Abstract
Inactivation of ARID1A and other components of the nuclear SWI/SNF protein complex occurs at very high frequencies in a variety of human malignancies, suggesting a widespread role for the SWI/SNF complex in tumour suppression1. However, the underlying mechanisms remain poorly understood. Here we show that ARID1A-containing SWI/SNF complex (ARID1A–SWI/SNF) operates as an inhibitor of the pro-oncogenic transcriptional coactivators YAP and TAZ2. Using a combination of gain- and loss-of-function approaches in several cellular contexts, we show that YAP/TAZ are necessary to induce the effects of the inactivation of the SWI/SNF complex, such as cell proliferation, acquisition of stem cell-like traits and liver tumorigenesis. We found that YAP/TAZ form a complex with SWI/SNF; this interaction is mediated by ARID1A and is alternative to the association of YAP/TAZ with the DNA-binding platform TEAD. Cellular mechanotransduction regulates the association between ARID1A–SWI/SNF and YAP/TAZ. The inhibitory interaction of ARID1A–SWI/SNF and YAP/TAZ is predominant in cells that experience low mechanical signalling, in which loss of ARID1A rescues the association between YAP/TAZ and TEAD. At high mechanical stress, nuclear F-actin binds to ARID1A–SWI/SNF, thereby preventing the formation of the ARID1A–SWI/SNF–YAP/TAZ complex, in favour of an association between TEAD and YAP/TAZ. We propose that a dual requirement must be met to fully enable the YAP/TAZ responses: promotion of nuclear accumulation of YAP/TAZ, for example, by loss of Hippo signalling, and inhibition of ARID1A–SWI/SNF, which can occur either through genetic inactivation or because of increased cell mechanics. This study offers a molecular framework in which mechanical signals that emerge at the tissue level together with genetic lesions activate YAP/TAZ to induce cell plasticity and tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Mass spectrometry data can be found in Supplementary Table 1. Source Data for Figs. 1, 2, 4 and Extended Data Figs. 2–5, 7, 8 can be found in the online version of the paper. Uncropped images of immunoblots can be found in Supplementary Fig. 1. All relevant data are included in the manuscript as Source Data or Supplementary Information; all other data are available from the corresponding authors upon reasonable request.
References
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Totaro, A., Panciera, T. & Piccolo, S. YAP/TAZ upstream signals and downstream responses. Nat. Cell Biol. 20, 888–899 (2018).
Panciera, T., Azzolin, L., Cordenonsi, M. & Piccolo, S. Mechanobiology of YAP and TAZ in physiology and disease. Nat. Rev. Mol. Cell Biol. 18, 758–770 (2017).
Rafiee, M. R., Girardot, C., Sigismondo, G. & Krijgsveld, J. Expanding the circuitry of pluripotency by selective isolation of chromatin-associated proteins. Mol. Cell 64, 624–635 (2016).
Ege, N. et al. Quantitative analysis reveals that actin and Src-family kinases regulate nuclear YAP1 and its export. Cell Syst. 6, 692–708 (2018).
Chen, H. I. & Sudol, M. The WW domain of Yes-associated protein binds a proline-rich ligand that differs from the consensus established for Src homology 3-binding modules. Proc. Natl Acad. Sci. USA 92, 7819–7823 (1995).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Wang, H. et al. BRCA1/FANCD2/BRG1-driven DNA repair stabilizes the differentiation state of human mammary epithelial cells. Mol. Cell 63, 277–292 (2016).
Cordenonsi, M. et al. The Hippo transducer TAZ confers cancer stem cell-related traits on breast cancer cells. Cell 147, 759–772 (2011).
Zanconato, F., Cordenonsi, M. & Piccolo, S. YAP/TAZ at the roots of cancer. Cancer Cell 29, 783–803 (2016).
Eroglu, E. et al. SWI/SNF complex prevents lineage reversion and induces temporal patterning in neural stem cells. Cell 156, 1259–1273 (2014).
