Abstract
SARS-CoV-2, the virus responsible for the COVID-19 pandemic, has been associated with substantial global morbidity and mortality. Despite a tropism that is largely confined to the airways, COVID-19 is associated with multiorgan dysfunction and long-term cognitive pathologies. A major driver of this biology stems from the combined effects of virus-mediated interference with the host antiviral defences in infected cells and the sensing of pathogen-associated material by bystander cells. Such a dynamic results in delayed induction of type I and III interferons (IFN-I and IFN-III) at the site of infection, but systemic IFN-I and IFN-III priming in distal organs and barrier epithelial surfaces, respectively. In this Review, we examine the relationship between SARS-CoV-2 biology and the cellular response to infection, detailing how antagonism and dysregulation of host innate immune defences contribute to disease severity of COVID-19.
Similar content being viewed by others
Introduction
The evolutionary success of a virus hinges on its ability to enter a cell and gain access to the raw genetic material, amino acids and basic cellular machinery that are required for de novo generation of progeny viruses. Given the value of these resources, cells have evolved surveillance strategies to detect their usage. In vertebrates, actively replicating viruses are identified through the recognition of foreign RNA or DNA structures. Virus-derived RNA can be discerned as a result of missing modifications (for example, 2′-O-methylation), inclusion of certain virus-distinguishing elements (for example, an exposed 5′-triphosphate) and/or the presence of extensive secondary structures (for example, double-stranded RNA (dsRNA))1. By contrast, virus-derived DNA can be identified by its location within the cell (for example, endosomal or cytosolic DNA) or by distinct aspects of sequence and/or structure not present in host DNA (for example, unmethylated CpG motifs)2. In both examples, cellular detection of these so-called pathogen-associated molecular patterns (PAMPs) within the infected cell is either direct or aided by autophagy, a process by which the cell engulfs and hydrolyses an internal portion of itself and subsequently releases the content back into the cytoplasm3. As these PAMPs represent unavoidable by-products of replication, viruses are under constant evolutionary pressure to minimize their levels, prevent their detection and/or block any consequential downstream biology4.
Viral strategies to subvert cellular detection, if successful, not only enable productive infection to occur in the cognate host, but can also facilitate zoonotic events, as many cellular defence components are conserved among phylogenetically related species5. For this reason, encroachment of one species into the ecosystem of another is a common source of zoonotic diseases and is a dynamic common to population overgrowth — as evidenced by the fact that many emergent viruses have been the product of deforestation6,7,8,9 (Box 1). Close contact with other vertebrate species through animal husbandry, live wet markets or the acquisition of bush meat is also a primary source for zoonotic transmission and is believed to have enabled the pandemic spread of human immunodeficiency virus and countless influenza viruses10,11. Zoonoses-enabling environments such as live wet markets also contributed to the emergence of two highly pathogenic coronaviruses, Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) in 2003 and SARS-CoV-2 in 2019, the latter being responsible for the coronavirus disease 2019 (COVID-19) pandemic12,13.
Despite the rapid development and deployment of vaccines and antivirals, SARS-CoV-2 caused more than 650 million infections and 6.6 million deaths from the beginning of 2020 to the end of 2022 (ref. 14). As global spread continues, SARS-CoV-2 undergoes countless rounds of selection: adapting to our biology, circumventing our immune defences and improving transmission efficiency. Individual strains that have acquired a competitive fitness advantage because of this selection have been defined as variants of concern (VOCs) and designated with letters of the Greek alphabet15. In this Review, we summarize the mechanisms employed by SARS-CoV-2 to circumvent our first line of innate antiviral defences and discuss how this relates to COVID-19 severity. For comparable reviews that focus on the later stages of the adaptive immune response to SARS-CoV-2, we refer the reader to several well-written and comprehensive summaries available elsewhere16,17.
Cellular response to SARS-CoV-2 infection
The genome of SARS-CoV-2, like all members of its genus, is ~30,000 nucleotides in length, with an invariant gene order. Starting at the 5′ end are the genes for replicase and other non-structural components (ORF1a and ORF1ab), followed by the structural genes for the spike (S), envelope (E), membrane (M) and nucleocapsid (N) proteins, amid a number of intergenic accessory factors13,18,19. The SARS-CoV-2 positive-sense, single-stranded RNA genome encodes ~30 proteins. These include the ORF1a and ORF1ab polyproteins (pp1a and pp1ab), which are cleaved by viral proteases (PLpro and 3CLpro) into 16 non-structural proteins (Nsps) that largely form the RNA-dependent viral replicase, four structural proteins that form the physical virion and nine accessory proteins not required for viral replication, but essential for hijacking aspects of cell biology13,18,19.
As with most viral infections, a crucial first step in the host innate immune response to coronavirus infections is the production of type I and type III interferons (IFN-I and IFN-III, respectively), as well as pro-inflammatory cytokines and chemokines20. IFN-I and IFN-III are produced by a variety of cell types following recognition of viral PAMPs and/or host danger-associated molecular patterns (DAMPs), such as exposed mitochondrial DNA, via specialized cellular pattern recognition receptors (PRRs)20. For SARS-CoV-2, RNA-based replication intermediates are thought to represent the major viral PAMP, recognized by the RIG-I-like and Toll-like receptors (RLRs and TLRs, respectively)21. The RLRs comprise a family of RNA-binding helicases that include two central intracellular sentinels, RIG-I and MDA5, in addition to a third RLR called LGP2, the function of which remains unclear22. Although they exhibit structural similarities, RLRs have different binding preferences, allowing them to make up a diversified and complementary defence system22. Generally, RIG-I is thought to sense RNAs with an exposed 5′-triphosphate, a common feature of negative-strand RNA viruses, whereas MDA5 activation is mediated by engaging dsRNA, a common motif formed during replication of all viruses22. In contrast to these intracellular sentinels, TLRs sample the extracellular milieu for the presence of PAMPs or DAMPs resulting from distal infections23. As it relates to the sensing of an RNA virus infection, extracellular dsRNA, single-stranded RNA or DNA can be sensed by TLR3, TLR7/TLR8 or TLR9, respectively23. Although all of these canonical sensing mechanisms, as well as some non-canonical systems, have been suggested to contribute to host recognition of SARS-CoV-2, MDA5 is thought to be most critical in this process24,25,26,27,28,29,30.
Cellular recognition of SARS-CoV-2 promotes the recruitment of adaptor proteins and production of IFN-I and IFN-III21. In the absence of interference, RLR activation would result following the production of viral subgenomic RNA (sgRNA), which can readily form dsRNA structures by engaging the genomic template21,31. MDA5 aggregates on dsRNA structures and subsequently engages the mitochondrial antiviral signalling protein (MAVS), leading to the recruitment of ubiquitin ligases and serine/threonine kinases that coordinate the activation of two central transcription factor family members, nuclear factor-κB (NF-κB, comprising subunits p50 and p65) and interferon regulatory factors (IRFs) such as IRF3 (refs. 21,31). Similarly, TLR recognition of an appropriate viral ligand initiates recruitment of either MyD88 or TRIF cellular adaptor proteins and triggers activation of NF-κB and IRFs through a similar signalling cascade32. Activation of these transcription factors, in addition to other factors associated with cellular stress, causes the formation of a large complex termed the enhanceosome, which binds upstream of the transcriptional start site of both IFN-I members (notably IFNβ) and IFN-III members (such as IFNλ-1, IFNλ-2 and IFNλ-3) to initiate the antiviral response33. Despite inducing similar transcriptional outputs, several characteristics give these two antiviral cytokine families distinct biologies, including the makeup and distribution of cell surface receptors for IFN-I versus IFN-III, variations in protein stabilities, and the magnitude, kinetics and anatomical location of each response34. Whereas IFN-I can prime almost any cell to induce an antiviral state, IFN-III activity is more selective and thought to be limited to epithelial barrier tissues, including the respiratory and gastrointestinal tracts as well as the blood–brain barrier34.
IFN-I and IFN-III signalling functions in both autocrine and paracrine manners to promote upregulation of interferon-stimulated genes (ISGs) such as IRF7, a transcription factor related to IRF3 that exhibits more promiscuous DNA-binding activity35. The addition of IRF7 expands the transcriptional output resulting from virus recognition to include more members of the IFN-I family, most notably the IFNα variants36,37. Induction of IRF7 and other ISGs is mediated by IFN-I-dependent or IFN-III-dependent receptor dimerization that culminates in the activation and assembly of a transcriptional complex termed IFN-stimulated gene factor 3 (ISGF3), which comprises members of the signal transducer and activator of transcription (STAT1 and STAT2) family, as well as an additional IRF family member (IRF9)34,38. Activated ISGF3 then migrates to the nucleus, where it orchestrates the transcription of hundreds of ISGs that exert antiviral effects directly and indirectly through a variety of mechanisms, including repression of viral replication, inhibition of viral transcription/translation and degradation of viral nucleic acids34.
The importance of IFN signalling as it relates to the host response to coronaviruses was first illustrated by studies in animals lacking expression of the IFN-I receptor (IFNAR1−/−), in which strains of mouse hepatitis virus (MHV) that normally produce mild illness became fatal39,40. The basis for this phenotype stems from the fact that coronaviruses provoke a systemic IFN-I response in the host that serves to protect distal organs from subsequent infection39,40. In the absence of this response, distal tissues are left susceptible to low levels of circulating virus, which is often associated with enhanced disease41. This dynamic is observed in young patients with life-threatening COVID-19 who were found to possess mutations in IFNAR1 or TLR genes42,43 and in individuals of advanced age for whom evidence of IFN-I-autoantibodies has been reported44,45. Differences in the anatomical expression of IFN-I or IFN-III, ISGs and PRRs during infection have also been associated with disease severity in patients with COVID-19 in an age-dependent manner46,47,48.
In addition to inducing IFN-I or IFN-III, vertebrates also generate pro-inflammatory cytokines and chemokines to combat virus infections. Unlike IFN-I, which represents a ‘call to arms’ at the site of infection, cytokines and chemokines aid in coordinating a ‘call for reinforcements’ from more distal sites16,17. When functional, these two complementary pathways work to slow replication at the site of infection and provide time for the adaptive immune response to develop the capacity for antigen-specific recognition, establishing a formidable barrier to the evolutionary success of any virus. Unfortunately, viruses have evolved countless strategies that interfere with these host defences, leading to often unpredictable disease outcomes that result from the virus directly, from an aberrant host response, or from a combination of the two. In the following section, we describe the current state of knowledge regarding SARS-CoV-2-mediated manipulation of host biology and provide insight as to how this viral interference contributes to disease severity.
SARS-CoV-2 evasion of host defences
As obligate intracellular parasites, all viruses depend on the host for energy, raw materials and access to complex biologic machines. To establish a productive infection, a virus must usurp or inactivate extensive host pathways. SARS-CoV-2 invests substantial resources to block the establishment of the antiviral response (Table 1). These strategies are divided into five broad categories and are described in detail below.
