Abstract
Nucleosomes cover most of the genome and are thought to be displaced by transcription factors in regions that direct gene expression. However, the modes of interaction between transcription factors and nucleosomal DNA remain largely unknown. Here we systematically explore interactions between the nucleosome and 220 transcription factors representing diverse structural families. Consistent with earlier observations, we find that the majority of the studied transcription factors have less access to nucleosomal DNA than to free DNA. The motifs recovered from transcription factors bound to nucleosomal and free DNA are generally similar. However, steric hindrance and scaffolding by the nucleosome result in specific positioning and orientation of the motifs. Many transcription factors preferentially bind close to the end of nucleosomal DNA, or to periodic positions on the solvent-exposed side of the DNA. In addition, several transcription factors usually bind to nucleosomal DNA in a particular orientation. Some transcription factors specifically interact with DNA located at the dyad position at which only one DNA gyre is wound, whereas other transcription factors prefer sites spanning two DNA gyres and bind specifically to each of them. Our work reveals notable differences in the binding of transcription factors to free and nucleosomal DNA, and uncovers a diverse interaction landscape between transcription factors and the nucleosome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Andrews, A. J. & Luger, K. Nucleosome structure(s) and stability: variations on a theme. Annu. Rev. Biophys. 40, 99–117 (2011).
Segal, E. & Widom, J. What controls nucleosome positions? Trends Genet. 25, 335–343 (2009).
Richmond, T. J. & Davey, C. A. The structure of DNA in the nucleosome core. Nature 423, 145–150 (2003).
McGinty, R. K. & Tan, S. Nucleosome structure and function. Chem. Rev. 115, 2255–2273 (2015).
Jin, J. et al. Synergistic action of RNA polymerases in overcoming the nucleosomal barrier. Nat. Struct. Mol. Biol. 17, 745–752 (2010).
Raveh-Sadka, T. et al. Manipulating nucleosome disfavoring sequences allows fine-tune regulation of gene expression in yeast. Nat. Genet. 44, 743–750 (2012).
Teves, S. S., Weber, C. M. & Henikoff, S. Transcribing through the nucleosome. Trends Biochem. Sci. 39, 577–586 (2014).
Hartzog, G. A. Transcription elongation by RNA polymerase II. Curr. Opin. Genet. Dev. 13, 119–126 (2003).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Zaret, K. S. & Mango, S. E. Pioneer transcription factors, chromatin dynamics, and cell fate control. Curr. Opin. Genet. Dev. 37, 76–81 (2016).
Segal, E., Raveh-Sadka, T., Schroeder, M., Unnerstall, U. & Gaul, U. Predicting expression patterns from regulatory sequence in Drosophila segmentation. Nature 451, 535–540 (2008).
Mirny, L. A. Nucleosome-mediated cooperativity between transcription factors. Proc. Natl Acad. Sci. USA 107, 22534–22539 (2010).
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Roy, S. et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Stanojevic, D., Small, S. & Levine, M. Regulation of a segmentation stripe by overlapping activators and repressors in the Drosophila embryo. Science 254, 1385–1387 (1991).
Yan, J. et al. Transcription factor binding in human cells occurs in dense clusters formed around cohesin anchor sites. Cell 154, 801–813 (2013).
Yin, Y. et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
Jolma, A. et al. DNA-dependent formation of transcription factor pairs alters their binding specificity. Nature 527, 384–388 (2015).
Vaquerizas, J. M., Kummerfeld, S. K., Teichmann, S. A. & Luscombe, N. M. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 10, 252–263 (2009).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Nodelman, I. M. et al. Interdomain communication of the Chd1 chromatin remodeler across the DNA gyres of the nucleosome. Mol. Cell 65, 447–459.e6 (2017).
Edayathumangalam, R. S., Weyermann, P., Gottesfeld, J. M., Dervan, P. B. & Luger, K. Molecular recognition of the nucleosomal “supergroove”. Proc. Natl Acad. Sci. USA 101, 6864–6869 (2004).
Faial, T. et al. Brachyury and SMAD signalling collaboratively orchestrate distinct mesoderm and endoderm gene regulatory networks in differentiating human embryonic stem cells. Development 142, 2121–2135 (2015).
Lolas, M., Valenzuela, P. D. T., Tjian, R. & Liu, Z. Charting Brachyury-mediated developmental pathways during early mouse embryogenesis. Proc. Natl Acad. Sci. USA 111, 4478–4483 (2014).
Kundaje, A. et al. Ubiquitous heterogeneity and asymmetry of the chromatin environment at regulatory elements. Genome Res. 22, 1735–1747 (2012).
Sherwood, R. I. et al. Discovery of directional and nondirectional pioneer transcription factors by modeling DNase profile magnitude and shape. Nat. Biotechnol. 32, 171–178 (2014).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997).
Isaac, R. S. et al. Nucleosome breathing and remodeling constrain CRISPR-Cas9 function. eLife 5, e13450 (2016).
Poirier, M. G., Bussiek, M., Langowski, J. & Widom, J. Spontaneous access to DNA target sites in folded chromatin fibers. J. Mol. Biol. 379, 772–786 (2008).
Chaney, B. A., Clark-Baldwin, K., Dave, V., Ma, J. & Rance, M. Solution structure of the K50 class homeodomain PITX2 bound to DNA and implications for mutations that cause Rieger syndrome. Biochemistry 44, 7497–7511 (2005).
Stirnimann, C. U., Ptchelkine, D., Grimm, C. & Müller, C. W. Structural basis of TBX5-DNA recognition: the T-box domain in its DNA-bound and -unbound form. J. Mol. Biol. 400, 71–81 (2010).