Panciera, T. et al. Induction of expandable tissue-specific stem/progenitor cells through transient expression of YAP/TAZ. Cell Stem Cell 19, 725–737 (2016).
Zhang, N. et al. The Merlin/NF2 tumor suppressor functions through the YAP oncoprotein to regulate tissue homeostasis in mammals. Dev. Cell 19, 27–38 (2010).
Benhamouche, S. et al. Nf2/Merlin controls progenitor homeostasis and tumorigenesis in the liver. Genes Dev. 24, 1718–1730 (2010).
Bakiri, L. & Wagner, E. F. Mouse models for liver cancer. Mol. Oncol. 7, 206–223 (2013).
Plessner, M., Melak, M., Chinchilla, P., Baarlink, C. & Grosse, R. Nuclear F-actin formation and reorganization upon cell spreading. J. Biol. Chem. 290, 11209–11216 (2015).
Grosse, R. & Vartiainen, M. K. To be or not to be assembled: progressing into nuclear actin filaments. Nat. Rev. Mol. Cell Biol. 14, 693–697 (2013).
Baarlink, C., Wang, H. & Grosse, R. Nuclear actin network assembly by formins regulates the SRF coactivator MAL. Science 340, 864–867 (2013).
Rando, O. J., Zhao, K., Janmey, P. & Crabtree, G. R. Phosphatidylinositol-dependent actin filament binding by the SWI/SNF-like BAF chromatin remodeling complex. Proc. Natl Acad. Sci. USA 99, 2824–2829 (2002).
Miralles, F., Posern, G., Zaromytidou, A. I. & Treisman, R. Actin dynamics control SRF activity by regulation of its coactivator MAL. Cell 113, 329–342 (2003).
Zanconato, F. et al. Genome-wide association between YAP/TAZ/TEAD and AP-1 at enhancers drives oncogenic growth. Nat. Cell Biol. 17, 1218–1227 (2015).
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013).
Benham-Pyle, B. W., Pruitt, B. L. & Nelson, W. J. Mechanical strain induces E-cadherin-dependent Yap1 and β-catenin activation to drive cell cycle entry. Science 348, 1024–1027 (2015).
Skibinski, A. et al. The Hippo transducer TAZ interacts with the SWI/SNF complex to regulate breast epithelial lineage commitment. Cell Rep. 6, 1059–1072 (2014).
Komuro, A., Nagai, M., Navin, N. E. & Sudol, M. WW domain-containing protein YAP associates with ErbB-4 and acts as a co-transcriptional activator for the carboxyl-terminal fragment of ErbB-4 that translocates to the nucleus. J. Biol. Chem. 278, 33334–33341 (2003).
Azzolin, L. et al. Role of TAZ as mediator of Wnt signaling. Cell 151, 1443–1456 (2012).
Sif, S., Saurin, A. J., Imbalzano, A. N. & Kingston, R. E. Purification and characterization of mSin3A-containing Brg1 and hBrm chromatin remodeling complexes. Genes Dev. 15, 603–618 (2001).
Guan, B. et al. Mutation and loss of expression of ARID1A in uterine low-grade endometrioid carcinoma. Am. J. Surg. Pathol. 35, 625–632 (2011).
Maherali, N. et al. A high-efficiency system for the generation and study of human induced pluripotent stem cells. Cell Stem Cell 3, 340–345 (2008).
Stewart, S. A. et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA 9, 493–501 (2003).
Hughes, C. S. et al. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 10, 757 (2014).
Gao, X. et al. ES cell pluripotency and germ-layer formation require the SWI/SNF chromatin remodeling component BAF250a. Proc. Natl Acad. Sci. USA 105, 6656–6661 (2008).
Schuler, M., Dierich, A., Chambon, P. & Metzger, D. Efficient temporally controlled targeted somatic mutagenesis in hepatocytes of the mouse. Genesis 39, 167–172 (2004).
