Abstract
The stem cell niche and the size of the root meristem in plants are maintained by intercellular interactions and signalling networks involving a peptide hormone, root meristem growth factor 1 (RGF1)1. Understanding how RGF1 regulates the development of the root meristem is essential for understanding stem cell function. Although five receptors for RGF1 have been identified2,3,4, the downstream signalling mechanism remains unknown. Here we report a series of signalling events that follow RGF1 activity. We find that the RGF1-receptor pathway controls the distribution of reactive oxygen species (ROS) along the developmental zones of the Arabidopsis root. We identify a previously uncharacterized transcription factor, RGF1-INDUCIBLE TRANSCRIPTION FACTOR 1 (RITF1), that has a central role in mediating RGF1 signalling. Manipulating RITF1 expression leads to the redistribution of ROS along the root developmental zones. Changes in ROS distribution in turn enhance the stability of the PLETHORA2 protein, a master regulator of root stem cells. Our results thus clearly depict a signalling cascade that is initiated by RGF1, linking this peptide to mechanisms that regulate ROS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All RNA-seq data from this study have been deposited in the National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO), with the accession number GSE108730. Source data for all graphs have been provided. A previous version of this work was deposited in the preprint depository server bioRxiv at https://doi.org/10.1101/244947. Source Data for Figs. 1–4 and Extended Data Figs. 1, 3, 5–10 are provided with the paper. All other data are available from the corresponding author upon reasonable request.
Code availability
All code from this study is available upon request.
References
Matsuzaki, Y., Ogawa-Ohnishi, M., Mori, A. & Matsubayashi, Y. Secreted peptide signals required for maintenance of root stem cell niche in Arabidopsis. Science 329, 1065–1067 (2010).
Ou, Y. et al. RGF1 INSENSITIVE 1 to 5, a group of LRR receptor-like kinases, are essential for the perception of root meristem growth factor 1 in Arabidopsis thaliana. Cell Res. 26, 686–698 (2016).
Song, W. et al. Signature motif-guided identification of receptors for peptide hormones essential for root meristem growth. Cell Res. 26, 674–685 (2016).
Shinohara, H., Mori, A., Yasue, N., Sumida, K. & Matsubayashi, Y. Identification of three LRR-RKs involved in perception of root meristem growth factor in Arabidopsis. Proc. Natl Acad. Sci. USA 113, 3897–3902 (2016).
Dunand, C., Crèvecoeur, M. & Penel, C. Distribution of superoxide and hydrogen peroxide in Arabidopsis root and their influence on root development: possible interaction with peroxidases. New Phytol. 174, 332–341 (2007).
Tsukagoshi, H., Busch, W. & Benfey, P. N. Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 143, 606–616 (2010).
Whitford, R. et al. GOLVEN secretory peptides regulate auxin carrier turnover during plant gravitropic responses. Dev. Cell 22, 678–685 (2012).
Meng, L., Buchanan, B. B., Feldman, L. J. & Luan, S. CLE-like (CLEL) peptides control the pattern of root growth and lateral root development in Arabidopsis. Proc. Natl Acad. Sci. USA 109, 1760–1765 (2012).
Aida, M. et al. The PLETHORA genes mediate patterning of the Arabidopsis root stem cell niche. Cell 119, 109–120 (2004).
Ishida, T. et al. SUMO E3 ligase HIGH PLOIDY2 regulates endocycle onset and meristem maintenance in Arabidopsis. Plant Cell 21, 2284–2297 (2009).
Li, S., Yamada, M., Han, X., Ohler, U. & Benfey, P. N. High-resolution expression map of the Arabidopsis root reveals alternative splicing and lincRNA regulation. Dev. Cell 39, 508–522 (2016).
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003).
Zuo, J., Niu, Q. W. & Chua, N. H. Technical advance: an estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J. 24, 265–273 (2000).
Galinha, C. et al. PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 449, 1053–1057 (2007).
Waszczak, C. et al. Sulfenome mining in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 111, 11545–11550 (2014).
Licausi, F., Ohme-Takagi, M. & Perata, P. APETALA2/ethylene responsive factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. New Phytol. 199, 639–649 (2013).