Minimizing and masking inflammatory RNA
The first step of the coronavirus infection cycle involves binding of the trimeric S glycoprotein to the host cell receptor, which leads to either an ‘early’ entry pathway of direct fusion with the cellular plasma membrane or a ‘late’ entry pathway of receptor-mediated uptake via endocytosis49,50. In the case of SARS-CoV-2, both pathways are mediated through binding of the S trimer to host angiotensin-converting enzyme 2 (ACE2)51. For direct fusion with the host membrane to occur, the S protein must be cleaved during egress of the virion and then again during the entry process. For SARS-CoV-2, this first event is largely mediated by the furin protease, and the second by transmembrane serine protease 2 (TMPRSS2) at the cell surface51. Alternatively, dual cleavage of the S protein can also be mediated by cathepsin proteases during receptor-mediated endocytosis51. In either event, once the genomic viral ribonucleoprotein complex enters the cytoplasm, the genome dissociates from the viral N protein and can be directly translated by the host ribosome thereafter50.
Initial translation of the viral genomic RNA must conform to the rules of host translation, both for functionality and to avoid host detection. To this end, coronaviruses modify the 5′-triphosphate (pppA) of their genomic RNA and sgRNAs through capping and methylation to ensure ribosomal loading and avoid host detection (Fig. 1). Formation of the coronaviral cap structure entails four sequential enzymatic reactions52. First, a 5′ RNA triphosphatase removes the γ-phosphate from the pppA of the nascent mRNA to form 5′-diphosphate (ppA). Next, a guanylyltransferase generates the cap core structure by transferring a guanosine monophosphate (GMP) to the ppA. After the core structure is formed, the guanine is methylated at position N7 by an N7-methyltransferase. Finally, a 2′-O-methyltransferase methylates the ribose at the 2′-O position of the first transcribed nucleotide, producing the final cap structure: 7MeGpppA2′OMe. Studies performed with SARS-CoV-2 and other coronaviruses have demonstrated that Nsp13, Nsp14 and Nsp16 function in the viral capping process as a 5′ RNA triphosphatase, N7-methyltransferase and 2′-O-methyltransferase, respectively53,54,55,56,57,58. SARS-CoV-2 Nsp12 has also been suggested to participate as a guanylyltransferase in this process52,57. Additional studies have shown that Nsp10 may participate in viral capping by serving as a cofactor for the activities of Nsp14 and Nsp16, a function that is conserved among coronaviruses58,59,60,61.
SARS-CoV-2 enters into a host cell by binding angiotensin-converting enzyme 2 (ACE2) on the cell surface, a process that can be facilitated by transmembrane protease serine 2 (TMPRSS2), which provides proteolytic cleavage of the viral spike (S) protein to promote virus–host fusion. Following internalization of the viral particle, the capped and polyadenylated genomic viral RNA is released into the cytoplasm where it can be directly translated (stage 1). Initial translation of viral genomic RNA results in the production of the ORF1a and ORF1ab polyproteins (pp1a and pp1ab) that are subsequently processed by viral proteases to form the replicase and non-structural proteins (Nsps; depicted in yellow) necessary to establish replication organelles (ROs). Nsp3 and Nsp4 mediate the modification of endoplasmic reticulum (ER) membranes into convoluted membranes (CM) and double-membrane vesicles (DMVs) that make up ROs, whereas Nsp6 forms a zippered molecular tether between ROs and the ER that enables the flow of lipids (stage 2). Nascent viral RNA is modified by Nsp enzymes (depicted in blue) to mimic host transcripts and minimize the ability of the cell to induce a defence. First, Nsp13 (a 5′ RNA triphosphatase) removes the phosphate from the 5′ end of the viral RNA. This is followed by the transfer of a guanosine monophosphate to the 5′ end by Nsp12 (a guanylyltransferase) to yield the cap core. Subsequently, Nsp14 (an N7-methyltransferase) and Nsp16 (a 2′-O-methyltransferase) assisted by the Nsp10 capping cofactor catalyse the final methylation steps necessary to complete the viral cap (stage 3). As viral replication proceeds, negative-sense RNA (−ssRNA) and double-stranded RNA (dsRNA) intermediates are sequestered inside ROs to prevent host detection. In parallel, the positive-sense, single-stranded genomic and subgenomic RNAs (+ssRNA) needed for translation of viral proteins and de novo virion assembly are chaperoned from the replication organelles. As replication intensifies, viral RNAs accumulate outside of ROs, and are masked and/or minimized by the SARS-CoV-2 nucleocapsid protein (N) and/or Nsp15, depicted in orange (stage 4).
Because viral mRNA is disguised as host mRNA, translation and assembly of the replicase should not initially elicit a cellular response; however, as viral RNA synthesis proceeds, dsRNA intermediates inevitably begin to form that can trigger host immune activation. The reason for this is that both genomic RNAs and sgRNAs must proceed through negative-strand intermediates. This accumulation of RNA transcripts with opposing polarities and complementary sequences increases the potential for dsRNA formation. For SARS-CoV-2 and other coronaviruses, early detection of dsRNA dictates the overall fitness of the virus. Host induction of antiviral defences before the completion of the virus life cycle would be enough to confine the infection to only those cells that were initially infected. Therefore, before dsRNA production, the virus must ensure that strategies are in place to either minimize its accumulation or hide it from host sensors: SARS-CoV-2 utilizes both strategies in parallel. The highly conserved Nsp15 has been reported to be responsible for minimizing the accumulation of negative-stranded RNA and dsRNA via its endonuclease activity, which targets negative-sense transcripts62,63. In parallel, SARS-CoV-2 induces the assembly of double-membrane vesicles (DMVs), which have been suggested in other coronaviruses, such as MHV, to compartmentalize viral replication away from cellular sensors64,65. These so-called replication organelles (ROs) associate with the endoplasmic reticulum and have been observed during infection with MHV, SARS-CoV, Middle East respiratory syndrome coronavirus (MERS-CoV) and SARS-CoV-2 (refs. 64,66,67,68,69). The process of establishing ROs is also a means of diminishing host capacity to recognize the presence of the virus infection by interfering with and/or exploiting autophagy biology70,71. RO assembly during SARS-CoV-2 infection is mediated by Nsp3 and Nsp4 function, with Nsp6 forming a molecular tether to the endoplasmic reticulum that enables the flow of lipids69 (Fig. 1). As it takes time to accumulate sufficient levels of Nsp3, Nsp4, Nsp6 and Nsp15, optimal infection is achieved with a low multiplicity of infection so that replicase assembly and early genome replication do not generate large amounts of dsRNA before the formation of DMVs or the translation of adequate levels of Nsp15. Moreover, differences in the innate immune activation of emerging VOCs may be observed as a result of non-synonymous changes that affect the abundance, expression or function of viral proteins involved in this process69,72.
Blocking host recognition
Following successful establishment of viral ROs, SARS-CoV-2 transcripts exit the DMVs through specialized molecular pores and enter the cytosol, where they undergo translation and begin virion assembly73. Outside the protection of the sealed ROs, viral RNAs are subject to potential recognition by cellular sensors (Fig. 2). As replication intensifies, avoiding host detection by excluding any inflammatory viral RNAs would rapidly become impossible. Consequently, coronaviruses also devote substantial resources to inhibit cell signalling pathways that ensue following recognition.
Virus infection generates replication intermediates and/or induces the formation of stress granules that serve as platforms for RIG-I-like receptor (RIG-I or MDA5) activation. Host recognition of viral pathogen-associated molecular patterns, such as single-stranded RNA (ssRNA) with an exposed 5′-triphosphate or double-stranded RNA (dsRNA), promotes the assembly of a mitochondria-localized signalling hub orchestrated by mitochondrial antiviral signalling protein (MAVS), and culminates in the activation of host kinases IKKα, IKKβ and TBK1. Kinase activation induces the production of interferon-β (IFNβ) through cooperative engagement of the ATF2–JUN, interferon regulatory factor 3 (IRF3) and nuclear factor-κB (NF-κB) transcription factors. Secreted IFNβ functions in an autocrine or paracrine manner to promote an antiviral state in cells. On binding of IFNβ, the type I IFN receptor subunits on the cell surface dimerize, bringing together the receptor-associated kinases, Janus kinase 1 (JAK1) and tyrosine kinase 2 (TYK2), which subsequently activate each other via transphosphorylation and promote the recruitment and phosphorylation of the signal transducer and activator of transcription (STAT) molecules, STAT1 and STAT2. Phosphorylated STAT1 and STAT2 form a stable complex with interferon regulatory factor 9 (IRF9) that translocates into the nucleus, where it promotes the transcription of IFN-stimulated genes (ISGs). Each of these processes is the target of SARS-CoV-2 interference, as illustrated here and further described in Table 1. Viral proteins that inhibit aspects of host recognition and the associated signalling pathways are shown in purple, whereas those that block components of the IFN signalling pathway are depicted in teal. Owing to the ability of Nsp1 to more generally inhibit protein synthesis, its role in specifically blocking elements of these pathways remains uncertain (question marks). M, membrane protein; N, nucleocapsid protein; P, phosphorylation; S, spike protein.
To navigate this advanced surveillance system, several SARS-CoV-2 proteins have been suggested to target viral sensing processes and delay the production of host antiviral defences (Fig. 2). A central viral component proposed to be involved in this role is the N protein. As one of the most abundant viral proteins produced during infection, this RNA-binding protein is critical for packaging viral genome into virions18,74. As it relates to immune antagonism, the inherent ability of N to associate with free RNA may also serve to mask it from detection or prevent it from forming extensive base pairs with complementary sequences75,76,77,78. Supporting this hypothesis, in vitro studies using the Alpha (B.1.1.7) VOC suggested that the enhanced IFN antagonism observed with this variant may correlate with increased N protein expression compared with earlier viral lineages; additional explanations such as an overall reduction in dsRNA production or increases in the expression of other non-structural genes remain possible79,80. A recent study also suggested that fragments of the N protein produced after cleavage by host caspase 6 may themselves dampen the host IFN response, further adding to the growing evidence that N can act as an IFN antagonist81. The culmination of this potential viral interference by N would be a delay in innate immune activation, providing SARS-CoV-2 critical time to replicate and spread in an optimal cellular environment.
Another mechanism for inducing RLR signalling involves the formation of stress granules31 (Fig. 2). These membrane-less cytoplasmic aggregates of translationally stalled mRNAs and RNA-binding proteins form under various stress conditions, including viral infection, and act as platforms for RLR signalling pathways31. Nsp5, which is the main protease of SARS-CoV-2, can inhibit the formation of these stress granules independently of its enzymatic activity82. The SARS-CoV-2 N protein can also block formation of stress granules by binding and sequestering both RNA and a specific factor called G3BP1, which not only functions as the nucleating protein for stress granules but also serves as a liaison between stress granules and the RLRs31,82.