Zhu, Y. et al. Ablation of NF1 function in neurons induces abnormal development of cerebral cortex and reactive gliosis in the brain. Genes Dev. 15, 859–876 (2001).
Azzolin, L. et al. YAP/TAZ incorporation in the β-catenin destruction complex orchestrates the Wnt response. Cell 158, 157–170 (2014).
Morsut, L. et al. Negative control of Smad activity by ectodermin/Tif1γ patterns the mammalian embryo. Development 137, 2571–2578 (2010).
Rempe, D. et al. Synapsin I Cre transgene expression in male mice produces germline recombination in progeny. Genesis 44, 44–49 (2006).
Acknowledgements
We thank A. Fujimura for help with neuron preparation; G. Della Giustina for micropattern fabrication; V. Guzzardo for histology; C. Frasson and G. Basso for FACS; D. M. Livingston for HMECs and plasmids; D. J. Pan, M. Giovannini, Z. Wang, P. Chambon, and I. De Curtis and R. Brambilla for gifts of mice; R. Treisman for ACTB (encoding β-actin) cDNAs; L. Naldini for plasmids; S. Dupont for performing the initial experiments leading to biochemical identification of SWI/SNF and for the protocol to perform F-actin pull-down; Gianluca Grenci and Mona Suryana (MBI-Singapore) and the MBI microfabrication facility team for the supply of quartz masks. This work is supported by AIRC Special Program Molecular Clinical Oncology ‘5 per mille’, by an AIRC PI-Grant, by a MIUR-FARE grant, and by Epigenetics Flagship project CNR-MIUR grants to S.P. This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (DENOVOSTEM grant agreement No 670126 to S.P.).
Reviewer information
Nature thanks M. Sudol, P. Wade and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
L.C. carried out experiments in vitro, and L.A. carried out experiments on mice. Roles of other coauthors: D.D.B., molecular biology and IFs; D.D.B. and R.L.X., liver experiments; G.Ba. and F.Z., molecular biology and preparation of samples for ChIP–MS; L.C. and T.P., neuronal reprogramming; S.G., hydrogel preparation; G.S. and J.K. for mass spectroscopy; M.F., histology and histopathological evaluations; G.Br. and A.G., microfabrication. S.P. and M.C. conceived the initial hypothesis and experimental design, and planned, discussed and organized the work. L.C., L.A., F.Z., M.C. and S.P. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Interaction between YAP and the SWI/SNF complex.
a, Proteomic analyses of endogenous YAP/TAZ-binding partners reveal interactions with endogenous components of the SWI/SNF complex (green). Red, the used bait. R1 and R2 are the results from n = 2 biologically independent samples. See Supplementary Table 1. b, YAP(5SA) was immunoprecipitated from lysates of MCF10A cells stably expressing Flag-tagged YAP(5SA) using an anti-Flag antibody, and co-precipitating endogenous components of the SWI/SNF complex were detected by western blot. As a negative control, immunoprecipitation (IP) was repeated with cells transduced with empty vector. GAPDH serves as a loading control for inputs (right). c, HEK293T cells were transfected with independent siRNAs against the indicated genes (ARID1A in lanes 5 and 6; BAF53A in lanes 7 and 8; SNF5 in lanes 9 and 10) and control siRNAs ((siCo) lanes 1–4) and with plasmids encoding HA–YAP(5SA) (all lanes) and Flag–BRG1 (lanes 3–10), as indicated. Cell lysates were subjected to anti-Flag immunoprecipitation and co-precipitating proteins were checked by western blot. ARID1A depletion impairs the interaction between YAP and BRG1, but it had no effect on the association of BRG1 with BAF53A (lanes 5 and 6). Depletion of BAF53A (lanes 7 and 8) or SNF5 (lanes 9 and 10) had no effect on the interaction between YAP and BRG1. ARID1A blot, top band represents the full-length ARID1A. Input ARID1A was from a separate gel. d, Western blots of the inputs of the immunoprecipitation experiment shown in Fig. 1b. HEK293T cells were transfected with control (Co.) siRNA or siRNA against ARID1A and with plasmids encoding HA–YAP(5SA) and Flag–BRM, as indicated. e, Western blots of the inputs of the immunoprecipitation experiment shown in Fig. 