Shaikhali, J. et al. The redox-sensitive transcription factor Rap2.4a controls nuclear expression of 2-Cys peroxiredoxin A and other chloroplast antioxidant enzymes. BMC Plant Biol. 8, 48 (2008).
Dietz, K. J., Vogel, M. O. & Viehhauser, A. AP2/EREBP transcription factors are part of gene regulatory networks and integrate metabolic, hormonal and environmental signals in stress acclimation and retrograde signalling. Protoplasma 245, 3–14 (2010).
Licausi, F. et al. Oxygen sensing in plants is mediated by an N-end rule pathway for protein destabilization. Nature 479, 419–422 (2011).
Welsch, R., Maass, D., Voegel, T., Dellapenna, D. & Beyer, P. Transcription factor RAP2.2 and its interacting partner SINAT2: stable elements in the carotenogenesis of Arabidopsis leaves. Plant Physiol. 145, 1073–1085 (2007).
Wang, Z. P. et al. Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol. 16, 144 (2015).
Maeda, H. et al. Fluorescent probes for hydrogen peroxide based on a non-oxidative mechanism. Angew. Chem. Int. Edn 43, 2389–2391 (2004).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank I. Taylor, J. Dickinson, E. Pierre-Jerome, K. Lehner and C. Winter for comments on the manuscript; C. Wilson for help with generating overexpression lines; G. Yang for help in identifying CRISPR mutants; K. Sugimoto for HYP2-GFP seeds; Y. Matsubayashi for rgfr1/2/3 seeds; R. Heidstra for gPLT2-YFP and pPLT2-CFP seeds; N.-H. Chua for the pMDC7 vector; The Duke Genome Sequencing Center for sequencing Illumina libraries; the Plant Tech Core Facility in the Agricultural Biotechnology Research Center for generating the CRISPR construct; and the Transgenic Plant Laboratory at Academia Sinica for transforming the CRISPR construct into plants. This work was funded by the Howard Hughes Medical Institute and the Gordon and Betty Moore Foundation (through grant GBMF3405), the US National Institutes of Health (MIRA 1R35GM131725) to P.N.B., and Academia Sinica, Taiwan, to M.Y.
Author information
Authors and Affiliations
Contributions
M.Y. and P.N.B. conceptualized the study; M.Y. performed all experiments; X.H. performed the computational analyses; all authors wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Yoshikatsu Matsubayashi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Expression of meristematic-zone marker and transcriptome analysis upon RGF1 treatment.
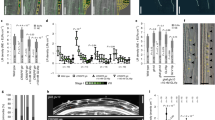
a, b, Confocal images of HPY2-GFP roots 24 h after treatment with water (mock; a) or 20 nM RGF1 (b). Seedlings were grown on MS plates for seven days before treatment. Left, PI-stained roots; right, GFP signals. White and blue arrowheads indicate the junction between the meristematic and elongation zones. Scale bar, 50 µm. c, Area of HPY2-GFP expression (in µm2; n = 8 independent roots; P < 2.1 × 10−7). Bar graphs show means. Error bars are ± s.d. Dots indicate each data point. P values are calculated by two-sided Student’s t-test. d, Method of RNA extraction following RGF1 treatment.
Extended Data Fig. 2 GO categories that are enriched upon RGF1 treatment.
These highly significantly enriched GO categories within lists of genes are regulated by RGF1 (FDR-adjusted P < 0.001), and include glutathione transferase activity (FDR-adjusted P = 4.1 × 10−6, red) and oxidoreductase activity (FDR-adjusted P = 0.00039, red). See Supplementary Table 2 (enriched GO categories upon RGF1 treatment). P values for GO enrichment analysis are based on Fisher’s exact test, with the sample size being all genes in the genome and using a Benjamini–Yekutieli FDR for multiple testing correction.
Extended Data Fig. 3 H2O2 levels after inducible overexpression of RITF1 and RGF1 treatment.
a, Confocal images of H2O2-BES-Ac stained roots, with or without XVE–RITF1 expression, in a wild-type (Col-0) background 4 h after treatment with water (mock), 10 μM oestradiol or 100 nM RGF1. b, Quantification of H2O2-BES-Ac intensity in the meristematic zone (n = 6 independent samples; *P < 0.0005). Bar graphs show means. Error bars are ± s.d. Dots indicate each data point. P values are calculated by two-sided Student’s t-test.