In addition to enacting mechanisms to sequester viral PAMPs, SARS-CoV-2 proteins can also block PRR activation. The N protein has been shown to bind to the DExD/H box RNA helicase domain of RIG-I, interrupting its interaction with TRIM25, a cellular ubiquitin ligase that potentiates RLR signalling through K63-ubiquitin-mediated activation of RIG-I83,84,85. However, the conclusions of these studies are based on overexpression systems and do not account for the fact that both genomic and subgenomic SARS-CoV-2 RNAs are capped or that RIPLET, not TRIM25, is believed to be the more relevant adaptor for RIG-I activation86,87. Similarly, the Nsp3 protein, also known as the SARS-CoV-2 papain-like cysteine protease (PLpro), has been reported to antagonize conjugation of an ISG, called ISG15, to MDA5, but these data also rely on overexpression experiments and focus on an activation pathway that has unclear physiological relevance88,89. Nsp3 may play a more central role in host immune evasion owing to its general ability to interfere with other protein modifications, including ubiquitin and/or ADP-ribose conjugations90,91,92,93,94. Despite sequence divergence, the ability of Nsp3 to alter post-translational modifications is largely conserved among coronaviruses, suggesting an important global role for this protein in host evasion92,95.
As cellular recognition of virus infection is unlikely to be completely prevented, other viral components target host factors further downstream in the signalling cascade. This strategy enables the virus to interfere with multiple antiviral surveillance systems that converge on the activation of certain cellular adaptors. For example, overexpression studies of N, M, Nsp5 and ORF7b have all generated data supporting the capacity of SARS-CoV-2 to interfere with MAVS biology82,83,84,85,96,97,98,99 (Fig. 2). Although the mechanisms of action against MAVS have not been fully elucidated, recent evidence has suggested that the N protein may inhibit MAVS polyubiquitination and aggregation in a manner dependent on the N dimerization domain100. Notably, this domain is also required for the liquid–liquid phase separation (LLPS) of N on its interaction with viral RNA100. In addition to its probable role in mediating viral assembly, LLPS of N has also been associated with its ability to interact with stress granules and components of the NF-κB signalling pathway, providing additional evidence supporting its role in immune evasion and modulation101,102,103. Other sensor-targeting strategies employed by SARS-CoV-2 proteins include direct ubiquitination and proteasome-mediated degradation of MAVS, an activity mediated by Nsp5 (ref. 96). Similarly, the M protein of SARS-CoV-2 has been reported to interfere with the ability of MAVS to establish the necessary scaffolding required for downstream transcription factor activation99. SARS-CoV-2 M, Nsp6, Nsp13, ORF7a and ORF9b proteins have also been shown to target and inhibit the function of additional host factors involved in MAVS signalling such as TBK1, the kinase responsible for IRF3 activation97,104,105,106,107,108,109,110,111. In particular, Nsp6, Nsp13 and ORF9b have each been suggested to bind to TBK1, preventing its phosphorylation-mediated activation105,106,107, whereas M and ORF7a have been reported to reduce TBK1 expression97,104.
One unique aspect of SARS-CoV-2 as it relates to PRR signalling is that the virus induces NF-κB activation, despite its involvement in antiviral signalling (Box 2). The basis for this activity is thought to relate to NF-κB-dependent transcriptional outputs that are not related to cellular defences but are required by the virus112. Moreover, as SARS-CoV-2 specifically targets IRF3, active NF-κB signalling is not sufficient to restore IFN-I induction. The S, Nsp3 and Nsp5 viral proteins have all been reported to reduce the expression of IRF3 through a variety of proposed models91,113,114,115,116,117. Nsp5, Nsp12, ORF3b and M also block the nuclear translocation of IRF3 following PRR signalling events114,118,119,120. For Nsp5 and Nsp12, this suppression was independent of both the phosphorylation state of IRF3 and the respective protease and polymerase activities of Nsp5 and Nsp12 (refs. 114,118). Inhibition of IRF3 phosphorylation has also been reported for Nsp1, although the mechanism may involve a more general shutoff of global translation, which depletes the cellular factors necessary for this process to occur, as further discussed below121.
Blocking interferon signalling
Single-cell RNA sequencing of SARS-CoV-2-infected cells has revealed that more than 60% of the total mRNA found in an infected cell can be virus-derived, illustrating the efficiency with which the virus usurps the cell112,122. This outcome is the product of high levels of viral genomic RNA and sgRNA transcription, coupled to the Nsp1-mediated suppression of host mRNA123. Further contributing to this hostile cellular takeover is the targeted inhibition of host translation and RNA splicing mediated by Nsp10 and Nsp14 (refs. 124,125). However, even with these potent viral countermeasures in place, SARS-CoV-2 ultimately induces cell death through multiple mechanisms126, enabling detection of the virus by phagocytic cells127. It is this dynamic that is likely responsible for the high levels of IFN-I and IFN-III observed in response to SARS-CoV-2 infection, especially in patients with severe COVID-19 (refs. 128,129,130). As this biology cannot be easily blocked, SARS-CoV-2 also benefits from targeting the signalling cascade responsible for responding to IFN-I and IFN-III and inducing ISG expression (Fig. 2).
SARS-CoV-2 Nsp13 and Nsp14 interfere with IFN-I signalling by reducing the expression of the IFNAR1 receptor subunit131. In addition, Nsp13 and the S protein both interact with STAT1, preventing its docking to the receptor and subsequent phosphorylation-mediated activation119,132. Similarly, reduction of host transcripts mediated by Nsp1, Nsp10 and Nsp14 also contributes to blocking ISG production121,124. Interference with ISG induction is especially important for viruses, as infection of a cell primed with IFN-I or IFN-III is both non-productive and serves to amplify the global host response at a time when the virus is still vulnerable to detection133.
Blocking nuclear transport
Whether through blocking the translocation of transcription factors or by preventing the export of host mRNA, interfering with the nuclear transport machinery can provide an immediate selective advantage for a cytoplasmic pathogen. The nuclear pore complex (NPC) is a large structure that bridges the inner and outer membranes of the nuclear envelope and forms an aqueous channel through which nucleocytoplasmic transport is regulated134,135. The NPC is composed of various protein subunits, called nucleoporins (Nups), that interact with soluble nuclear transport receptors of the karyopherin protein family (importins and exportins) responsible for shuttling specifically tagged proteins between the nucleus and the cytoplasm. Viruses from many different families co-opt this machinery to allow specific viral proteins to gain access to the nucleus as needed, and/or to block host components from carrying out functions that restrict viral replication134. Analyses of protein–protein interactions that occur between SARS-CoV-2 and host factors revealed that Nsp1, Nsp9, Nsp15, ORF6 and M are all capable of interacting with the host nuclear transport machinery119,136,137,138 (Fig. 3).
Cellular induction of an antiviral response is dependent on bidirectional trafficking through the nuclear pore complex (NPC). With the aid of nuclear transport receptors (importins), cellular transcription factors that are induced in response to infection translocate through the NPC into the nucleus and bind sequences within antiviral genes to drive their expression. Following de novo transcription and capping of the nascent messenger RNA (mRNA), the host spliceosome assembles at RNA splicing sites and promotes intron excision to yield translationally competent mature transcripts. These transcripts associate with nuclear transport receptors (exportins) and are exported through the NPC into the cytoplasm, where they are translated by host ribosomes and routed for proper folding and cellular localization based on recognition of their signal peptide by the signal recognition particle (SRP). SARS-CoV-2 encodes several proteins that block nuclear transport (depicted in purple), including non-structural proteins (Nsps), the open reading frame 6 (ORF6) accessory protein and the membrane structural protein (M). This inhibition is facilitated by interactions with the host proteins indicated (KPNA2, karyopherin subunit α2; KPNA6, karyopherin subunit α6; NTF2, nuclear transport factor 2; Nup62, nucleoporin 62; Nup98, nucleoporin 98; NXF1, nuclear RNA export factor 1; NXT1, nuclear transport factor 2-like export factor 1; Rae1, ribonucleic acid export factor 1). SARS-CoV-2 also encodes proteins that ultimately shut off translation (depicted in red) by inhibiting host RNA splicing, preferentially blocking host RNAs for nuclear export in favour of viral RNAs, interfering with ribosomal function and preventing protein trafficking. As Nsp1 is also capable of more generally inhibiting protein synthesis, its role in preferential targeting of viral RNAs remains uncertain (question marks).
Numerous independent research groups have demonstrated that ORF6 inhibits nucleocytoplasmic transport by binding to the complex formed by the IFN-inducible nucleoporins, Nup98 and ribonucleic acid export factor 1 (Rae1), and drawing them away from the NPC104,136,139,140,141,142. Crystal structure data analysis of this interaction demonstrated that ORF6 outcompetes RNA for access to the mRNA-binding groove of the Nup98–Rae1 complex143. By targeting this complex, ORF6 blocks nuclear translocation of transcription factors, including IRF3 and the individual components of ISGF3, and it prevents mRNAs, such as those encoding IFN-I, IFN-III or antiviral ISGs, from entering the cytoplasm for translation139,140,141,142,144. Notably, ORF6 may also contribute to observable differences between SARS-CoV and SARS-CoV-2, as the ORF6 from SARS-CoV-2 generates a modified structure associated with stronger anti-IFN activity than the ORF6 from SARS-CoV142,145.
Shutting off translation
In addition to preventing an infected cell from utilizing the resources of its own defence system, many viruses also target aspects of protein synthesis to enhance fitness146. During eukaryotic translation, the 5′ cap and poly-A structures of mature mRNA transcripts are recognized by the 40S ribosomal subunit, a process regulated by eukaryotic initiation factors147. On scanning of the AUG initiator codon, the 60S ribosomal subunit joins the complex to form the 80S ribosome, and elongation of the polypeptide chain proceeds147. As translation of viral proteins also relies on the host machinery, virus-mediated interference with these processes must retain some selectivity to be successful.
Nsp1 is the best-characterized example of a coronavirus protein that shuts off host translation, although equivalent proteins from different viruses seem to use distinct strategies for inhibition148. Several groups have demonstrated that SARS-CoV-2 Nsp1 blocks the mRNA entry channel to the ribosome by binding to the 18S ribosomal RNA component of the 40S ribosomal subunit149,150,151,152,153 (Fig. 3). In fact, the C terminus of Nsp1 was found to exhibit structural similarity to two known ribosome inhibitors, SERBP1 and Stm1, which also prevent mRNA from accessing the entry channel of the 40S ribosome149. Moreover, when the C-terminal domain of Nsp1 was mutated to ablate its interaction with the ribosome, a loss of translational repression was observed106,149,150,153. This interaction between Nsp1 and the ribosome has been associated with a robust reduction in the translation of endogenous proteins in human cells, even in the absence of other SARS-CoV-2 proteins106,149,150,151. Although some groups have reported that mRNAs containing the 5′ viral leader sequence were largely protected from Nsp1-mediated translation inhibition149,154, others found this activity to be non-discriminatory150,153,155. In each of these studies, the 5′ and/or 3′ untranslated region of SARS-CoV-2 was fused to a reporter gene, so it is possible that other features of viral mRNA may play a role in escaping translational shutdown. One study additionally demonstrated that inclusion of the viral 5′ untranslated region resulted in a fivefold increase in the translation of reporter mRNA, suggesting that viral RNA may be translated more efficiently than host mRNAs, thus compensating for the translational block150. Alternatively, the Nsp1 protein may induce degradation of mRNAs that lack the 5′ viral leader sequence, thus allowing viral mRNAs to be selectively translated over cellular mRNAs155. As the Nsp1 protein has been implicated in several processes related to the evasion of host innate immunity, it is important to consider how its ability to globally suppress protein translation may contribute to its other reported functions.
