Left, whole-mount depicting Tbx21-Cre-dependent expression of AAV PHP.eB hSynapsin1-FLEX-axon-GCaMP6s in OB projection neuron axons. GCaMP6s fluorescence is broadly distributed across piriform cortex. Right, coronal sections depicting GCaMP6s signal (green) in the mitral cell layer across the entire anterior-posterior extent of the olfactory bulb and cortex. Inset, bottom, GCaMP6s-labelled axons shown coursing through PCx L1a. Bottom left, en face image of L1a depicts dense and uniform distribution of axonal boutons. b, Difference heat map of a typical field-of-view (FOV) depicting baseline and odour-driven fluctuations in GCaMP6s signal. The strongest activation (light colour) is associated with axonal boutons. c, Time-averaged fluorescence signal of FOV in b. Overlay shows segmented ROIs corresponding to axonal boutons depicting increases (red) or decreases (blue) in fluorescence, averaged over multiple presentations of a single odour from the tiled odour set. d, Example average fluorescence from several boutons in a. Grey bar indicates odour delivery period, scale bar indicates response amplitude. For clarity, fluorescence time courses for each example bouton are offset along the y axis. e, Example bouton responses for the tiled odour set. Each row represents the trial-averaged response of a single bouton for two seconds during and after odour exposure (columns) depicted as z-scored ∆F/F0; rows are sorted hierarchically using correlation distance and average linkage. The functional group and carbon chain-length associated with each odour are indicated below each column; light-to-saturated gradient indicates progression from short-chain to long-chain odours. Note that, as has been observed previously for OB projection neurons, boutons exhibit a substantial amount of odour-driven suppression.
Extended Data Fig. 7 Bouton odour response properties.
a, Probability density distributions for boutons, PCx L2, and PCx L3 for signal correlations. b, Left, as in a, but for ensemble correlations. Right, for the top 5% most similar odour pairs identified in boutons, correlation for the same odour pairs in PCx. Ensemble responses in both PCx L2 and PCx L3 exhibit stronger similarity than boutons. c, d, Probability density distributions for boutons, PCx L2 and PCx L3, for lifetime and population sparseness. e, Cumulative neural variance explained with increasing numbers of principal components, indicating relatively higher dimensionality in boutons compared to PCx (that is, more uniform distribution of variance across principal components). f, Probability density distributions for boutons, PCx L2 and PCx L3 for coefficient of variation representing trial-to-trial response variability across cell–odour pairs. These data demonstrate that observed odour responses in boutons are more reliable than similar responses in the cortex. For a–f, only the tiled odour set is used. For lifetime sparseness, 1 = perfectly odour selective, 0 = completely non-selective. For population sparseness, 1 = few neurons responsive, 0 = all neurons equally responsive. Distributions are built using all responsive neurons/boutons (significant response to at least one odour by auROC analysis; boutons: 3160 ROIs across 6 subjects, PCx L2: 427 neurons across 3 subjects. PCx L3: 334 neurons across 3 subjects). Asterisk indicates significant difference between boutons and either L2 or L3: a, vs L2 P < 10−27; vs L3 P = 0.02; b, vs L2 P < 10−20; vs L3 P < 0.005; c, vs L2 P < 10−9; vs L3 P = 0.93; d, vs L2 P < 10−7 vs L3 P < 10−4; f, vs L2: P < 10−20; vs L3: P < 10−23; two-sided Wilcoxon rank sum test for all comparisons. g, Single-trial Z-scored ∆F/F0 for 1,000 boutons recorded in PCx L1a during presentation of 22 odours belonging to the tiled odour set indicated by black lines. Redder colours indicating excitatory transients, and bluer colours indicate odour-evoked suppression. h, Response types observed in boutons (tiled odour set). Individual panels correspond to clusters identified using a Gaussian mixture model (Methods). Grey traces correspond to trial-averaged bouton-odour pairs. Coloured overlays represent mean response time course associated with each cluster. Blue vertical lines mark periods of odour presentation. i, Fraction of total odour-driven bouton variance in each individual mouse that can be attributed to the shared across-mouse structure as quantified by distance covariance analysis (Methods).