Extended Data Fig. 4 Localization of gPLT2–YFP, NBT and H2O2-BES-Ac staining after RGF1 treatment.
a–t, Localization of gPLT2–YFP (a–h), NBT staining (i–n) and H2O2-BES-Ac staining (o–t), 4 h after treatment with water (mock; a) or 100 nM RGF1 (b), 6 h after treatment with water (mock; c, i, o) or 100 nM RGF1 (d, j, p), 8 h after treatment with water (mock; e, k, q) or 100 nM RGF1 (f, l, r), or 10 h after treatment with water (mock; g, m, s) or 100 nM RGF1 (h, n, t). Blue arrowheads indicate the junction between the meristematic and elongation zones. Scale bar, 50 µm. Seedlings were grown on MS agar plates for seven days before treatment. Experiments were independently repeated three times with similar results.
Extended Data Fig. 5 Time course of gPLT2–YFP localization and NBT and H2O2-BES-Ac staining.
a, Distance (in μm) of gPLT2–YFP localization from quiescent-centre cells (n = 5 independent roots; P < 5.7 × 10−6). b, Total intensity of NBT staining in the meristematic zone (n = 8 independent roots; P < 0.0003). c, Average intensity of H2O2-BES-Ac staining in the elongation zone (n = 5 independent roots; P < 0.003). Bar graphs show means. Error bars are ± s.d. Dots indicate each data point. P values are calculated by two-sided Student’s t-test. Experiments were independently repeated three times with similar results.
Extended Data Fig. 6 Phenotype of rift mutants.
a, Root growth (in mm) of wild-type (Col), ritf1-1 (CRISPR mutant) and ritf1-2 (SALK line) seedlings from 4 to 8 days after germination (n = 21 independent roots). b, Confocal images of wild-type, ritf1-1 (CRISPR mutant) and ritf1-2 (Salk line) roots stained with PI. c, Percentage increase (in which 100% is the number of cells in the mock-treated case) in the number of cells in the meristematic zone of wild-type, ritf1-1 and ritf1-2 roots 24 h after mock treatment or 5 nM RGF1 treatment (n = 7 independent roots, *P < 5.4 × 10−6). d, Light microscope images of roots of wild-type, ritf1-3 and ritf1-2 roots stained with NBT 24 h after treatment with 5 nM RGF1. Scale bars, 50 µm. Blue arrowheads show the junction between the meristematic and elongation zones. e, Quantification of NBT staining intensity in the meristematic zone in wild-type, ritf1-3 and ritf1-2 roots after treatment with 5 nM RGF1 (n = 8 independent roots; *P < 0.003). Scale bars, 50 μm. Blue and white arrowheads show the junction between the meristematic and elongation zones. Bar and line graphs show means. Error bars are ± s.d. Dots indicate each data point. P values are calculated by two-sided Student’s t-test.
Extended Data Fig. 7 Expression of pPLT2-CFP and gPLT2-YFP upon RGF1 treatment.
a, b, Confocal images showing pPLT2-CFP expression (cyan; a) and gPLT2-YFP expression (green; b) 24 h after treatment with 20 nM RGF1. Red, PI staining. Scale bar, 50 µm. Arrow heads show the junction between the meristematic and elongation zones. c, Extent (in µm) of gPLT2-YFP expression from quiescent-centre cells (n = 5 independent roots; P < 2.5 × 10−13). Bar graphs show means. Error bars are ± s.d. Dots indicate each data point. P values are calculated by two-sided Student’s t-test.
Extended Data Fig. 8 Localization of PLT2 protein after RGF1 and/or H2O2 treatment.
a–d, Confocal images showing gPLT2–YFP expression 24 h after treatment with water (mock), 20 nM RGF1, 20 nM RGF1 with 500 µM H2O2, or 500 µM H2O2. gPLT2-YFP seedlings were grown for seven days on MS agar plates before treatment. Scale bar, 50 µm. Arrowheads show the extent of gPLT2-YFP expression. e, Distance (in µm) of PLT2 localization as measured from quiescent-centre cells (n = 6, *P < 0.0002). Bar graphs show means. Error bars are ± s.d. Dots indicate each data point. P values calculated by two-sided Student’s t-test.