Coll, M., Seidman, J. G. & Müller, C. W. Structure of the DNA-bound T-box domain of human TBX3, a transcription factor responsible for ulnar-mammary syndrome. Structure 10, 343–356 (2002).
Cui, F. & Zhurkin, V. B. Rotational positioning of nucleosomes facilitates selective binding of p53 to response elements associated with cell cycle arrest. Nucleic Acids Res. 42, 836–847 (2014).
Li, Q. & Wrange, O. Accessibility of a glucocorticoid response element in a nucleosome depends on its rotational positioning. Mol. Cell. Biol. 15, 4375–4384 (1995).
McGinty, R. K. & Tan, S. Recognition of the nucleosome by chromatin factors and enzymes. Curr. Opin. Struct. Biol. 37, 54–61 (2016).
Zhou, B. R. et al. Structural mechanisms of nucleosome recognition by linker histones. Mol. Cell 59, 628–638 (2015).
Iwafuchi-Doi, M. et al. The pioneer transcription factor FoxA maintains an accessible nucleosome configuration at enhancers for tissue-specific gene activation. Mol. Cell 62, 79–91 (2016).
Struhl, K. & Segal, E. Determinants of nucleosome positioning. Nat. Struct. Mol. Biol. 20, 267–273 (2013).
Collings, C. K., Fernandez, A. G., Pitschka, C. G., Hawkins, T. B. & Anderson, J. N. Oligonucleotide sequence motifs as nucleosome positioning signals. PLoS ONE 5, e10933 (2010).
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998).
Ramachandran, S. & Henikoff, S. Transcriptional regulators compete with nucleosomes post-replication. Cell 165, 580–592 (2016).
Li, M. et al. Dynamic regulation of transcription factors by nucleosome remodeling. eLife 4, e06249 (2015).
Sekiya, T., Muthurajan, U. M., Luger, K., Tulin, A. V. & Zaret, K. S. Nucleosome-binding affinity as a primary determinant of the nuclear mobility of the pioneer transcription factor FoxA. Genes Dev. 23, 804–809 (2009).
Hayes, J. J. & Wolffe, A. P. Histones H2A/H2B inhibit the interaction of transcription factor IIIA with the Xenopus borealis somatic 5S RNA gene in a nucleosome. Proc. Natl Acad. Sci. USA 89, 1229–1233 (1992).
Jolma, A. et al. Multiplexed massively parallel SELEX for characterization of human transcription factor binding specificities. Genome Res. 20, 861–873 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Henikoff, J. G., Belsky, J. A., Krassovsky, K., MacAlpine, D. M. & Henikoff, S. Epigenome characterization at single base-pair resolution. Proc. Natl Acad. Sci. USA 108, 18318–18323 (2011).
Kasinathan, S., Orsi, G. A., Zentner, G. E., Ahmad, K. & Henikoff, S. High-resolution mapping of transcription factor binding sites on native chromatin. Nat. Methods 11, 203–209 (2014).
Chiu, T. P. et al. DNAshapeR: an R/Bioconductor package for DNA shape prediction and feature encoding. Bioinformatics 32, 1211–1213 (2016).
Chiu, T. P., Rao, S., Mann, R. S., Honig, B. & Rohs, R. Genome-wide prediction of minor-groove electrostatic potential enables biophysical modeling of protein-DNA binding. Nucleic Acids Res. 45, 12565–12576 (2017).
Polach, K. J. & Widom, J. Mechanism of protein access to specific DNA sequences in chromatin: a dynamic equilibrium model for gene regulation. J. Mol. Biol. 254, 130–149 (1995).
Anderson, J. D. & Widom, J. Sequence and position-dependence of the equilibrium accessibility of nucleosomal DNA target sites. J. Mol. Biol. 296, 979–987 (2000).
Li, G., Levitus, M., Bustamante, C. & Widom, J. Rapid spontaneous accessibility of nucleosomal DNA. Nat. Struct. Mol. Biol. 12, 46–53 (2005).
Privalov, P. L., Dragan, A. I. & Crane-Robinson, C. The cost of DNA bending. Trends Biochem. Sci. 34, 464–470 (2009).
Ye, Z. et al. Genome-wide analysis reveals positional-nucleosome-oriented binding pattern of pioneer factor FOXA1. Nucleic Acids Res. 44, 7540–7554 (2016).
Cirillo, L. A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol. Cell 9, 279–289 (2002).
Acknowledgements
We thank F. Zhong, A. Jolma, J. Zhang and J. Toivonen for valuable suggestions; E. Inns for proofreading; T. Kivioja for critical review of the manuscript and L. Hu, J. Liu and S. Augsten for technical assistance. This work was funded by the EU Horizon 2020 project MRGGrammar (664918), Cancerfonden (120529, 150662), Knut and Alice Wallenberg Foundation (2013.0088), Vetenskapsrådet (D0815201), Academy of Finland CoE (312042) (J.T.); DFG (SFB860, SPP1935), ERC AdG TRANSREGULON (693023), Volkswagen Foundation (P.C.) and EMBO fellowship ALTF 949-2016 (S.D.).
Reviewer information
Nature thanks T. Hughes, B. F. Pugh and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
J.T., F.Z. and P.C. conceived the experiments. F.Z. performed most experiments and analyses. L.F. produced the histone octamers. B.S. and E.K. contributed to generation and analysis of the MNase–seq and ChIP–seq data, respectively. B.W. and S.O.D. performed SOX EMSA and the binding assay with nuclear proteins, respectively. Y.Y. contributed to protein production and motif analysis. M.T., K.R.N. and E.M. contributed to design and analysis of sequencing and structure data. F.Z. and J.T. interpreted the data and wrote the manuscript. All authors discussed the findings and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
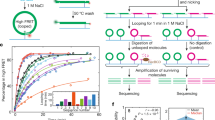
Extended Data Fig. 1 Experiment design and data analysis strategy of NCAP–SELEX.