1c. HEK293T cells were transfected with control siRNAs or with a siRNA mix against BRG1 and BRM. f, DNase-treated nucleus preparations from HEK293T cells were subjected to sequential salt extraction and fractions were analysed by western blot (left, lanes 1–4). The unsonicated, chromatin-free 0-mM NaCl fraction was incubated with GST–YAP or GST protein (negative control), immobilized on a glutathione resin, and proteins that were pulled down were analysed by western blot (right, lanes 5 and 6). g, HEK293T cells were transfected with siRNAs against the indicated genes and with plasmids encoding HA–YAP(5SA) and Flag–BRG1, as indicated. Cell lysates were subjected to anti-Flag immunoprecipitation and co-precipitating proteins were checked by western blot. h, Western blot of recombinant V5–ARID1A pulled down by GST–YAP or GST–TAZ, immobilized on a glutathione resin. GST protein was used as a negative control. Input, a fraction of V5–ARID1A used for the pull-down experiments. i, HEK293T cells were transfected with plasmids encoding empty vector (e.v.) or Flag–YAP(WT) or WW-domain mutants, as indicated. Cell lysates were subjected to anti-Flag immunoprecipitation and western blot analysis of endogenous ARID1A. GAPDH serves as a loading control in inputs. j, Flag–TAZ was immunoprecipitated from lysates of HEK293T cells transfected with Flag-tagged TAZ(WT) or TAZ(ΔWW) using an anti-Flag antibody, and co-precipitating endogenous ARID1A was detected by western blot only with TAZ(WT). As a negative control, immunoprecipitation was repeated using HEK293T cells transfected with empty vector. k, HEK293T cells were transfected with plasmids encoding Flag–YAP(WT) (all lanes) and either V5–ARID1A(WT) or V5–ARID1A(PPxA) mutant, as indicated. Cell lysates were subjected to anti-Flag immunoprecipitation and western blot analysis of V5–ARID1A. We notice that other SWI/SNF components (such as BRG1 or SNF5) also carry PPxY motifs; although these components are by themselves not essential for the association with YAP/TAZ, the presence of a second WW motif in YAP (although not in TAZ) raises the possibility of stronger, cooperative associations between YAP and other elements of the SWI/SNF complex. b, c, f–k Panels display representative experiments, repeated independently two (c, f–k) or three (b) times, all with similar results.
Extended Data Fig. 2 Effect of ARID1A depletion on YAP/TAZ levels, localization and activity.
a, Results of luciferase assays with the 8×GTIIC-Lux reporter in HEK293 cells transfected with empty or YAP-expressing vectors and the indicated siRNAs. Data are normalized to control siRNA- and empty vector-transfected cells and are presented as mean + s.d. of n = 3 biologically independent samples. b, Results of luciferase assays with the 8×GTIIC-Lux reporter in HEK293 cells reconstituted with either YAP(WT) or YAP(WW1mut) and transfected with the indicated siRNAs. Data are normalized to control siRNA-transfected cells and are presented as mean + s.d. of n = 3 biologically independent samples. c, qPCR analyses of the YAP/TAZ targets ANKRD1, CYR61 and PTX3 in MCF10A cells transfected as indicated. Data are mean + s.d. of n = 3 biologically independent samples. d, Western blot analysis of ARID1A, E-cadherin and vimentin from lysates of MCF10A cells transfected with the indicated siRNAs. e, qPCR analyses of CTGF (left) and ARID1B (right) expression in MCF10A cells transfected as indicated. Data are mean + s.d. of n = 3 biologically independent samples. f, Representative confocal images (left) and quantifications (right; >100 cells per condition) of YAP/TAZ localization in MCF10A cells transfected with the indicated siRNAs. g, Western blot analysis of YAP, TAZ and YAP phosphorylated at the key Hippo/LATS target site (p-YAP S127) in lysates of MCF10AT cells transfected with the indicated siRNAs. P values were determined by unpaired two-sided Student’s t-test; n.s., not significant. All panels display representative experiments, repeated independently two (d, e, g) or three (a–c, f) times with similar results.