Extended Data Fig. 8 Habituation-dishabituation test for assessing perceptual similarity of odour pairs.
a, Left, mice presented with new odours exhibit investigation that diminishes over several consecutive presentations of the same odorant. Subsequent presentation of a perceptually different odour reinstates investigation, and presentation of a similar odour has little effect. The extent to which two odorants are perceptually related is assessed by the magnitude of rekindled interest in the second odour after habituation has occurred to the first. b, Investigation times for two different odour triplets. Data are mean ± s.e.m. (n = 7 and n = 8 mice, respectively). After habituation to heptanal, investigation of the closely related octanal (1-carbon difference) only modestly increases. Presentation of butanal after habituation to octanal (4-carbon difference) induces greater investigation. For the second triplet, presentation of heptanal after habituation to heptanone (0-carbon difference, different functional group) induces greater investigation, whereas subsequent presentation of octanal after habituation to heptanal (1-carbon difference, same functional group) induces much less investigation.
Extended Data Fig. 9 Inhibition of the associative network through cell-autonomous expression of tetanus toxin light chain in excitatory PCx neurons.
a, Uniform infection of excitatory pyramidal neurons in PCx L2 and L3 with AAV-hSyn-FLEX-TeLC-P2A-NLS-dTom in an Emx1-Cre mouse. b, Left, coronal section through PCx indicating placement of recording electrode. Right, single-unit odour-evoked activity (grand-average of all excitatory responses deemed as significant by auROC analysis) in Emx1-Cre mice expressing TeLC or wild-type controls. Disruption of cortical recurrent excitation enhances odour-evoked excitation, consistent with disruption of feedback inhibition. Grey bar indicates odour presentation (n = 121 cell–odour pairs from two Emx1-Cre mice expressing TeLC; n = 229 cell–odour pairs from four mice). c–g, Probability density distributions for the TeLC experiment for signal and ensemble correlations, lifetime and population sparseness, and coefficient of variation (constructed as in Extended Data Fig. 7, here only for the tiled odour set). For lifetime sparseness, 1 = perfectly odour selective, 0 = completely non-selective. For population sparseness, 1 = few neurons responsive, 0 = all neurons equally responsive. Distributions are built using all responsive neurons (significant response to at least one odour by auROC analysis; TeLC L2: 435 neurons across 3 subjects. TeLC L3: 590 neurons across 3 subjects. PCx L2: 427 neurons across 3 subjects. PCx L3: 334 neurons across 3 subjects). Asterisk indicates TeLC is significantly different from PCx L2 or L3: c, L2 P < 10−8; L3 P < 10−198; d, L2 P < 10−46; L3 P < 10−55; e, L2 P < 10−05; L3 P < 10−37; f, L2 P < 10−7 L3 P < 10−8; g, L2: P < 10−10; L3: P < 10−4; two-sided Wilcoxon rank sum test for all comparisons.
Extended Data Fig. 10 Passive odour experience modifies odour relationships.
a, Correlation distance matrices for the tiled odour set obtained from odour-naive (same data as in Figs. 1–4) mice as well as mice passively exposed to a target mixture of two short-chain aldehydes and two short-chain ketones in the home cage (Methods, Fig. 4e, f). Passive experience with the mixture increases odour similarity specifically between mixture components (target comparisons indicated in the legend in blue), but not between target ketones and long-chain aldehydes or short-chain esters and short-chain acids with which mice had no previous experience (off-target comparisons indicated in legend in black, nave: 334 neurons, n = 3 mice; exposed: 742 neurons, n = 3 mice).
Supplementary information
Supplementary Table 1
Physiochemical descriptor sets that maximize correspondence between chemistry and neural activity. Sets of physiochemical descriptors identified for each cortical odour set experiment and the bouton experiment with the tiled odour set. Terms reflecting the semantic meaning of a subset of these descriptors are highlighted, with terms associated with functional groups shown in blue, and those associated with molecular weight shown in red; identified features can be positively or negatively weighted, and therefore be either correlated or anti-correlated with neural activity driven by each of the odour sets. These results demonstrate that both cortex and boutons encode information about wide array of chemical features.
Rights and permissions
About this article
Cite this article
Pashkovski, S.L., Iurilli, G., Brann, D. et al. Structure and flexibility in cortical representations of odour space. Nature 583, 253–258 (2020). https://doi.org/10.1038/s41586-020-2451-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2451-1
This article is cited by
-
Decomposition of an odorant in olfactory perception and neural representation
Nature Human Behaviour (2024)
-
Distinct information conveyed to the olfactory bulb by feedforward input from the nose and feedback from the cortex
Nature Communications (2024)
-
High-precision mapping reveals the structure of odor coding in the human brain
Nature Neuroscience (2023)
-
Olfactory perception prediction model inspired by olfactory lateral inhibition and deep feature combination
Applied Intelligence (2023)
-
The mnemonic basis of subjective experience
Nature Reviews Psychology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.