In addition to the well-documented involvement of Nsp1, other viral proteins such as Nsp10, Nsp14 and Nsp16 also play a role in translational shutoff during SARS-CoV-2 infection (Fig. 3). In order for nascent RNA transcripts to be converted into translationally competent mature mRNAs, intervening sequences (introns) must be cutout so that expressed sequences (exons) can be joined together156. This process, known as RNA splicing, is mediated by an RNA–protein complex termed the spliceosome, which comprises a group of small, non-coding, nuclear RNAs and protein splicing factors156. As disruption in splicing can often equate to a selective advantage for a virus that does not require this biology, this is another commonly observed target for RNA viruses, including SARS-CoV-2. In this regard, SARS-CoV-2 Nsp16 has been shown to be capable of binding to the mRNA recognition domains of U1 and U2, two of the major small nuclear RNA subunits that make up the spliceosome, resulting in global inhibition of host mRNA splicing during infection149. Similar interference with global host expression that has also been found to impact splicing has been reported for Nsp10 and Nsp14 (refs. 124,125). Perturbations in host splicing activity as a result of SARS-CoV-2 infection have been observed in infected lung samples from patients with COVID-19 (ref. 157).
To ensure proper folding and trafficking of newly formed proteins, nascent ribosome-associated peptide chains are continually scanned by the signal recognition particle (SRP) for hydrophobic signal peptides that identify products destined for secretion or insertion into a host membrane158. On signal recognition, the SRP triggers the ribosome to translocate to the endoplasmic reticulum, where those proteins can be properly formed and routed. SARS-CoV-2 Nsp8 and Nsp9 have each been found to bind to the 7SL RNA scaffold component of the SRP complex, disrupting protein trafficking and resulting in degradation of newly translated proteins149. Specifically, Nsp8 binds to the region of 7SL bound by the SRP54 protein, which is the component of SRP responsible for signal peptide recognition, whereas Nsp9 binds to the region of 7SL that interacts with the SRP19 protein, which is required for proper folding and assembly of SRP itself149. This interference with SRP structure and function was associated with reduced protein integration into the cell membrane of SRP-dependent membrane proteins149. As a number of immune mediators, including IFN-I and IFN-III family members, are known to be secreted in this way, this study suggests an important role for Nsp8 and Nsp9 in evasion of host immunity (Fig. 3).
Systemic inflammation and clinical presentation
Despite the ability of SARS-CoV-2 to block and evade innate immunity in the context of directly infected cells, these immune mechanisms remain intact in uninfected bystander cells, which are capable of sensing and responding to the debris from dying or damaged cells112. Ironically, this dynamic leads to an abnormally robust innate immune response, albeit delayed, as SARS-CoV-2 replication can initially proceed unabated in the airways until the infected cells begin to die and new progeny virions are released by the thousands159. As a result, initial engagement of the host response initiates at a time of relatively high virus load when viral PAMPs become readily detectable29,112. This dynamic results in elevated production of IFN-I and IFN-III, which spread beyond the airways and begin inducing inflammatory processes in all distal tissues. Even in the absence of virus infection, this systemic response can impose substantial stress on the body and materialize into the diverse clinical presentations that have been associated with COVID-19.
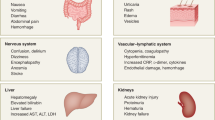
Arguably one of the paradoxes of SARS-CoV-2 biology is the capacity of this respiratory virus to bring about extrapulmonary clinical manifestations, especially gastrointestinal and/or cardiovascular ones160,161,162. Countless clinical case studies have documented virus-associated transcriptional changes in every organ of the body during acute infection, a phenomenon that can be phenocopied in the hamster model of COVID-19 (refs. 41,163,164,165). Moreover, a notable proportion of individuals with COVID-19 who clear the virus can retain protracted disease symptoms involving multiple organs162. Although an understanding of the underlying biology for the acute and persistent presentations of COVID-19 remains incomplete, the host response to SARS-CoV-2 infection undoubtedly initiates these processes.
It is tempting to speculate that the diverse clinical presentations of SARS-CoV-2 may be a product of some unique aspect of the virus. Although possibly true for a subset of the many documented systemic conditions, it should be noted that several disease outcomes associated with COVID-19 are also commonly observed in response to other respiratory infections, such as influenza A virus (IAV), suggesting that the extent of disease may instead be a by-product of strain-specific attributes, such as viral fidelity or interference, that might impact the production of inflammatory material or the kinetics by which this material can be sensed166,167. This concept is supported by recent publications demonstrating that infection of different small-animal models with either SARS-CoV-2 or an H1N1 IAV strain can result in comparable transcriptional signatures in organs distal to the initial site of infection168,169. Isolation of infectious virus outside the airways in non-immunocompromised individuals infected with either virus is rare, suggesting that virus-mediated damage to the airways results in a substantial host response that travels through the circulation and stimulates every organ system. As underlying conditions in any one of those organs could be exacerbated by such a stimulus, diverse clinical presentations would be anticipated.
In contrast to the observed similarities in host immune response to IAV versus SARS-CoV-2, there are some clinical presentations that suggest SARS-CoV-2 does exhibit unique attributes, especially regarding its ability to induce post-acute sequelae (also known as long COVID). Although IAV and SARS-CoV-2 enact similar strategies to gain cellular access, the viral fusion proteins responsible for these activities utilize different cellular receptors. The resulting difference in cell tropism could help explain the propensity for SARS-CoV-2 to lead to some conditions associated with long COVID, including anosmia (loss of smell), which is a commonly observed clinical presentation and unique feature of COVID-19. SARS-CoV-2 productively infects sustentacular cells in the olfactory epithelium, resulting in the same host response and cell death as that observed in the airways170. Following viral infection, there is evidence that dying sustentacular cells release dsRNA-laden material capable of triggering microglial and macrophage activation and robust engagement of antiviral defences in the olfactory system, events that are not observed in response to IAV infection170,171. The likely resulting production of IFN-I, IFN-III and pro-inflammatory cytokines from these bystander cells could then induce immune priming in the olfaction system in a dynamic akin to how respiratory infection can prime distal organs. An important distinction in the olfaction system, however, is that when olfactory neurons respond to these types of immune cue, their normal function is disrupted, which may explain the development of anosmia170. This same phenomenon is also a probable link to the underlying cause of other neurological dysfunctions following infection with SARS-CoV-2 (ref. 169). Together, these data suggest that the pathobiology of a given virus infection can be defined by the magnitude of inflammatory material available for sensing, in combination with the physical location in which this material is generated and/or deposited.
Host-targeted immunotherapeutics for COVID-19
RNA virus populations undergo logarithmic expansion and generate a landscape of small mutations in the process, enabling them to escape the selective pressures imposed by our immune defences and/or administration of antiviral drugs. One strategy to circumvent this evasion is by targeting a host factor that is required for virus biology, but not for host biology. For example, host factors involved in, but not critical for, membrane trafficking, lysosome regulation and chromatin remodelling have been identified via whole genome CRISPR–Cas screens as being essential for SARS-CoV-2 infection49,172,173. Small compounds designed to competitively engage such factors would thus impose a formidable selective pressure on the virus. A complementary approach is to directly target the host response to virus infection. Common examples of this strategy include compounds that diminish pain, fever and/or malaise. In addition, pathogenesis resulting from an imbalanced host response to viral infection can be diminished using immune-modulating compounds. Below, we summarize host-targeted immunotherapeutics that have demonstrated clinical value in treating hospitalized patients with COVID-19 (Table 2).
Corticosteroids
Multiple randomized controlled clinical trials (RCTs) have demonstrated the effectiveness of systemic corticosteroid treatment in a subset of hospitalized patients with severe COVID-19, presumably due to its ability to generally mitigate SARS-CoV-2-induced inflammation. In a large (n = 6,425), open-label RCT (RECOVERY), low-dose dexamethasone decreased mortality in patients receiving either invasive mechanical ventilation or oxygen alone, but not among those receiving no respiratory support174. A WHO meta-analysis of seven RCTs including a total of 1,703 critically ill patients with COVID-19 also reported a reduction in mortality following administration of dexamethasone175. As a result of these findings, current NIH guidelines recommend the standard use of dexamethasone or other systemic corticosteroids in hospitalized patients with COVID-19 who require mechanical ventilation or oxygen support176.
IL-1 and IL-6 inhibitors
Early studies characterizing the host responses to SARS-CoV-2 highlighted the development of a life-threatening hyperinflammatory response that was characterized by the robust induction of both IL-1 and IL-6. This finding drew parallels to cytokine release syndrome, an aberrant and life-threating immune response commonly associated with various cancer treatments177,178,179,180. As a result, inhibitors of IL-1 (anakinra and canakinumab) or IL-6 (tocilizumab and sarilumab), which are routinely co-administered to combat cytokine release syndrome, were tested individually to assess their therapeutic value for COVID-19 (refs. 181,182). Although some clinical trials reported improvements in inflammatory biomarkers and amelioration of lung pathology, others were inconclusive or demonstrated only modest value in reducing progression to more severe disease post-hospitalization183,184,185,186,187. Despite insufficient evidence to support monotherapy with IL-6 inhibitors, their use in the context of corticosteroid treatment in a subset of hospitalized patients requiring ventilation was clinically beneficial184,188. Efforts are ongoing to assess the value of simultaneously blocking IL-1 and IL-6 pathways (Clinicaltrials.gov; NCT05279391)189.
JAK inhibitors
In response to SARS-CoV-2 infection, a variety of immune-related activities are triggered through cytokine-mediated induction of Janus kinase (JAK)–STAT-dependent signalling events. A subset of therapeutic inhibitors of this pathway have shown benefit in the treatment of individuals hospitalized with COVID-19. The COV-BARRIER RCT (n = 1,525), which looked at patients with COVID-19 pneumonia and at least one elevated inflammatory marker, demonstrated increased survival with baricitinib (JAK1 and JAK2 inhibitor) when included with corticosteroids in standard care190. Clinical benefit of baricitinib in hospitalized patients with oxygen requirements was also seen in the ACTT-2 RCT (n = 1,033), although the drug was not evaluated in combination with corticosteroids191. Additionally, the double-blind, placebo-controlled STOP-COVID RCT of 289 hospitalized patients on low-flow oxygen demonstrated a reduction in risk of respiratory failure and death with use of tofacitinib (JAK1, JAK2 and JAK3 inhibitor) in combination with dexamethasone192.