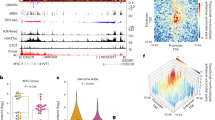
a, Expression of the recombinant histones from Xenopus laevis. For each lane 3 μg histone is loaded. Similar purifications for untagged H2A, H2B, H3, and H4 have been repeated at least three times. The SBP–H2A purification was performed once. b, Size-exclusion chromatogram of the histone octamer. Octamer formation was performed twice and the results were highly consistent. c, EMSA result showing the reconstituted nucleosomes using lig147 and lig200. The original ligands are also loaded as reference. The asterisks indicate the nucleosome bands. Similar results are seen in four independent nucleosome reconstitutions. For gel source data see Supplementary Fig. 1. d, Oligonucleotide periodicity in the library enriched by nucleosome. As a quality control of nucleosome reconstitution, we verified whether nucleosome by itself is enriching the previously reported approximately 10-bp periodic oligonucleotide signal41,42. Nucleosome SELEX (without TF) were carried out for four cycles to enrich nucleosome-favouring ligands. The counts of each single and di-nucleotide across each individual ligand were Fourier transformed and summed up for the whole library. A clear peak around 0.1 bp−1 (corresponding to the reported approximately 10-bp periodicity) is visible for most mono- and dinucleotides. e, The C/G/CG preferences of nucleosome. All 9-mers were counted for the nucleosome-favoured (bound) and the nucleosome-disfavoured (unbound) libraries. The point representing each 9-mer is coloured according to its C/G/CG content (top), and the count ratios between the bound and the unbound libraries are summarized for 9-mers of different C/G/CG contents (bottom). For the box plots grouped by C/G content, the sample sizes of the boxes are 19,683, 59,049, 78,732, 61,236, 30,618, 10,206, 2,268, 324, 27 and 1, respectively for 9-mer groups containing 0 to 9 C/G. For the box plots grouped by CG dinucleotide content, the sample sizes of the boxes are 151,316, 91,824, 17,784, 1,200 and 20, respectively, for 9-mer groups containing 0 to 4 CG. The line within each box represents the median; the lower and upper boundaries of the box indicate the first and third quartiles and the whiskers represent the 1.5-fold interquartile range. More extreme values are indicated with dots. f, Analysis pipeline for the ligands enriched in NCAP–SELEX. g, E-MI strength comparison for libraries with and without TF signals. The E-MI heat maps represent signals in the input (cycle 0) library, in the cycle 4 library of nucleosome-favoured sequences (Nucl. SELEX), and in the NCAP– and high-throughput (HT)-SELEX cycle 4 libraries. The libraries enriched with TF (NCAP and HT) have much stronger E-MI signals compared to the cycle 0 and the nucleosome-SELEX library. The detected dimer signals of HSF1 in HT-SELEX is boxed. h, Family-wise coverage of TFs tried in NCAP–SELEX.
Extended Data Fig. 2 NCAP–SELEX with lig200.
a, Hierarchical clustering of the E-MI diagonals for NCAP–SELEX with the 200-bp ligand (lig200). The E-MI diagonal for each TF is oriented radially. The randomized region is 154 bp and contains 149 windows for MI calculation between neighbouring 3-mers. The names of the TFs are coloured by family with the colouring scheme indicated on the centre. TFs from the same family tend to be clustered together (for example, SOX, indicated). Because of the gradient of nucleosome occupancy, the penetration of the E-MI signal into the centre of the E-MI diagonals (E-MI penetration; see Supplementary Methods for details) reflects the ability of each TF to bind to nucleosomal DNA. Note that almost all TFs have lower E-MI towards the centre of lig200, indicating their lower affinity to nucleosomal DNA than to free DNA. The decrease of E-MI towards the centre is rarely observed in the absence of the nucleosome. Note that the binding inhibition of TF to nucleosomal DNA occurs in the absence of higher-order effects, such as chromatin compaction, remodelling or histone modification. This result directly verifies the mutually antagonistic role of TFs and the nucleosome13,43,44, which has been biochemically validated in only a few cases45,46. The E-MI diagonals shown are scaled for each TF (see Supplementary Methods). Owing to the fixed adaptor sequences, TFs may prefer one end of the lig200 over the other end. b, E-MI penetration of individual TFs on lig200. TFs are ordered according to their E-MI penetration depth towards the centre of the ligand. This order reflects the ability of TFs to bind nucleosome-occupied DNA. Note that the penetration of E-MI into the ligand centre (E-MI penetration; see Supplementary Methods for details) varies strongly between the TFs. TFs representing either of the two ends are coloured red and exemplified in c. c, The diagonal of E-MI for TFs with high (above dotted line) and low (below dotted line) E-MI penetrations. Because HT (blue) and NCAP–SELEX (black) may differ in stringency, each E-MI diagonal is normalized by dividing its maximum value. On lig200 the central 94 bp (shaded grey) is always occupied by a nucleosome. d, Correlation between E-MI penetration and the capability of TFs to bind nucleosomal DNA in vivo. Per base-pair coverage of MNase fragments (>140 bp) at ChIP–seq peaks of the TFs (x axis) is plotted against their E-MI penetration (y axis) in NCAP–SELEX. The calculation of Pearson’s r and the correlation test is performed for n = 20 TFs. The observed correlation suggests that the ability of TFs to bind nucleosomal DNA in NCAP–SELEX (E-MI