Extended Data Fig. 3 YAP and TAZ are required for the biological effects of SWI/SNF depletion in HMECs.
a, b, HMECs were transduced with the indicated shRNA-encoding vectors and collected for protein extraction (a) or RNA extraction (b). a, Western blot of BRG1, TAZ and epithelial (ECAD) and mesenchymal (vimentin) markers. b, qPCR analyses of mesenchymal (TWIST1) and epithelial (KRT18) markers. Data are mean + s.d. of n = 3 biologically independent samples. Continuation of Fig. 2a. c–e, HMECs were transduced with the indicated shRNA-encoding vectors and/or transfected with the indicated siRNAs and collected for RNA extraction. qPCR analyses of the indicate genes are shown. Data are mean + s.d. of n = 3 biologically independent samples. f, Mammospheres formed by HMECs transduced with the indicated shRNAs and transfected with indicated siRNAs. Data are mean + s.d. of n = 6 biologically independent samples. g, HMECs were transduced with the indicated shRNA-encoding vectors and analysed for their CD44 and CD24 immunophenotype. Quantification of the percentage of cells that displayed either a CD44highCD24low (stem-like mesenchymal cells) or CD44lowCD24high (differentiated epithelial cells) profile8. P values were determined by unpaired two-sided Student’s t-test. All panels display representative experiments, repeated independently three times with similar results.
Extended Data Fig. 4 SWI/SNF depletion potentiates YAP-induced reprogramming of neurons into NSCs.
a, Efficiency of Brm and Arid1a downregulation in neurons transduced with the indicated shRNA-encoding vectors, as measured by qPCR. Data are mean + s.d. of n = 3 biologically independent samples. A representative experiment repeated twice with similar results is shown. b, c, Related to Fig. 2c. Neurons were infected with doxycycline-inducible YAP-encoding vectors or empty vector and the indicated shRNA-encoding lentiviral vectors. b, Representative images of the cultures after 14 days in NSC medium with doxycycline. Scale bar, 300 μm. c, Quantification of the emerging (P0) neurospheres. Data are mean + s.e.m. of four independent experiments; *P = 0.03 for comparisons between YAP(WT)-expressing neurons transduced with control shRNA (shCo.) and Brm shRNAs (shBrm) or between YAP(WT)-expressing neurons transduced with control shRNA and Arid1a shRNAs. d, e, Effect of Arid1a depletion on YAP-induced reprogramming of neurons. d, Syn1cre drives Arid1a knockout specifically in neurons as shown by genotyping. Genomic DNA from neurons was compared to genomic DNA from the tail of the same Syn1creArid1afl/+ mouse. PCR bands are shown for the indicated alleles. e, Control (Arid1a+/+) and Arid1a+/− (from Syn1creArid1afl/+ mice) neurons were infected with inducible YAP-encoding vectors. Left, Representative images of P0 neurospheres that emerged from these cultures after doxycycline treatment in NSC medium. Scale bar, 300 μm. Right, quantification of P0 neurospheres that emerged from these cultures after doxycycline treatment in NSC medium. Data are mean + s.e.m. of four independent experiments. YAPS94A serves as negative control. e complements Fig. 2c and Extended Data Fig. 4b, c, which show comparable results between shRNA and genetic attenuation of Arid1a. P values were determined by unpaired two-sided Student’s t-test (a) and by two-sided Mann–Whitney U-test (c, e).