At present, a direct antiviral targeting the main viral protease (Paxlovid, manufactured by Pfizer) is the preferred treatment for high-risk, symptomatic individuals with SARS-CoV-2 (ref. 176). If disease continues to develop and hospitalization is required, initial efforts are focused on preventing clotting while reducing general inflammation in the patient with inexpensive steroids. The immune-modulating drugs described above, as well as others still under evaluation, serve as experimental approaches that might be used should standard treatment prove ineffective.
Conclusions
The COVID-19 pandemic has imposed a major burden on global health, the full extent of which remains unclear. Despite being predominantly restricted to the airways, SARS-CoV-2 infection induces system-wide innate immune activation, which can manifest clinically as diverse extrapulmonary presentations alongside the more common respiratory complications165,169,193. In addition to the countless infections and millions of deaths worldwide, SARS-CoV-2 also harbours the unique capacity to generate persistent symptoms for weeks to months following infection — a clinical phenomenon colloquially referred to as long COVID. Based on our present understanding of SARS-CoV-2 biology, the underlying driver of many acute disease outcomes has become clearer, whereas others, especially those pertaining to long COVID, remain enigmatic.
Initial infection in the airways results in productive viral replication, transforming infected cells into viral factories and hampering first-line IFN-mediated defences required to slow this process177. Complicating things further, SARS-CoV-2 also engages the cell to promote a pro-viral environment, which incidentally activates aspects of later-stage innate immune defences112. This imbalanced response consequently recruits pro-inflammatory cells to the airways while virus replication progresses unchecked, causing extensive inflammation and respiratory damage. As infected cells die, inflammatory material associated with the virus begins to appear in the extracellular milieu, enabling the induction of the IFN response by local bystander cells. As the virus population expands, accumulation and detection of inflammatory debris accelerates, resulting in an IFN signature that can be documented in every organ of the body165,169. This response, which can persist for 7–10 days following infection, offers some protection against distal infections, but can also result in organ dysfunction should any underlying condition exist. In healthy individuals, the acute phase of infection can materialize with few consequences and result in successful virus neutralization and resolution of infection164.
Although the initial virus–host dynamics are now generally understood, the long-term consequences of SARS-CoV-2 infection remain unclear. Long COVID encompasses a myriad of clinical presentations with no clear root cause194. Initial efforts to understand long COVID have found evidence for localized and sustained inflammation in the olfactory system, changes in areas of the brain and anomalies in the blood168,195,196. When modelled in small animals, comparable observations further suggest that these events can result in changes to sensory perception, neural biology, renal function and cardiovascular performance169,197. Although many of these disease manifestations could be stochastic or driven by a common mechanism, at this time we still do not understand the basis of continued inflammation. Transcriptional changes observed in animal models and cadaver tissues suggest that material capable of inducing inflammation persists well beyond viral clearance169. Current leading theories relating to the source of this material include low-level virus replication, persistent defective viral genomes or even a secondary infection made possible by initial SARS-CoV-2 infection. It may be that all, or none, of these possibilities drive long COVID. Nevertheless, solving this scientific question should be a central focus of the community moving forward, as it will be essential for developing effective treatments for the many lives that continue to be impacted by SARS-CoV-2.
References
Kawai, T. & Akira, S. The roles of TLRs, RLRs and NLRs in pathogen recognition. Int. Immunol. 21, 317–337 (2009).
Ma, Z., Ni, G. & Damania, B. Innate sensing of DNA virus genomes. Annu. Rev. Virol. 5, 341–362 (2018).
Deretic, V., Saitoh, T. & Akira, S. Autophagy in infection, inflammation and immunity. Nat. Rev. Immunol. 13, 722–737 (2013).
Mogensen, T. H. Pathogen recognition and inflammatory signaling in innate immune defenses. Clin. Microbiol. Rev. 22, 240–273 (2009).
tenOever, B. R. The evolution of antiviral defense systems. Cell Host Microbe 19, 142–149 (2016).
Lloyd-Smith, J. O. et al. Epidemic dynamics at the human-animal interface. Science 326, 1362–1367 (2009).
Olivero, J. et al. Recent loss of closed forests is associated with Ebola virus disease outbreaks. Sci. Rep. 7, 14291 (2017).
Pernet, O. et al. Evidence for henipavirus spillover into human populations in Africa. Nat. Commun. 5, 5342 (2014).
Wolfe, N. D., Daszak, P., Kilpatrick, A. M. & Burke, D. S. Bushmeat hunting, deforestation, and prediction of zoonoses emergence. Emerg. Infect. Dis. 11, 1822–1827 (2005).
Sharp, P. M. & Hahn, B. H. Origins of HIV and the AIDS pandemic. Cold Spring Harb. Perspect. Med. 1, a006841 (2011).
Naguib, M. M. et al. Live and wet markets: food access versus the risk of disease emergence. Trends Microbiol. 29, 573–581 (2021).
Cui, J., Li, F. & Shi, Z. L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020). The first report of a complete viral genome of SARS-CoV-2 using next-generation sequencing, identifying it as a close relative of SARS-like coronaviruses.
Dong, E., Du, H. & Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect. Dis. 20, 533–534 (2020).
Rambaut, A. et al. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 5, 1403–1407 (2020).
Sette, A. & Crotty, S. Immunological memory to SARS-CoV-2 infection and COVID-19 vaccines. Immunol. Rev. https://doi.org/10.1111/imr.13089 (2022).
Moss, P. The T cell immune response against SARS-CoV-2. Nat. Immunol. 23, 186–193 (2022).
Arya, R. et al. Structural insights into SARS-CoV-2 proteins. J. Mol. Biol. 433, 166725 (2021).
Perlman, S. & Masters, P. S. in Fields Virology: Emerging Viruses Vol. 1 (eds Howley, P. M. & Knipe, D. M.) (Lippincott Williams & Wilkins, 2021).
Kasuga, Y., Zhu, B., Jang, K. J. & Yoo, J. S. Innate immune sensing of coronavirus and viral evasion strategies. Exp. Mol. Med. 53, 723–736 (2021).
Diamond, M. S. & Kanneganti, T. D. Innate immunity: the first line of defense against SARS-CoV-2. Nat. Immunol. 23, 165–176 (2022).
Thoresen, D. et al. The molecular mechanism of RIG-I activation and signaling. Immunol. Rev. 304, 154–168 (2021).
Lester, S. N. & Li, K. Toll-like receptors in antiviral innate immunity. J. Mol. Biol. 426, 1246–1264 (2014).
Bortolotti, D. et al. TLR3 and TLR7 RNA sensor activation during SARS-CoV-2 infection. Microorganisms https://doi.org/10.3390/microorganisms9091820 (2021).
Salvi, V. et al. SARS-CoV-2-associated ssRNAs activate inflammation and immunity via TLR7/8. JCI Insight https://doi.org/10.1172/jci.insight.150542 (2021).
Jung, H. E. & Lee, H. K. Current understanding of the innate control of toll-like receptors in response to SARS-CoV-2 infection. Viruses https://doi.org/10.3390/v13112132 (2021).
Yin, X. et al. MDA5 governs the innate immune response to SARS-CoV-2 in lung epithelial cells. Cell Rep. 34, 108628 (2021). A definitive study using both knock-out cells and validation assays with short interfering RNA to demonstrate that MDA5, and not RIG-I, is responsible for sensing SARS-CoV-2-derived PAMPs.
Wickenhagen, A. et al. A prenylated dsRNA sensor protects against severe COVID-19. Science 374, eabj3624 (2021).
Rodrigues, T. S. et al. Inflammasomes are activated in response to SARS-CoV-2 infection and are associated with COVID-19 severity in patients. J. Exp. Med. https://doi.org/10.1084/jem.20201707 (2021). The first study implicating inflammasome activation in disease severity and poor clinical outcomes in response to SARS-CoV-2.
Campbell, G. R., To, R. K., Hanna, J. & Spector, S. A. SARS-CoV-2, SARS-CoV-1, and HIV-1 derived ssRNA sequences activate the NLRP3 inflammasome in human macrophages through a non-classical pathway. iScience 24, 102295 (2021).
Onomoto, K., Onoguchi, K. & Yoneyama, M. Regulation of RIG-I-like receptor-mediated signaling: interaction between host and viral factors. Cell Mol. Immunol. 18, 539–555 (2021).
Fitzgerald, K. A. & Kagan, J. C. Toll-like receptors and the control of immunity. Cell 180, 1044–1066 (2020).
Panne, D., Maniatis, T. & Harrison, S. C. An atomic model of the interferon-β enhanceosome. Cell 129, 1111–1123 (2007).
Lazear, H. M., Schoggins, J. W. & Diamond, M. S. Shared and distinct functions of type I and type III interferons. Immunity 50, 907–923 (2019).
Lin, R., Génin, P., Mamane, Y. & Hiscott, J. Selective DNA binding and association with the CREB binding protein coactivator contribute to differential activation of alpha/beta interferon genes by interferon regulatory factors 3 and 7. Mol. Cell Biol. 20, 6342–6353 (2000).
Morin, P. et al. Preferential binding sites for interferon regulatory factors 3 and 7 involved in interferon-A gene transcription. J. Mol. Biol. 316, 1009–1022 (2002).
Marie, I., Durbin, J. E. & Levy, D. E. Differential viral induction of distinct interferon-α genes by positive feedback through interferon regulatory factor-7. EMBO J. 17, 6660–6669 (1998).
Platanitis, E. et al. A molecular switch from STAT2-IRF9 to ISGF3 underlies interferon-induced gene transcription. Nat. Commun. 10, 2921 (2019).
Cervantes-Barragan, L. et al. Type I IFN-mediated protection of macrophages and dendritic cells secures control of murine coronavirus infection. J. Immunol. 182, 1099–1106 (2009).
Ireland, D. D., Stohlman, S. A., Hinton, D. R., Atkinson, R. & Bergmann, C. C. Type I interferons are essential in controlling neurotropic coronavirus infection irrespective of functional CD8 T cells. J. Virol. 82, 300–310 (2008).
Boudewijns, R. et al. STAT2 signaling restricts viral dissemination but drives severe pneumonia in SARS-CoV-2 infected hamsters. Nat. Commun. 11, 5838 (2020). The first report demonstrating that disruption of IFN-I signalling, but not of IFN-III signalling, results in higher levels of SARS-CoV-2 RNA in distal organs.
Zhang, Q. et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science https://doi.org/10.1126/science.abd4570 (2020).
Pairo-Castineira, E. et al. Genetic mechanisms of critical illness in COVID-19. Nature 591, 92–98 (2021).
Bastard, P. et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science https://doi.org/10.1126/science.abd4585 (2020). The first report showing that autoantibodies against IFN-I may account for severe clinical outcomes in as many as 5% of hospitalized patients.