penetration) partially explains the nucleosome occupancy at the sites of TFs in K562 cells. Thus the biochemical ability of TFs to bind to nucleosomal DNA also affects their binding in vivo. e, Left, E-MI heat map of T (brachyury) in HT-SELEX using lig200. Pairwise E-MI for all 3-mer pairs is presented as a heat map. The signal is only visible near the diagonal, no E-MI signal is detected across approximately 80 bp. Right, the gyre-spanning mode (arrow) of T (brachyury) on lig147. The corresponding motif is derived with the indicated seed for a specific position (number in the parentheses) in the high E-MI region (arrow). Position weight matrix generation follows our previous method47 using multinomial 1. f, Type 2 binding of Brachyury (T) stabilizes nucleosome from dissociation. log2 ratio of E-MI between the bound and unbound libraries (cycle 5) is calculated for both the type 2 binding and for the background E-MI level (see Supplementary Methods for details) of Brachyury (T). Compared to the unbound, the bound library has stronger type 2 binding but a similar background. As a control, for 20 random TFs (Rnd), the log2 ratio of E-MI between the bound and unbound libraries is also calculated for both the type 2 binding (hypothetic) and for the background E-MI level. For these TFs the bound libraries have similar E-MI strength as the unbound in the region corresponding to the type 2 binding of Brachyury (T). Data are mean ± s.d.; two-sided t-test was used, 95% confidence intervals, 0.097 − 0.202 (T) and −0.008 − 0.004 (random TFs). The sample sizes are n = 20 libraries for random TFs and n = 4 independent SELEX replicates for Brachyury (T). The raw data for the random control TFs are listed in Supplementary Data 3. g, E-MI heat map of TBX2, ETV4 and ETV1 in NCAP–SELEX using lig147. The E-MI signals across approximately 80 (type 2) or 40 bp (type 1) are indicated with arrows. The corresponding motif of each binding type is derived with the indicated seed for a specific position (number in the parentheses) in the high E-MI regions (arrows). Note that the E-MI signals across approximately 40 bp are position-specific, with one binding event being observed near the dyad, and the other(s) on the opposite side of the nucleosome, with the two contacts separated by approximately 180°. This binding mode can be achieved by TF dimers that contact nucleosomal DNA in a pincer-like manner. However, as the individual TFs are located far from each other in this binding mode, it probably suggests that the nucleosome may have two allosteric states, or may form a higher-order complex with these TFs.
Extended Data Fig. 3 Control experiments with lig200.
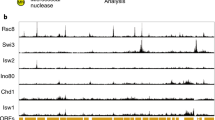
a, Determination of nucleosome positions for NCAP–SELEX libraries (lig200, all TFs). To examine if nucleosome has preferred positioning on lig200, nucleosomes were loaded onto the amplified cycle 4 NCAP–SELEX library of each TF. After digestion with MNase, the remaining DNA fragments were collected and sequenced. A titration was first carried out to find the appropriate concentration of MNase. As shown in the gel image (left, see Supplementary Fig. 1 for gel source image), 4.8, 2.4, 1.2, 0.6, 0.3, 0.15 U of MNase (lane 1–6) were added into each 25-μl reaction containing the purified nucleosome. According to the results, the condition marked by an asterisk was chosen for the reactions to determine nucleosome position. After sequencing, the fractions of MNase fragments that mapped to the variable region (grey) and to the adaptor-overlapping region (blue) of lig200 are visualized (middle, each row corresponds to a TF). To identify potential positional preference of nucleosome on lig200, the adaptor-overlapping fragments are analysed for their end distributions. Distributions of both the left end (cyan) and the right end (red) of the MNase-digested fragments on lig200 are shown (right, each row corresponds to a TF). Such distributions likely indicate that nucleosomes have two relatively preferred positions on lig200 (illustrated by cartoon in green). Note that most nucleosomes are not positioned by the adaptor (middle) thus are randomly distributed. b, E-MI diagonals for HT-SELEX with the 200-bp ligand (lig200). TFs are arranged according to the clustering for NCAP-SELEX libraries (Extended Data Fig. 2a) to facilitate comparison. TFs without a lig200 HT-SELEX control are left as blank. The E-MI diagonal for each TF is oriented radially and the names of the TFs are coloured by family as indicated. The E-MI diagonals are scaled for each TF. Some TFs show preferred positions on lig200, probably due to the fixed adaptors. c, TFs prefer free DNA to the edge of a nucleosome. For a few randomly chosen TFs, NCAP–SELEX was run using a ligand (Lig70Nlinker, sequence in Supplementary Table 2) that positions nucleosome at its centre by embedding a segment of Widom 601 sequence, and with randomized flankings. At a low resolution, the E-MI signal of TFs decreases monotonically towards the nucleosome-occupied region. Thus the higher E-MI at the flankings of lig200 (Extended Data Fig. 2a) suggests the preference of TFs for free DNA, rather than for the edge of a nucleosome. E-MI diagonals are scaled for each TF. d, E-MI diagonals for TFs at doubled concentrations. The concentration effect on the E-MI diagonal of TFs is explored by running NCAP–SELEX at doubled (2×) concentrations for a few randomly chosen TFs. Compared to the E-MI diagonal with the original TF concentrations (1×), the change in the E-MI pattern is minor.