Extended Data Fig. 5 Effect of ARID1A depletion in hepatocytes on tumour formation.
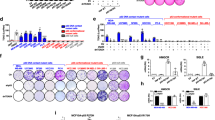
a, qPCR analysis of Nf2 and Arid1a expression in the livers of control (n = 6 mice), and Nf2 (n = 6 mice), Arid1a (n = 7 mice) and Nf2/Arid1a (n = 7 mice) liver mutant (LKO) mice, four months after tamoxifen treatment. All animals were included. Mean and data for individual mice are shown. b, Livers of control (Arid1afl/fl) and Arid1a LKO (AlbcreERT2Arid1afl/fl) mice were collected two weeks after tamoxifen treatment, and genomic DNA and proteins were extracted using standard procedures. Representative results are shown, experiments were repeated on four mice for each genotype. Left, PCR analysis of the indicated alleles. Right, western blots of GAPDH (loading control) and ARID1A. c, YAP immunohistochemistry (IHC) staining in control and Nf2 mutant livers. Scale bars, 40 μm. Representative images of experiments that were independently replicated using three mice for each genotype, with similar results. d, Continuation of Fig. 2e. Representative cytokeratin (CK; top) and Ki-67 (bottom) stainings of sections of livers of the indicated genotypes (same genotypes as in Fig. 1d, e and Extended Data Fig. 5a). Note intrahepatic cholangiocarcinomas (iCCA; CK+Ki-67+) and hepatocellular carcinomas (HCC; Ki-67+CK−) were found only in livers from Nf2/Arid1a LKO mice. Scale bars, 100 μm. Representative images are shown, experiments independently replicated for all of the mice of each genotype described in a, with similar results. e, qPCR analysis of selected genes of livers of mice with the indicated genotypes. All animals were included. Data are normalized to Nf2/Arid1a LKO mice. Data are mean + s.d. for same number of mice per genotype as in a. f, Continuation of Fig. 2f. Control, Arid1a LKO and Arid1a/Yap/Taz LKO mice were treated with tamoxifen and were then fed a DDC-containing diet for six weeks. CK (top; scale bars, 40 μm) and Ki-67 (bottom; scale bars, 20 μm) stainings of liver sections from the indicated mice. Note the presence of early cholangiocarcinoma lesions (CK+Ki-67+) in the Arid1a LKO mice and their absence upon concomitant YAP/TAZ loss (that is, in the Arid1a/Yap/Taz LKO mice). Asterisks indicate porfirin deposits, which are typically present in the liver of mice treated with DDC. Representative images are shown, experiments were independently replicated for all of the mice of each genotype (same number of mice as in Fig. 2f), with similar results. g, Representative qPCR analysis of Afp expression in the livers of control (n = 4), Arid1a LKO (n = 5), Arid1a/Yap/Taz LKO (n = 5) mice treated with tamoxifen and then DDC. Data are normalized to livers of mice not treated with DDC (n = 4). Data are mean + s.d. of the indicated number of mice. This experiment was independently repeated three times with similar results, analysing, in total, at least 10 mice for each genotype. h, Representative E-cadherin staining showing that CCA lesions retain an epithelial morphology in sections of the liver of the indicated genotype. Scale bar, 30 μm. Experiments were independently repeated on three DDC-treated Arid1a LKO mice, with similar results. P values were determined by one-way ANOVA with Dunnett’s multiple comparisons test (a) or with Tukey’s multiple comparisons test (e, g).
Extended Data Fig. 6 Interaction of SWI/SNF with F-actin and YAP is mutually exclusive.