Manry, J. et al. The risk of COVID-19 death is much greater and age dependent with type I IFN autoantibodies. Proc. Natl Acad. Sci. USA 119, e2200413119 (2022).
Sposito, B. et al. The interferon landscape along the respiratory tract impacts the severity of COVID-19. Cell 184, 4953–4968.e16 (2021).
Cheemarla, N. R. et al. Dynamic innate immune response determines susceptibility to SARS-CoV-2 infection and early replication kinetics. J. Exp. Med. https://doi.org/10.1084/jem.20210583 (2021). A unique study that showcases the relationship between timing of IFN-I engagement and the extent of SARS-CoV-2 replication.
Loske, J. et al. Pre-activated antiviral innate immunity in the upper airways controls early SARS-CoV-2 infection in children. Nat. Biotechnol. 40, 319–324 (2022).
Daniloski, Z. et al. Identification of required host factors for SARS-CoV-2 infection in human cells. Cell 184, 92–105.e16 (2021).
Prydz, K. & Saraste, J. The life cycle and enigmatic egress of coronaviruses. Mol. Microbiol. https://doi.org/10.1111/mmi.14907 (2022).
Jackson, C. B., Farzan, M., Chen, B. & Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 23, 3–20 (2022).
Yan, L. et al. Cryo-EM structure of an extended SARS-CoV-2 replication and transcription complex reveals an intermediate state in cap synthesis. Cell 184, 184–193.e10 (2021). First report on the mechanism by which the SARS-CoV-2 replicase achieves capping of its genomic and subgenomic RNAs.
Ivanov, K. A. et al. Multiple enzymatic activities associated with severe acute respiratory syndrome coronavirus helicase. J. Virol. 78, 5619–5632 (2004).
Chen, Y. et al. Functional screen reveals SARS coronavirus nonstructural protein nsp14 as a novel cap N7 methyltransferase. Proc. Natl Acad. Sci. USA 106, 3484–3489 (2009).
Snijder, E. J. et al. Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage. J. Mol. Biol. 331, 991–1004 (2003).
Decroly, E. et al. Coronavirus nonstructural protein 16 is a cap-0 binding enzyme possessing (nucleoside-2’O)-methyltransferase activity. J. Virol. 82, 8071–8084 (2008).
Walker, A. P. et al. The SARS-CoV-2 RNA polymerase is a viral RNA capping enzyme. Nucleic Acids Res. 49, 13019–13030 (2021).
Wilamowski, M. et al. 2’-O methylation of RNA cap in SARS-CoV-2 captured by serial crystallography. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2100170118 (2021).
Krafcikova, P., Silhan, J., Nencka, R. & Boura, E. Structural analysis of the SARS-CoV-2 methyltransferase complex involved in RNA cap creation bound to sinefungin. Nat. Commun. 11, 3717 (2020).
Yan, L. et al. Coupling of N7-methyltransferase and 3′-5′ exoribonuclease with SARS-CoV-2 polymerase reveals mechanisms for capping and proofreading. Cell 184, 3474–3485.e11 (2021).
Silva, J. R. A. et al. Exploring the catalytic mechanism of the RNA cap modification by nsp16-nsp10 complex of SARS-CoV-2 through a QM/MM approach. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23010300 (2021).
Frazier, M. N. et al. Characterization of SARS2 Nsp15 nuclease activity reveals it’s mad about U. Nucleic Acids Res. 49, 10136–10149 (2021).
Hackbart, M., Deng, X. & Baker, S. C. Coronavirus endoribonuclease targets viral polyuridine sequences to evade activating host sensors. Proc. Natl Acad. Sci. USA 117, 8094–8103 (2020).
Versteeg, G. A., Bredenbeek, P. J., van den Worm, S. H. & Spaan, W. J. Group 2 coronaviruses prevent immediate early interferon induction by protection of viral RNA from host cell recognition. Virology 361, 18–26 (2007).
Zhou, H. & Perlman, S. Mouse hepatitis virus does not induce beta interferon synthesis and does not inhibit its induction by double-stranded RNA. J. Virol. 81, 568–574 (2007).
Angelini, M. M., Akhlaghpour, M., Neuman, B. W. & Buchmeier, M. J. Severe acute respiratory syndrome coronavirus nonstructural proteins 3, 4, and 6 induce double-membrane vesicles. mBio https://doi.org/10.1128/mBio.00524-13 (2013).
Oudshoorn, D. et al. Expression and cleavage of Middle East respiratory syndrome coronavirus nsp3-4 polyprotein induce the formation of double-membrane vesicles that mimic those associated with coronaviral RNA replication. mBio https://doi.org/10.1128/mBio.01658-17 (2017).
Zerbib, Y. et al. Pathology assessments of multiple organs in fatal COVID-19 in intensive care unit vs. non-intensive care unit patients. Front. Med. 9, 837258 (2022).
Ricciardi, S. et al. The role of NSP6 in the biogenesis of the SARS-CoV-2 replication organelle. Nature https://doi.org/10.1038/s41586-022-04835-6 (2022). One of the first studies demonstrating the roles of Nsp6 in organizing and providing resources for double-membrane vesicle clusters during SARS-CoV-2 replication.
Levine, B., Mizushima, N. & Virgin, H. W. Autophagy in immunity and inflammation. Nature 469, 323–335 (2011).
Chen, D. & Zhang, H. Autophagy in SARS-CoV-2 infection. Curr. Opin. Physiol. https://doi.org/10.1016/j.cophys.2022.100596 (2022).
Guo, K. et al. Interferon resistance of emerging SARS-CoV-2 variants. Proc. Natl Acad. Sci. USA 119, e2203760119 (2022).
Wolff, G. et al. A molecular pore spans the double membrane of the coronavirus replication organelle. Science 369, 1395–1398 (2020).
Finkel, Y. et al. The coding capacity of SARS-CoV-2. Nature 589, 125–130 (2021). One of the earliest reports using ribosomal profiling to identify a high-resolution map of coding regions within the SARS-CoV-2 genome.
Min, Y. Q. et al. Immune evasion of SARS-CoV-2 from interferon antiviral system. Comput. Struct. Biotechnol. J. 19, 4217–4225 (2021).
Caruso, I. P., Sanches, K., Da Poian, A. T., Pinheiro, A. S. & Almeida, F. C. L. Dynamics of the SARS-CoV-2 nucleoprotein N-terminal domain triggers RNA duplex destabilization. Biophys. J. 120, 2814–2827 (2021).
Lu, S. et al. The SARS-CoV-2 nucleocapsid phosphoprotein forms mutually exclusive condensates with RNA and the membrane-associated M protein. Nat. Commun. 12, 502 (2021).
Cubuk, J. et al. The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA. Nat. Commun. 12, 1936 (2021).
Thorne, L. G. et al. Evolution of enhanced innate immune evasion by SARS-CoV-2. Nature 602, 487–495 (2022). A study leveraging genomic and proteomic techniques that revealed the importance of mutations outside the spike coding region in the adaptation of SARS-CoV-2 VOCs.
Parker, M. D. et al. Altered subgenomic RNA abundance provides unique insight into SARS-CoV-2 B.1.1.7/Alpha variant infections. Commun. Biol. 5, 666 (2022).
Chu, H. et al. Coronaviruses exploit a host cysteine-aspartic protease for replication. Nature 609, 785–792 (2022).
Zheng, Y. et al. SARS-CoV-2 NSP5 and N protein counteract the RIG-I signaling pathway by suppressing the formation of stress granules. Signal. Transduct. Target. Ther. 7, 22 (2022).
Chen, K. et al. SARS-CoV-2 nucleocapsid protein interacts with RIG-I and represses RIG-mediated IFN-betaβ production. Viruses https://doi.org/10.3390/v13010047 (2020).
Oh, S. J. & Shin, O. S. SARS-CoV-2 nucleocapsid protein targets RIG-I-like receptor pathways to inhibit the induction of interferon response. Cells https://doi.org/10.3390/cells10030530 (2021).
Gori Savellini, G., Anichini, G., Gandolfo, C. & Cusi, M. G. SARS-CoV-2 N protein targets TRIM25-mediated RIG-I activation to suppress innate immunity. Viruses https://doi.org/10.3390/v13081439 (2021).
Lai, M. M. Coronavirus: organization, replication and expression of genome. Annu. Rev. Microbiol. 44, 303–333 (1990).
Cadena, C. et al. Ubiquitin-dependent and -independent roles of E3 ligase RIPLET in innate immunity. Cell 177, 1187–1200.e16 (2019).
Liu, G. et al. ISG15-dependent activation of the sensor MDA5 is antagonized by the SARS-CoV-2 papain-like protease to evade host innate immunity. Nat. Microbiol. 6, 467–478 (2021).
Dzimianski, J. V., Scholte, F. E. M., Bergeron, E. & Pegan, S. D. ISG15: it’s complicated. J. Mol. Biol. 431, 4203–4216 (2019).
Klemm, T. et al. Mechanism and inhibition of the papain-like protease, PLpro, of SARS-CoV-2. EMBO J. 39, e106275 (2020).
Shin, D. et al. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 587, 657–662 (2020).
Alhammad, Y. M. O. et al. The SARS-CoV-2 conserved macrodomain is a mono-ADP-ribosylhydrolase. J. Virol. https://doi.org/10.1128/JVI.01969-20 (2021).
Frick, D. N., Virdi, R. S., Vuksanovic, N., Dahal, N. & Silvaggi, N. R. Molecular basis for ADP-ribose binding to the Mac1 domain of SARS-CoV-2 nsp3. Biochemistry 59, 2608–2615 (2020).
Michalska, K. et al. Crystal structures of SARS-CoV-2 ADP-ribose phosphatase: from the apo form to ligand complexes. IUCrJ 7, 814–824 (2020).
Mielech, A. M., Chen, Y., Mesecar, A. D. & Baker, S. C. Nidovirus papain-like proteases: multifunctional enzymes with protease, deubiquitinating and deISGylating activities. Virus Res. 194, 184–190 (2014).
Liu, Y. et al. SARS-CoV-2 Nsp5 demonstrates two distinct mechanisms targeting RIG-I and MAVS to evade the innate immune response. mBio 12, e0233521 (2021).
Kouwaki, T., Nishimura, T., Wang, G. & Oshiumi, H. RIG-I-like receptor-mediated recognition of viral genomic RNA of severe acute respiratory syndrome coronavirus-2 and viral escape from the host innate immune responses. Front. Immunol. 12, 700926 (2021).
Shemesh, M. et al. SARS-CoV-2 suppresses IFNβ production mediated by NSP1, 5, 6, 15, ORF6 and ORF7b but does not suppress the effects of added interferon. PLoS Pathog. 17, e1009800 (2021).
Fu, Y. Z. et al. SARS-CoV-2 membrane glycoprotein M antagonizes the MAVS-mediated innate antiviral response. Cell Mol. Immunol. 18, 613–620 (2021).