Extended Data Fig. 4 Nucleosome breaks the rotational symmetry of DNA.
a, Density plot representing the orientational asymmetry of all TFs in NCAP–SELEX and in HT-SELEX. In NCAP–SELEX, more TFs bind with high orientational asymmetry than in HT-SELEX. A few TFs can also prefer different ends of the ligand for the two binding directions in HT-SELEX; this is likely induced by the adaptor sequences. However, there are more TFs with higher orientational asymmetry in NCAP–SELEX libraries, despite the fact that for most TFs their signals are stronger in HT-SELEX libraries. b, Orientation asymmetry of ELF2 revealed by using top 8-mers. Each row of the heat map corresponds to the counts distribution of a top 8-mer (non-palindromic) across the positions of the SELEX ligand. Hits of the top 8-mers occur at different ends for different strands of nucleosomal DNA (that is, an 8-mer and its reverse-complement prefer different ends), whereas their distribution is relatively homogeneous for free DNA. c, Orientation asymmetry of CREB TFs. CREB TFs have different motif density distributions for the two strands of nucleosomal DNA. The motif used for matching is indicated above. The minus strand profile is from the density of the reverse-complement motif. d, Break of the two-fold rotational symmetry of DNA induces preferred orientation of TFs. Left, free DNA has a pseudo-two-fold axis (red ellipse) perpendicular to the helix axis. Motifs in two orientations are symmetric with each other with respect to a 180° rotation centred on the axis. Right, for motifs on nucleosomal DNA, if the other strand of DNA or the histone proteins (green) affect binding, the two-fold axis of DNA no longer exists, as a 180° rotation centred on the axis no longer generates an identical conformation (the rotated image not superimposable with the original one). The break of rotational symmetry occurs also on the linker DNA that immediately flanks the nucleosome (f). e, Top, the orientational asymmetry of ELF1 in NCAP–SELEX of lig200. Bottom, the asymmetric nucleosome distribution around genomic ELF1 sites (top). Such asymmetry is not observed for the same ELF1 sites after a 30 min 500 mM KCl treatment to mobilize the nucleosome (bottom). ELF1 motif matches are positioned at the centre. Frequency of the centre of MNase-fragments (140–170 bp) is visualized for nearby regions to represent the nucleosome occupancy. Each profile (n = 999 data points) is LOESS smoothed with a span of 0.05 and the shaded band indicates the s.e.m. f, The orientational binding of ELF occurs on both the nucleosomal DNA and the nearby linker region. The motif matches of ELF on lig147 (top) suggest that the orientational binding occurs on nucleosomal DNA. In addition, the motif matches of ELF on the 293-bp ligand (bottom; nucleosome positioned at the centre, ligand schematic in Extended Data Fig. 3c) indicates that the orientational binding also occurs on nearby linker DNA regions.
Extended Data Fig. 5 TFs can bind nucleosomal DNA without substantial motif change.
a, Cartoons showing that TFs are theoretically able to contact grooves of the bent nucleosomal DNA from the solvent-exposed side. The left panel for each TF shows the structures (Protein Data Bank (PDB); PITX2: 2LKX, TBX5: 2X6V). For the right panels of each TF, the PDB structure of the TF is aligned to the nucleosome structure (3UT9) as described in the Supplementary Methods (section ‘FFT analysis and structure alignment’). The corresponding base pairs of the nucleosomal DNA were replaced with Coot48 according to the DNA sequence in the PDB structure of each TF. The models are visualized with UCSF Chimera49. b, TFAP binds nucleosomal DNA with slightly different specificity than free DNA. The scatter plot (top) shows the counts of gapped 9-mers from SELEX libraries of TFAP2B, enriched with NCAP–SELEX (x axis) and HT-SELEX (y axis). The examined 9-mers consist of three segments of trimers interspaced with two gaps (0–5 bp). Only the most enriched 9-mers (top 300 in each library and in the combined library) are shown for clarity. For comparison, the most differentially enriched gapped 9-mers were also used as seeds to derive the corresponding motifs from both libraries (right). The heat map (bottom) shows the pairwise E-MI for all combinations of positions on lig147, in the presence (left) and absence (right) of nucleosome. The arrowheads indicate the additional signals developed in the presence of nucleosome.
Extended Data Fig. 6 NCAP–SELEX with lig147.
a, E-MI diagonals for HT-SELEX with the 147-bp ligand (lig147). TFs are arranged according to the clustering for NCAP–SELEX libraries (Fig. 3a) to facilitate comparison. The E-MI diagonal for each TF is oriented radially and scaled. The names of the TFs are coloured by family as indicated. b, The top five principal components (PCs) and the components from non-negative matrix factorization (NMF) with rank equal to five. The E-MI diagonals of lig147 (n = 195 TFs) were used in the dimension reduction. For visualization purposes, each component is centred and scaled. Note that the five principal components (left) correspond well to the three identified positional preferences of TFs on nucleosomal DNA (end: dim 1, 2; periodic: dim 3, 4; dyad: dim 5). c, Comparison between the scores from principal-component classifiers and custom classifiers. Red points indicate the TFs defined as displaying respective preferences according to custom classifiers. The PC classifiers are well in accordance with custom classifiers for the end and the dyad preferences (left), but not for the periodic preference (right). Because the phase of periodic preference can vary continuously whereas principal components can only capture discrete values, the custom FFT-based classifier is more natural for such purposes. The libraries of n = 195 TFs were used in the analyses. The correlation coefficients (Pearson’s r) are also indicated. d, E-MI diagonal and motif-matching results for the bZIP factor CEBPB. In HT-SELEX (without nucleosome), the binding signal is more distributed across the ligand. e, Pearson’s correlation between the E-MI penetrations of TFs on lig200 and on lig147. The libraries of n = 155 TFs, which are successful with both lig200 and lig147, were used in this analysis. The end preference of TFs on lig200 reveals that they prefer free DNA to nucleosomal DNA. The free-DNA preference also probably explains the end preference of TFs on lig147 owing to the observed correlation of E-MI penetrations. For each TF, the E-MI penetration values differ between lig147 and lig200 because free-DNA regions are expected near the ends of lig200, but not present on lig147. f, Correspondence between the E-MI patterns of TFs on lig147 and on 1ig200. The E-MI diagonals of RFX5 and SHOX on lig200 and those on lig147 are plotted together for comparison. The peaks on lig200 that illustrate the central preference of RFX5 and periodic preference of SHOX are indicated with red arrowheads. The weaker preference patterns on lig200 are due to the delocalization of the nucleosome on lig200, however they are still visible because the two fixed adaptors dictate two weakly preferred nucleosome positions.