Extended Data Fig. 9 Phenotypes resulting from RITF1 overexpression in plt2 mutants.
a, NBT-stained roots, with or without XVE–RITF1 expression, in a wild-type or plt2 background 24 h after treatment with water (mock) or 10 μM oestradiol. b, Quantification of NBT staining intensity in the differentiation zone with or without XVE–RITF1 in wild-type and plt2 roots (n = 8 independent roots; *P < 5.4 × 10−6). c, Confocal images of PI-stained roots with or without XVE-RITF1, in a wild-type or plt2 background, 24 h after mock treatment or treatment with 10 μM oestradiol. d, Number of cells in the meristematic zone, with or without XVE–RITF1, in wild-type and plt2 roots 24 h after mock or 10 μM oestradiol treatment (n = 7 independent roots; *P < 4.3 × 10−5). Scale bars, 50 μm. White and blue arrowheads indicate the junction between the meristematic and elongation zones. Bar graphs show means. Error bars are ± s.d. Dots indicate each data point. P values are calculated by two-sided Student’s t-test.
Extended Data Fig. 10 Phenotype of plt2 roots upon RGF1 treatment.
a, Confocal images of PI-stained wild-type and plt2 roots 24 h after treatment with water (mock) or 20 nM RGF1. Scale bar, 50 µm. White arrowheads show junctions between the meristematic and elongation zones. b, Number of cells in the meristematic zone 24 h after mock or 5 nM RGF1 treatment (n = 6 independent roots; *P < 4.9 × 10−7). c, Light microscope images of roots from wild-type and plt2 roots stained with NBT. Seedlings were grown on MS agar plates for 7 days before treatment with water (mock) or 20 nM RGF1. d, Total intensity of NBT staining in the differentiation zone of wild-type and plt2 roots 24 h after treatment with water (mock) or 20 nM RGF1 (n = 8 independent roots; *P < 0.0003). Scale bars, 50 μm. White arrowheads show the junction between the meristematic and elongation zones. Bar graphs show means. Error bars are ± s.d. Dots indicate each data point. P values are calculated by two-sided Student’s t-test.
Supplementary information
Supplementary Table 1:
Differential expression upon RGF1 treatment. Expression of 583 genes is significantly altered in the RNA-seq data between RGF1 and mock treatment in three biological replicates (FDR-adjusted p value < 0.1). The differentially expressed genes were identified using the statistical library DEseq2, which models the read counts with a negative binomial distribution. The p-value is adjusted for multiple testing using Benjamini-Hochberg FDR procedure. The 3 biological replicates are typical for RNA-seq study, with both statistical and practical considerations.
Supplementary Table 2:
Enriched GO categories upon RGF1 treatment. List of significant Gene Ontology (GO) categories (FDR≤ 1) and AGI numbers in the GO categories. The p-value of GO enrichment analysis was based on Fisher’s exact test, with the sample size of all genes in the genome and Benjamini–Yekutieli FDR for multiple testing correction.
Supplementary Table 3:
All gene expression upon RGF1 treatment. Expression of all annotated Arabidopsis genes in mock or RGF1 treatment (Value is FPKM).
Source data
Rights and permissions
About this article
Cite this article
Yamada, M., Han, X. & Benfey, P.N. RGF1 controls root meristem size through ROS signalling. Nature 577, 85–88 (2020). https://doi.org/10.1038/s41586-019-1819-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1819-6
This article is cited by
-
CAFRI-Arabidopsis: An Intuitive Web-Based Functional Redundancy Inspector in Arabidopsis
Journal of Plant Biology (2024)
-
Chemical imaging reveals diverse functions of tricarboxylic acid metabolites in root growth and development
Nature Communications (2023)
-
Dosage differences in 12-OXOPHYTODIENOATE REDUCTASE genes modulate wheat root growth
Nature Communications (2023)
-
Cross-species transcriptomic analyses reveals common and opposite responses in Arabidopsis, rice and barley following oxidative stress and hormone treatment
BMC Plant Biology (2022)
-
Genome-wide identification and expression analysis of the plant-specific PLATZ gene family in Tartary buckwheat (Fagopyrum tataricum)
BMC Plant Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.