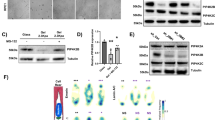
a, Related to Fig. 3a. HEK293T cells were transfected with Flag–NLS–β-actin(R62D). Representative anti-Flag immunofluorescence images to visualize transfected Flag–NLS–β-actin. Nuclei were counterstained with DAPI. Scale bar, 10 μm. b, c, Related to the PLAs shown in Fig. 3b. b, Negative controls for the PLA of Fig. 3b: in the absence of one of the two partners, no dots can be seen. c, In HEK293T cells, by PLA, endogenous BRM interacts with Flag-tagged NLS–β-actin(WT), but not with Flag-tagged NLS–β-actin(R62D), indicating that the association is specific to filamentous, and not monomeric, β-actin. d, Western blots of the inputs of the experiment shown in Fig. 3c. e, Sequential salt extraction of HEK293T cells treated with either phalloidin (Phall) or latrunculin A (Lat.A). Western blots of the indicated proteins are shown. H3 was loaded on a different blot. f, Western blots of the inputs of the experiment shown in Fig. 3d. MCF10AT cells were transfected with control siRNAs (siCo., lanes 1 and 2) or siRNAs against ARID1A (si1A; lane 3) and treated with phalloidin (lane 1) or latrunculin A (lanes 2 and 3), as indicated. g, Continuation of Fig. 3e. A PLA was carried out to detect the interaction between endogenous BRM and NLS–YAP in MCF10A cells. Control untreated cells, 0% PLA-positive cells; cells treated with the Src inhibitor dasatinib (that is, a low-mechanics condition in addition to those shown in Figs. 3e), 14.5% PLA-positive cells. h, Co-immunoprecipitation and western blot analysis of MCF10AT lysates showing endogenous ARID1A bound to endogenous YAP but not to TEAD1 and TEAD4. As a specificity control, immunoprecipitation with unrelated rabbit IgG was repeated using the same lysates. i, j, Related to Fig. 3f. i, Representative PLA images detecting the interaction between endogenous TEAD and NLS–YAP in MCF10A cells. The YAP–TEAD1 association is lost in C3-treated cells (that is, in cells with attenuated mechanotransduction (low mechanics) upon C3-mediated inhibition of RhoGTPases), but rescued after depletion of ARID1A (PLA-positive cells: 43.4%). j, Specificity controls of single antibodies for the PLA shown in i and in Fig. 3f. a–c, e, g–j are representative experiments, repeated independently two (e, h) or three (a–c, g, i, j) times, with similar results.
Extended Data Fig. 7 Loss of SWI/SNF restores YAP/TAZ transcriptional activity in mechanically inhibited cells.
a, Representative confocal images (left) and quantification (right; >100 cells per conditions) of YAP/TAZ localization in MCF10A cells transfected with the indicated siRNAs and replated on a soft ECM. b, MCF10A cells were transfected with the indicated siRNAs, and left untreated (control) or treated with anti-integrin-β1 antibodies, the Rho-inhibitors C3 and cerivastatin, the Src-inhibitor dasatinib or the ROCK inhibitor fasudil. qPCR analyses of CTGF expression (mean + s.d. of n = 3 biologically independent samples). Anti-integrin-β1 and fasudil were part of the same experiment and thus share the same control repeated in their corresponding graphs. c, HaCaT cells were transfected with the indicated siRNAs and replated to obtain either sparse (high mechanics) or dense monolayers (low mechanics). qPCR analyses of CTGF expression. Data are mean + s.d. of n = 3 biologically independent samples. d, Efficiency of Arid1a downregulation in Arid1afl/fl fibroblasts after transduction with Adeno-Cre, measured by qPCR (data are normalized to adeno-GFP-transduced cells and presented as mean + s.d. of n = 3 biologically independent samples) and western blot (in which GAPDH was used as a loading control). e, MCF10A cells were transfected with the indicated siRNAs and replated at very high density (see Methods). qPCR analyses of CTGF expression. Data are mean + s.d. of n = 3 biologically independent samples. All panels display representative experiments, repeated independently three times with similar results. P values were determined by unpaired two-sided Student’s t-test.