Extended Data Fig. 7 TFs with periodic preferences.
a, Density plot showing the periodicity strength of all TFs in NCAP–SELEX (orange) and HT-SELEX (blue). Note that the overall periodicity of E-MI is stronger for the NCAP–SELEX library compared to the free-DNA HT-SELEX library. b, A minor-groove binder prefers exposed minor grooves (m) on nucleosomal DNA. The E-MI diagonal of EOMES (T-box) is out of phase with the TA dinucleotide peaks, suggesting that it binds positions where the minor groove of nucleosomal DNA is facing outside (TA peaks indicate nucleosome–DNA contacts, whereas E-MI visualizes TF–DNA contacts, see Supplementary Methods for details). Accordingly, the TBX5 (T-box) structure (PDB entry 2X6V) shows contacts with DNA principally in the minor groove. Cartoon representation to the right shows that the steric hindrance is minimal when TBX5 (blue) binds out of phase with TA (orange) on the nucleosome structure (PDB entry 3UT9). c, Strength and phase of the approximately 10-bp periodicity of the TA dinucleotide in NCAP–SELEX and HT-SELEX libraries. For the library (lig147) enriched by a specific TF, the strength and phase information is derived from FFT of the TA counts at each position of the library. In the polar plot, each dot represents the library of one TF. The overall periodicity is stronger in the NCAP–SELEX libraries (yellow) than in the HT-SELEX libraries (blue), suggesting an enrichment of nucleosome signal. The TA phases in the NCAP–SELEX libraries of all TFs are similar, thus the rotational positioning of nucleosome on the SELEX ligand is similar for the libraries of all TFs. By contrast, the phase of the E-MI periodicity is much more dispersed (Fig. 4a), suggesting the preference of TFs towards different grooves of DNA. d, Cartoon representations of the 3D structures of PITX2 (PDB entry 2LKX) and TBX5 (T-box, PDB entry 2X6V) in complex with nucleosomal DNA. TBX5 structures were shown to illustrate the groove preferences of EOMES (T-box). The DNA ligand in the nucleosome structure (PDB entry 3UT9) contains phased TA steps (orange). Consistent with the SELEX result, PITX is more compatible with nucleosomal DNA when it binds in phase with TA, whereas T-box is more compatible when it binds out of phase with TA. Therefore, when a TF binds nucleosomal DNA according to the identified patterns, the steric conflict between TF and the histones is minimized. e, E-MI diagonal and motif-matching results for SHOX in NCAP–SELEX and HT-SELEX. The E-MI diagonal agrees with the motif-matching result. f, The approximately 10-bp periodicity for the preferred spacing of SHOX dimers on nucleosomal DNA. In NCAP–SELEX libraries of many periodic binders (SHOX as an example), enrichment of the most abundant 3-mer tandem repeats oscillates as a function of the spacing between the repeats. The enrichment is evaluated by the log2 ratio between the observed and expected occurrences. The observed approximately 10-bp periodicity with dimer spacing originates from the periodic availability of nucleosomal DNA. However, in most cases binding appears not to be cooperative, on the basis of the fact that the observed frequency of ligands with two motifs can be well estimated by the frequency of ligands that contain only one motif (data not shown). g, Homeodomain TFs from mouse liver prefer periodic positions on nucleosomal DNA. Motif hits of homeodomain TFs show a periodic pattern for both the nucleosome-bound and nucleosome-dissociated (unbound) libraries after incubation with mouse liver nuclear extract; however, the unbound library has more motif hits, indicating that binding events to the presented motif facilitate the dissociation of nucleosome. To more clearly visualize the approximately 10-bp periodicity, the Fourier-transformed spectra for both libraries are also shown to the right. The arrowhead indicates the peaks for the approximately 10-bp periodicity.
Extended Data Fig. 8 TFs with the dyad preference.