Wang, S. et al. Targeting liquid–liquid phase separation of SARS-CoV-2 nucleocapsid protein promotes innate antiviral immunity by elevating MAVS activity. Nat. Cell Biol. 23, 718–732 (2021).
Savastano, A., Ibanez de Opakua, A., Rankovic, M. & Zweckstetter, M. Nucleocapsid protein of SARS-CoV-2 phase separates into RNA-rich polymerase-containing condensates. Nat. Commun. 11, 6041 (2020). First report that SARS-CoV-2 N protein can induce liquid–liquid phase separation with its cognate RNA, aiding in viral evasion of innate immunity.
Wu, Y. et al. RNA-induced liquid phase separation of SARS-CoV-2 nucleocapsid protein facilitates NF-κB hyper-activation and inflammation. Signal. Transduct. Target. Ther. 6, 167 (2021).
Iserman, C. et al. Genomic RNA elements drive phase separation of the SARS-CoV-2 nucleocapsid. Mol. Cell 80, 1078–1091.e6 (2020).
Sui, L. et al. SARS-CoV-2 membrane protein inhibits type I interferon production through ubiquitin-mediated degradation of TBK1. Front. Immunol. 12, 662989 (2021).
Xia, H. et al. Evasion of type I interferon by SARS-CoV-2. Cell Rep. 33, 108234 (2020).
Vazquez, C. et al. SARS-CoV-2 viral proteins NSP1 and NSP13 inhibit interferon activation through distinct mechanisms. PLoS ONE 16, e0253089 (2021).
Han, L. et al. SARS-CoV-2 ORF9b antagonizes type I and III interferons by targeting multiple components of the RIG-I/MDA-5-MAVS, TLR3-TRIF, and cGAS-STING signaling pathways. J. Med. Virol. 93, 5376–5389 (2021).
Sharma, S. et al. Triggering the interferon antiviral response through an IKK-related pathway. Science 300, 1148–1151 (2003).
Brandherm, L. et al. Phosphorylation of SARS-CoV-2 Orf9b regulates its targeting to two binding sites in TOM70 and recruitment of Hsp90. Int. J. Mol. Sci. https://doi.org/10.3390/ijms22179233 (2021).
Jiang, H. W. et al. SARS-CoV-2 Orf9b suppresses type I interferon responses by targeting TOM70. Cell Mol. Immunol. 17, 998–1000 (2020).
Gao, X. et al. Crystal structure of SARS-CoV-2 Orf9b in complex with human TOM70 suggests unusual virus-host interactions. Nat. Commun. 12, 2843 (2021).
Nilsson-Payant, B. E. et al. The NF-κB transcriptional footprint is essential for SARS-CoV-2 replication. J. Virol. 95, e01257-21 (2021). A definitive study showcasing the dependency of SARS-CoV-2 replication on NF-κB signalling using scRNA-seq and ATAC-seq.
Freitas, R. S., Crum, T. F. & Parvatiyar, K. SARS-CoV-2 spike antagonizes innate antiviral immunity by targeting interferon regulatory factor 3. Front. Cell Infect. Microbiol. 11, 789462 (2021).
Fung, S. Y., Siu, K. L., Lin, H., Yeung, M. L. & Jin, D. Y. SARS-CoV-2 main protease suppresses type I interferon production by preventing nuclear translocation of phosphorylated IRF3. Int. J. Biol. Sci. 17, 1547–1554 (2021).
Zhang, W. et al. SARS-CoV-2 3C-like protease antagonizes interferon-beta production by facilitating the degradation of IRF3. Cytokine 148, 155697 (2021).
Moustaqil, M. et al. SARS-CoV-2 proteases PLpro and 3CLpro cleave IRF3 and critical modulators of inflammatory pathways (NLRP12 and TAB1): implications for disease presentation across species. Emerg. Microbes Infect. 10, 178–195 (2021).
Chen, J. et al. SARS-CoV-2 nsp5 exhibits stronger catalytic activity and interferon antagonism than Its SARS-CoV ortholog. J. Virol. 96, e0003722 (2022).
Wang, W. et al. SARS-CoV-2 nsp12 attenuates type I interferon production by inhibiting IRF3 nuclear translocation. Cell Mol. Immunol. 18, 945–953 (2021).
Zhang, Q. et al. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) membrane (M) and spike (S) proteins antagonize host type I interferon response. Front. Cell Infect. Microbiol. 11, 766922 (2021).
Konno, Y. et al. SARS-CoV-2 ORF3b is a potent interferon antagonist whose activity is increased by a naturally occurring elongation variant. Cell Rep. 32, 108185 (2020).
Kumar, A. et al. SARS-CoV-2 nonstructural protein 1 inhibits the interferon response by causing depletion of key host signaling factors. J. Virol. 95, e0026621 (2021).
Fiege, J. K. et al. Single cell resolution of SARS-CoV-2 tropism, antiviral responses, and susceptibility to therapies in primary human airway epithelium. PLoS Pathog. 17, e1009292 (2021). The first report of scRNA-seq data in response to SARS-CoV-2 infection demonstrating a lack of IFN-I induction in infected cells.
Kamitani, W., Huang, C., Narayanan, K., Lokugamage, K. G. & Makino, S. A two-pronged strategy to suppress host protein synthesis by SARS coronavirus Nsp1 protein. Nat. Struct. Mol. Biol. 16, 1134–1140 (2009).
Hsu, J. C., Laurent-Rolle, M., Pawlak, J. B., Wilen, C. B. & Cresswell, P. Translational shutdown and evasion of the innate immune response by SARS-CoV-2 NSP14 protein. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2101161118 (2021).
Zaffagni, M. et al. SARS-CoV-2 Nsp14 mediates the effects of viral infection on the host cell transcriptome. eLife https://doi.org/10.7554/eLife.71945 (2022).
Bader, S. M., Cooney, J. P., Pellegrini, M. & Doerflinger, M. Programmed cell death: the pathways to severe COVID-19? Biochem. J. 479, 609–628 (2022).
Sariol, A. & Perlman, S. SARS-CoV-2 takes its Toll. Nat. Immunol. 22, 801–802 (2021).
Lee, J. S. et al. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci. Immunol. https://doi.org/10.1126/sciimmunol.abd1554 (2020).
Zhou, Z. et al. Heightened innate immune responses in the respiratory tract of COVID-19 patients. Cell Host Microbe 27, 883–890.e2 (2020).
Kim, Y. M. & Shin, E. C. Type I and III interferon responses in SARS-CoV-2 infection. Exp. Mol. Med. 53, 750–760 (2021).
Hayn, M. et al. Systematic functional analysis of SARS-CoV-2 proteins uncovers viral innate immune antagonists and remaining vulnerabilities. Cell Rep. 35, 109126 (2021).
Fung, S. Y. et al. SARS-CoV-2 NSP13 helicase suppresses interferon signaling by perturbing JAK1 phosphorylation of STAT1. Cell Biosci. 12, 36 (2022).
Lokugamage, K. G. et al. Type I interferon susceptibility distinguishes SARS-CoV-2 from SARS-CoV. J. Virol. https://doi.org/10.1128/JVI.01410-20 (2020). The first report comparing the impact of the host antiviral response on SARS-CoV versus SARS-CoV-2, demonstrating SARS-CoV-2 to be significantly more sensitive to IFN-I-mediated inhibition.
Cautain, B., Hill, R., de Pedro, N. & Link, W. Components and regulation of nuclear transport processes. FEBS J. 282, 445–462 (2015).
Jiang, D. Building the nuclear pore complex. Science 376, 1172–1173 (2022).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020). A landmark paper reporting on protein–protein interactions of the majority of the SARS-CoV-2 proteome and the use of this information to identify possible therapeutic interventions.
Zhang, K. et al. Nsp1 protein of SARS-CoV-2 disrupts the mRNA export machinery to inhibit host gene expression. Sci. Adv. https://doi.org/10.1126/sciadv.abe7386 (2021).
Makiyama, K. et al. NSP9 of SARS-CoV-2 attenuates nuclear transport by hampering nucleoporin 62 dynamics and functions in host cells. Biochem. Biophys. Res. Commun. 586, 137–142 (2022).
Kato, K. et al. Overexpression of SARS-CoV-2 protein ORF6 dislocates RAE1 and NUP98 from the nuclear pore complex. Biochem. Biophys. Res. Commun. 536, 59–66 (2021).
Addetia, A. et al. SARS-CoV-2 ORF6 disrupts bidirectional nucleocytoplasmic transport through interactions with Rae1 and Nup98. mBio https://doi.org/10.1128/mBio.00065-21 (2021).
Miorin, L. et al. SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proc. Natl Acad. Sci. USA 117, 28344–28354 (2020).
Kimura, I. et al. Sarbecovirus ORF6 proteins hamper induction of interferon signaling. Cell Rep. 34, 108916 (2021).
Li, T. et al. Molecular mechanism of SARS-CoVs Orf6 targeting the Rae1-Nup98 complex to compete with mRNA nuclear export. Front. Mol. Biosci. 8, 813248 (2021).
Gori Savellini, G., Anichini, G., Gandolfo, C. & Cusi, M. G. Nucleopore traffic is hindered by SARS-CoV-2 ORF6 protein to efficiently suppress IFN-β and IL-6 secretion. Viruses https://doi.org/10.3390/v14061273 (2022).
Riojas, M. A. et al. A rare deletion in SARS-CoV-2 ORF6 dramatically alters the predicted three-dimensional structure of the resultant protein. Preprint at bioRxiv https://doi.org/10.1101/2020.06.09.134460 (2020).
Stern-Ginossar, N., Thompson, S. R., Mathews, M. B. & Mohr, I. Translational control in virus-infected cells. Cold Spring Harb. Perspect. Biol. https://doi.org/10.1101/cshperspect.a033001 (2019).
Joshi, S., Kaur, S., Kroczynska, B. & Platanias, L. C. Mechanisms of mRNA translation of interferon stimulated genes. Cytokine 52, 123–127 (2010).
Nakagawa, K., Lokugamage, K. G. & Makino, S. Viral and cellular mRNA translation in coronavirus-infected cells. Adv. Virus Res. 96, 165–192 (2016).
Banerjee, A. K. et al. SARS-CoV-2 disrupts splicing, translation, and protein trafficking to suppress host defenses. Cell 183, 1325–1339.e21 (2020). First report that characterizes the interactions of SARS-CoV-2 proteins with host components of the spliceosome, the ribosome and the SRP.
Schubert, K. et al. SARS-CoV-2 Nsp1 binds the ribosomal mRNA channel to inhibit translation. Nat. Struct. Mol. Biol. 27, 959–966 (2020).
Thoms, M. et al. Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2. Science 369, 1249–1255 (2020). A comprehensive report demonstrating how the SARS-CoV-2 Nsp2 protein blocks the mRNA entry tunnel of the 40S ribosomal subunit to block host translation.
Yuan, S. et al. Nonstructural protein 1 of SARS-CoV-2 is a potent pathogenicity factor redirecting host protein synthesis machinery toward viral RNA. Mol. Cell 80, 1055–1066.e6 (2020).