Extended Data Fig. 8 Loss of SWI/SNF enables YAP-induced biological effects in mechanically inhibited cells.
a, MCF10A cells were transfected with the indicated siRNAs, and replated to obtain dense monolayers (low mechanics). After 24 h, cells were incubated for 1 h with a pulse of EdU to label cells undergoing DNA duplication. Cells were fixed and processed for EdU staining. Quantification of proliferation was measured as the relative number of EdU+ cells. Data are normalized to sparse cells (high mechanics) transfected with control siRNA. Data are mean + s.e.m. of at least n = 3 biologically independent samples. Statistics for rescue experiments at low mechanics: control siRNA (n = 3) versus siBRM/BRG1 mix A (n = 3), P = 0.0003; control siRNA versus siBRM/BRG1 mix B (n = 3), P = 0.0005; control siRNA versus siARID1A#1 (n = 3), P = 0.04; control siRNA versus siARID1A#2 (n = 3), P = 0.002. A representative experiment is shown, experiments were repeated independently twice with similar results. b, Neurons were plated on a stiff or soft ECM and infected with inducible YAP-encoding vectors. Quantification of neurospheres emerging from these cultures after doxycycline treatment in NSC medium. Data are mean + s.e.m. of all biological independent samples of three experiments, n = 9. c, d, Related to Fig. 4d. Neurons were plated on a soft ECM and infected with inducible YAP-encoding vectors or empty vector and the indicated shRNA-encoding lentiviral vectors. c, d, Representative images (c) and quantification (d) of neurospheres (P0) emerging after doxycycline treatment. Scale bar, 300 μm. Data are mean + s.e.m. of four independent experiments. *P= 0.03, control shRNA (shCo) versus Brm shRNA (shBrm#1 or shBrm#2) in neurons transduced with YAP(WT); P = 0.03, control shRNA versus Arid1a shRNA (shArid1a#1 or shArid1a#2) in neurons transduced with YAP(WT). e, Fold change in expression in neurospheres emerging from cultures of YAP-induced neurons transduced with the indicated shRNAs against Brm or Arid1a, and plated either on a stiff (high mechanics) or soft (low mechanics) ECM, with respect to the corresponding control shRNA-expressing cultures. Data are mean + s.e.m. of four independent experiments. *P = 0.03, for comparisons between Brm or Arid1a shRNA under high mechanical conditions and the corresponding samples under low mechanical conditions. P values were determined by unpaired two-sided Student’s t-test (a) and by two-sided Mann–Whitney U-test (b, d, e).
Supplementary information
Supplementary Figures
This file contains uncropped blots with MW markers.
Supplementary Table 1
This file contains Supplementary Table 1.
Supplementary Table 2
This file contains Supplementary Table 2.
Supplementary Table 3
This file contains Supplementary Table 3.
Rights and permissions
About this article
Cite this article
Chang, L., Azzolin, L., Di Biagio, D. et al. The SWI/SNF complex is a mechanoregulated inhibitor of YAP and TAZ. Nature 563, 265–269 (2018). https://doi.org/10.1038/s41586-018-0658-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0658-1
Keywords
This article is cited by
-
The role of SWI/SNF complexes in digestive system neoplasms
Medical Oncology (2024)
-
The role of mechano-regulated YAP/TAZ in erectile dysfunction
Nature Communications (2023)
-
Cardiomyocyte proliferation is suppressed by ARID1A-mediated YAP inhibition during cardiac maturation
Nature Communications (2023)
-
Control of stem cell renewal and fate by YAP and TAZ
Nature Reviews Molecular Cell Biology (2023)
-
AXL activates YAP through the EGFR–LATS1/2 axis and confers resistance to EGFR-targeted drugs in head and neck squamous cell carcinoma
Oncogene (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.