a, E-MI diagonal and motif-matching results for RFX5. The distribution of binding events is more spread in the absence of nucleosome (HT-SELEX). b, The design of the competition assay and the raw counts of RFX5 motif matches. Differently barcoded nucleosomal DNA (orange) and free DNA (blue) were mixed as input, and incubated with the TF protein. Purification for the TF-bound species was then performed. Matches of the indicated RFX5 motif was counted for both the nucleosomal DNA (orange) and the free DNA (blue), and for both of the input and the bound libraries. On nucleosomal DNA, more motif hits near the centre of the ligand are observed after purification. c, MNase–ChIP fragments near the binding sites of RFX5 and HOXB13. Motif matches within MNase–ChIP peaks of each TF are positioned at the centre. Counts of MNase–ChIP fragments are binned to 3 bp by 3 bp bins according to their lengths and centre positions. Nucleosome distribution is reflected by the signal intensity of the approximately 150-bp fragments (bracket). This visualization resembles the reported ‘V-plot’50. Length distribution of all ChIP fragments and that of fragments <300 bp from the TF sites are shown on the right. Note that HOXB13 enriches ChIP fragments of approximately 120 bp at its sites (middle), suggesting that, similarly to most TFs50,51, its binding sites in the genome are depleted of nucleosomes. By contrast, RFX5 enriches nucleosome-sized fragments (left). Most of the enriched fragments also have their centre positioned between the red ‘V’ lines, and thus overlap with the TF motifs. d, Nucleosome distribution near the binding sites of RFX5 and HOXB13 before transfection (no TF expression). MNase–seq fragments around the identified TF sites are visualized as in c. The sites later bound by exogenous RFX5 are located at the maximum of nucleosome occupancy (left). e, Nucleosome distribution near the binding sites of RFX5 and HOXB13 after transfection (with TF expression). The nucleosomes are now positioned beside the exogenous RFX5 sites (left). f, EMSA of SOX11 complexes with nucleosome and with free DNA. Nucleosome is reconstituted and purified using a modified Widom 601 sequence, which contains a SOX11 binding sequence (extracted from cycle 4 SELEX library) embedded close to the dyad. Each 40 μl reaction contains 1 μg DNA, together with SOX11 protein at a molar ratio of 0, 0.5, 1, 2 (indicated at the top of each lane) to DNA. Here the observed multiple shifts probably reveal the binding of SOX11 to additional weaker sites on the ligand (shown in g). For gel source data, see Supplementary Fig. 1. g, The score of SOX11 motif across the EMSA ligand (see Supplementary Methods for ligand sequence). The top three binding sites are indicated. h, DNA shape features around SOX11 motifs. DNA shape features were calculated using DNAshapeR52,53, for NCAP–SELEX (black), HT-SELEX (blue), and cycle 0 (input, grey) libraries. The black line is plotted last thus may hide other lines when all values are similar. The boundary of each motif is indicated with dashed vertical lines. Only the ligands with motifs around the centre (position range: 36–58) are included in the analysis.
Extended Data Fig. 9 TF binding affects the stability of nucleosome.
a, E-MI difference between the bound and the unbound cycle 5 libraries. The bound and the unbound libraries were collected either in the presence (left) or in the absence (right) of TFs. The heat maps visualize E-MI differences between the bound and unbound libraries for all position combinations of 3-mer pairs, and each pixel on the heat map is a mean of the E-MI difference of all the examined TFs at this pixel. For individual TFs, the value at each pixel is calculated as log2(E-MIunbound/E-MIbound). Testing nucleosome dissociation in the absence of the TF aimed to verify whether the TF motifs on lig147 by themselves can affect the stability of the nucleosome. Note that in general, binding events close to the centre of nucleosomal DNA more efficiently dissociated the nucleosome (left). This observation is in accordance with the mutually exclusive nature between TFs and the nucleosome. Although TFs generally have lower affinity to the centre of the lig147, it is also conceivable that TF binding close to the centre will more efficiently undermine the DNA–histone interactions, and in turn lead to a higher rate of nucleosome dissociation. TFs bound close to the ends could have decreased the flexibility of the DNA there and subsequently disfavour the dissociation of DNA ends from the histones, which in turn contributes to nucleosome stability. b, The efficiency of nucleosome dissociation induced by ETV1 is dependent on its binding specificity. To displace nucleosome, binding with the shorter motif is more efficient than binding with the longer motif, because the shorter motif is more enriched in the dissociated library (unbound). c, Differential E-MI diagonals for TFs at doubled concentrations. The ability of each TF to dissociate or stabilize nucleosome is revealed by the log ratio of E-MI between the unbound and the bound cycle 5 libraries (differential E-MI). The concentration effect on the differential E-MI diagonal of TFs is explored by running NCAP–SELEX followed by the dissociation assay at doubled (2×) concentrations of the TFs. The differential E-MI diagonals at 2× TF concentrations resemble those at the original (1×) TF concentrations. d, Differential E-MI diagonals for the four ETS family TFs indicated by asterisks in Fig. 5a.
Extended Data Fig. 10 Modes of TF–nucleosome interaction.