Lapointe, C. P. et al. Dynamic competition between SARS-CoV-2 NSP1 and mRNA on the human ribosome inhibits translation initiation. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2017715118 (2021).
Tidu, A. et al. The viral protein NSP1 acts as a ribosome gatekeeper for shutting down host translation and fostering SARS-CoV-2 translation. RNA https://doi.org/10.1261/rna.078121.120 (2020).
Finkel, Y. et al. SARS-CoV-2 uses a multipronged strategy to impede host protein synthesis. Nature 594, 240–245 (2021).
Matera, A. G. & Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 15, 108–121 (2014).
Wang, C. et al. Abnormal global alternative RNA splicing in COVID-19 patients. PLoS Genet. 18, e1010137 (2022).
Akopian, D., Shen, K., Zhang, X. & Shan, S. O. Signal recognition particle: an essential protein-targeting machine. Annu. Rev. Biochem. 82, 693–721 (2013).
Bar-On, Y. M., Flamholz, A., Phillips, R. & Milo, R. SARS-CoV-2 (COVID-19) by the numbers. eLife https://doi.org/10.7554/eLife.57309 (2020).
Guan, W. J. et al. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 382, 1708–1720 (2020).
Gupta, A. et al. Extrapulmonary manifestations of COVID-19. Nat. Med. 26, 1017–1032 (2020).
Mehandru, S. & Merad, M. Pathological sequelae of long-haul COVID. Nat. Immunol. 23, 194–202 (2022).
Goyal, P. et al. Clinical characteristics of Covid-19 in New York city. N. Engl. J. Med. 382, 2372–2374 (2020).
Imai, M. et al. Syrian hamsters as a small animal model for SARS-CoV-2 infection and countermeasure development. Proc. Natl Acad. Sci. USA 117, 16587–16595 (2020). One of the earliest and most comprehensive studies demonstrating the value in using the golden hamster as the small-animal model for studying SARS-CoV-2 biology.
Hoagland, D. A. et al. Leveraging the antiviral type I interferon system as a first line of defense against SARS-CoV-2 pathogenicity. Immunity 54, 557–570.e5 (2021).
Alon, R. et al. Leukocyte trafficking to the lungs and beyond: lessons from influenza for COVID-19. Nat. Rev. Immunol. 21, 49–64 (2021).
Sellers, S. A., Hagan, R. S., Hayden, F. G. & Fischer, W. A. II The hidden burden of influenza: a review of the extra-pulmonary complications of influenza infection. Influenza Other Respir. Viruses 11, 372–393 (2017).
Fernández-Castañeda, A. et al. Mild respiratory COVID can cause multi-lineage neural cell and myelin dysregulation. Cell 185, 2452–2468.e16 (2022). One of the first reports characterizing the neurobiological effects of SARS-CoV-2 infection in mice and humans, showing persistent neuropathological changes and dysregulation compared with other respiratory infections, and identifying a possible mechanism for some complications observed in patients with long COVID.
Frere, J. J. et al. SARS-CoV-2 infection in hamsters and humans results in lasting and unique systemic perturbations post recovery. Sci. Transl Med. https://doi.org/10.1126/scitranslmed.abq3059 (2022). One of the first reports to investigate the long-term impact of SARS-CoV-2 infection in the golden hamster, showing that persistent interferon signatures in the brain are unique to SARS-CoV-2 and corroborate these findings in long COVID cadaver tissue.
Zazhytska, M. et al. Non-cell-autonomous disruption of nuclear architecture as a potential cause of COVID-19-induced anosmia. Cell 185, 1052–1064.e12 (2022). A definitive report detailing the molecular basis for anosmia, and demonstrating that infection of sustentacular cells results in the death and activation of microglia, leading to interferon signalling and disruption of olfactory neuron function.
de Melo, G. D. et al. COVID-19-related anosmia is associated with viral persistence and inflammation in human olfactory epithelium and brain infection in hamsters. Sci. Transl Med. https://doi.org/10.1126/scitranslmed.abf8396 (2021).
Baggen, J. et al. Genome-wide CRISPR screening identifies TMEM106B as a proviral host factor for SARS-CoV-2. Nat. Genet. 53, 435–444 (2021).
Wei, J. et al. Genome-wide CRISPR screens reveal host factors critical for SARS-CoV-2 infection. Cell 184, 76–91.e13 (2021).
RECOVERY Collaborative Group et al. Dexamethasone in hospitalized patients with Covid-19. N. Engl. J. Med. 384, 693–704 (2021). Report describing the first therapeutic breakthrough against COVID-19, showing that dexamethasone, a low-cost steroid treatment, can reduce deaths of hospitalized COVID-19 patients with severe respiratory complications by up to one-third.
WHO Rapid Evidence Appraisal for COVID-19 Therapies (REACT) Working Group et al. Association between administration of systemic corticosteroids and mortality among critically ill patients with COVID-19: a meta-analysis. JAMA 324, 1330–1341 (2020).
COVID-19 Treatment Guidelines Panel. Coronavirus Disease 2019 (COVID-19) Treatment Guidelines. NIH https://www.covid19treatmentguidelines.nih.gov/ (2022).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045.e9 (2020). The first comprehensive transcriptional analysis of SARS-CoV-2 infection in cells, small-animal models and patient samples, demonstrating an aberrant induction of proinflammatory cytokines with a delayed interferon response.
Mehta, P. et al. COVID-19: consider cytokine storm syndromes and immunosuppression. Lancet 395, 1033–1034 (2020).
Santa Cruz, A. et al. Interleukin-6 is a biomarker for the development of fatal severe acute respiratory syndrome coronavirus 2 pneumonia. Front. Immunol. 12, 613422 (2021).
Galván-Román, J. M. et al. IL-6 serum levels predict severity and response to tocilizumab in COVID-19: an observational study. J. Allergy Clin. Immunol. 147, 72–80.e8 (2021).
Potere, N. et al. The role of IL-6 and IL-6 blockade in COVID-19. Expert Rev. Clin. Immunol. 17, 601–618 (2021).
Kyriazopoulou, E. et al. Early treatment of COVID-19 with anakinra guided by soluble urokinase plasminogen receptor plasma levels: a double-blind, randomized controlled phase 3 trial. Nat. Med. 27, 1752–1760 (2021).
Kow, C. S. & Hasan, S. S. The effect of tocilizumab on mortality in hospitalized patients with COVID-19: a meta-analysis of randomized controlled trials. Eur. J. Clin. Pharmacol. 77, 1089–1094 (2021).
RECOVERY Collaborative Group. Tocilizumab in patients admitted to hospital with COVID-19 (RECOVERY): a randomised, controlled, open-label, platform trial. Lancet 397, 1637–1645 (2021).
Salama, C. et al. Tocilizumab in patients hospitalized with Covid-19 pneumonia. N. Engl. J. Med. 384, 20–30 (2021).
Caricchio, R. et al. Effect of canakinumab vs placebo on survival without invasive mechanical ventilation in patients hospitalized with severe COVID-19: a randomized clinical trial. JAMA 326, 230–239 (2021).
CORIMUNO-19 Collaborative Group. Effect of anakinra versus usual care in adults in hospital with COVID-19 and mild-to-moderate pneumonia (CORIMUNO-ANA-1): a randomised controlled trial. Lancet Respir. Med. 9, 295–304 (2021).
REMAP-CAP Investigators et al. Interleukin-6 receptor antagonists in critically ill patients with Covid-19. N. Engl. J. Med. 384, 1491–1502 (2021).
US National Library of Medicine. ClinicalTrials.gov, https://clinicaltrials.gov/ct2/show/NCT05279391 (2022).
Marconi, V. C. et al. Efficacy and safety of baricitinib for the treatment of hospitalised adults with COVID-19 (COV-BARRIER): a randomised, double-blind, parallel-group, placebo-controlled phase 3 trial. Lancet Respir. Med. 9, 1407–1418 (2021).
Kalil, A. C. et al. Baricitinib plus remdesivir for hospitalized adults with Covid-19. N. Engl. J. Med. 384, 795–807 (2021).
Guimaraes, P. O. et al. Tofacitinib in patients hospitalized with Covid-19 pneumonia. N. Engl. J. Med. 385, 406–415 (2021).
Frere, J. J. & tenOever, B. R. Cardiometabolic syndrome — an emergent feature of Long COVID? Nat. Rev. Immunol. 22, 399–400 (2022).
Couzin-Frankel, J. Clues to long COVID. Science 376, 1261–1265 (2022).
Couzin-Frankel, J. Long Covid clues emerge from patients’ blood. Science 377, 803 (2022).
Douaud, G. et al. SARS-CoV-2 is associated with changes in brain structure in UK Biobank. Nature 604, 697–707 (2022). The first clear evidence that SARS-CoV-2 infection can result in lasting in vivo hallmarks of a degenerative spread of the disease through olfactory pathways.
Serafini, R. A. et al. SARS-CoV-2 airway infection results in time-dependent sensory abnormalities in a hamster model. Preprint at bioRxiv https://doi.org/10.1101/2022.08.19.504551 (2022).
Li, T. et al. SARS-CoV-2 Nsp14 activates NF-κB signaling and induces IL-8 upregulation. Preprint at bioRxiv https://doi.org/10.1101/2021.05.26.445787 (2021).
Wu, X. et al. Viral mimicry of interleukin-17A by SARS-CoV-2 ORF8. mBio 13, e0040222 (2022).
Olajide, O. A., Iwuanyanwu, V. U., Adegbola, O. D. & Al-Hindawi, A. A. SARS-CoV-2 spike glycoprotein S1 induces neuroinflammation in BV-2 microglia. Mol. Neurobiol. 59, 445–458 (2022).
Waters, C. N. et al. The Anthropocene is functionally and stratigraphically distinct from the Holocene. Science 351, aad2622 (2016).
Piret, J. & Boivin, G. Pandemics throughout history. Front. Microbiol. 11, 631736 (2020).
Liu, T., Zhang, L., Joo, D. & Sun, S. C. NF-κB signaling in inflammation. Signal Transduct. Target. Ther. https://doi.org/10.1038/sigtrans.2017.23 (2017).
Acknowledgements
The authors would like to thank the Zegar Family Foundation for supporting the SARS-CoV-2 research programme within the tenOever lab.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Microbiology thanks Michaela Gack and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Clinicaltrials.gov: https://clinicaltrials.gov/
NIH COVID-19 Treatment Guidelines: https://www.covid19treatmentguidelines.nih.gov/
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Minkoff, J.M., tenOever, B. Innate immune evasion strategies of SARS-CoV-2. Nat Rev Microbiol 21, 178–194 (2023). https://doi.org/10.1038/s41579-022-00839-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-022-00839-1
This article is cited by
-
Recent Advances in Electrochemical and Nanophotonic Biosensors for SARS-CoV-2 Detection
BioChip Journal (2024)