a, For each TF, the strengths of all identified TF–nucleosome interaction modes, together with its ability to dissociate nucleosome, are shown in the heat map. The displayed features include the positional preference of each TF (E, end; P, periodic; D, dyad) on nucleosomal DNA, gyre-spanning binding mode (Gs), orientational asymmetry (Asym), and the ability of each TF to dissociate nucleosome (Ds). TFs succeeding only in NCAP–SELEX with lig200 are presented to the right for their orientational asymmetry. In the heat map values are scaled into 0 to 1 for each mode, except for the dissociation, in which TFs that stabilize nucleosome are given negative values (green). The raw data are provided in Supplementary Table 5. b, All the identified modes can be explained by the structural features of nucleosome. TFs with the end preference (E) bind nucleosomal DNA close to the entry and exit positions. This preference is in line with the probability of spontaneous dissociation (breathing) of nucleosomal DNA, which decreases from the end to the centre54,55,56. TFs with a strong end preference are likely less compatible with nucleosomal DNA thus only bind to the dissociated regions. These TFs could be structurally hindered by nucleosome, because one side of the nucleosomal DNA is masked by the histones. Moreover, nucleosomal DNA is bent sharply, which could impair TF-DNA contacts if TFs have evolved to specifically bind to free DNA. TFs with the periodic preference (P) binds approximately every 10.2 bp positions on nucleosomal DNA. This preference arises also because nucleosomal DNA is accessible only from one side, which leads to significant accessibility change along each pitch (approximately 10.2 bp) of the DNA helix. TFs that bind to short motifs, or to discontinuous motifs, are still able to occupy the available periodic positions on nucleosomal DNA. TFs with the dyad preference (D) tend to bind close to the nucleosomal dyad. Structurally, the dyad is distinct from other regions of the nucleosomal DNA. The dyad contains only a single DNA gyre, and features the thinnest histone disk29,37. These characteristics of the dyad DNA reduce the steric barrier for TF binding. The relatively weak DNA–histone interaction around the dyad could allow TFs that bend DNA upon binding (for example, SOXs57) to deform DNA more easily at the dyad compared to other positions. In addition, the entry and exit of nucleosomal DNA are also close to the dyad; together with the dyad DNA, they provide a scaffold for specific configurations of TFs. FOXA has been suggested to make use of this scaffold to achieve highly specific positioning close to the dyad39,58. However, the dyad positioning of FOXA is not observed in this study using eDBD, potentially because the full length of FOXA is required for its interaction with the nucleosome59. A few T-box TFs were found to bind nucleosomal DNA with the gyre-spanning binding mode (Gs). This mode is observed because DNA grooves align across the two nucleosomal DNA gyres29. The parallel gyres could specifically associate with TF dimers, or TFs with long recognition helices or multiple DNA-binding domains. The dual-gyre binding is possible only on nucleosomal DNA, and it thus stabilizes the nucleosome from dissociation, and may therefore function to lock a nucleosome in place at a specific position. Many TFs such as ETS and CREB show an orientational asymmetry (Asym) upon binding to the nucleosomal DNA. The nucleosomal environment has induced such preference by breaking the local rotational symmetry of DNA. In accordance with the mutually exclusive nature of TF and nucleosome binding, most TFs were found to dissociate nucleosomes (Ds). While nucleosome weakens the affinity of incompatible TFs, binding of such TFs are expected to weaken the nucleosome–DNA contacts as well. The ability of TFs to dissociate nucleosome is required for them to open chromatin and to activate transcription. Moreover, we also observed TFs that both stabilize and destabilize nucleosomal DNA, depending on their relative position of binding. Such ability could be used to more precisely position local nucleosomes. All the identified TF–nucleosome interactions suggest that the TF–nucleosome interaction could be more complicated than the previously suggested pioneer/non-pioneer classification of TFs11. We observed that for eDBD of almost all TFs, including known pioneer factors such as FOX and SOX, free DNA was nonetheless preferred over nucleosomal DNA. However, some pioneer factors can bind relatively better to the interior of the nucleosome (for example, FOX and SOX). In addition, some other TFs prefer nucleosomal DNA at restricted positions, or with one of their multiple binding motifs. These strategies are likely related to the access of pioneer factors to nucleosomal DNA.
Supplementary information
Supplementary Information
This file contains the Methods section of the manuscript.
Supplementary Figures
This file contains Supplementary Figure 1 | Uncropped gel scans with size marker indications.
Supplementary Data
This file contains Supplementary Data 1 | The binding specificities of TFs in NCAP–SELEX and HT-SELEX libraries. The phylogenetic tree was generated based on the amino acid sequences of the TF DBDs. The primary motifs were curated for lig147 NCAP–SELEX and HT-SELEX libraries with manually curated seeds (see Method for details). Motif-matching results for each TF’s cycle 4 and cycle 0 libraries were illustrated. The published motifs were from our previous curations.
Supplementary Data
This file contains Supplementary Data 2 | Motif-based analysis for NCAP–SELEX. a, Normalized E-MI diagonals from NCAP–SELEX against those from the corresponding HT-SELEX. b, Motif-matching results for NCAP–SELEX with the 147-bp ligand (lig147). c, Motif hits ratio (log scale) between the bound and the unbound cycle 5 libraries. d, Motif-matching of NCAP–SELEX with the 200-bp ligand (lig200).
Supplementary Data
This file contains Supplementary Data 3 | Control TFs without the gyre-spanning binding mode. Data showing that the randomly chosen control TFs in Extended Data Fig. 2f are without the gyre-spanning mode as identified for Brachyury (T). The control TFs are thus used for evaluating the fluctuation of the background noise between the cycle 5 bound and unbound libraries.
Supplementary Tables
This file contains Supplementary Tables S1–S8. The table legends are as follows: Table S1 - Sequence information for proteins, Table S2 - Sequence information for DNA ligands, Table S3 - Collection of E-MI diagonals, Table S4 - Competition assay for RFX5, Table S5 - Nucleosome–TF interaction modes, Table S6 - PCA and NMF analysis of TF signals on lig147, Table S7 - Collection of PFM models, Table S8 - Peaks called for MNase-ChIP.
Rights and permissions
About this article
Cite this article
Zhu, F., Farnung, L., Kaasinen, E. et al. The interaction landscape between transcription factors and the nucleosome. Nature 562, 76–81 (2018). https://doi.org/10.1038/s41586-018-0549-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0549-5
Keywords
This article is cited by
-
Integrative analysis of transcriptomic and epigenomic data reveals distinct patterns for developmental and housekeeping gene regulation
BMC Biology (2024)
-
Cell-type-directed design of synthetic enhancers
Nature (2024)
-
The BAF chromatin remodeler synergizes with RNA polymerase II and transcription factors to evict nucleosomes
Nature Genetics (2024)
-
Protein-intrinsic properties and context-dependent effects regulate pioneer factor binding and function
Nature Structural & Molecular Biology (2024)
-
Structural mechanism of synergistic targeting of the CX3CR1 nucleosome by PU.1 and C/EBPα
Nature Structural & Molecular Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.