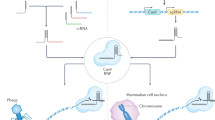
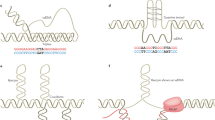
Abstract
All organisms must safeguard the integrity of their DNA to avoid deleterious consequences of genome instability, which have been linked to human diseases such as autoimmune disorders, neurodegenerative diseases and cancer. Traditionally, genome maintenance has been viewed largely in terms of DNA–protein interactions. However, emerging evidence points to RNA as a key modulator of genome stability, with seemingly opposing roles in promoting chromosomal instability and protecting genome integrity. Unravelling the mechanistic and contextual basis of this duality will not only improve our understanding of the interfaces between RNA and the genome but will also provide important insights into how disrupted RNA metabolism contributes to disease origin, laying the foundation for targeted intervention.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Jackson, S. P. & Bartek, J. The DNA-damage response in human biology and disease. Nature 461, 1071–1078 (2009).
Bunting, S. F. & Nussenzweig, A. End-joining, translocations and cancer. Nat. Rev. Cancer 13, 443–454 (2013).
McKinnon, P. J. Genome integrity and disease prevention in the nervous system. Genes. Dev. 31, 1180–1194 (2017).
Tubbs, A. & Nussenzweig, A. Endogenous DNA damage as a source of genomic instability in cancer. Cell 168, 644–656 (2017).
Bader, A. S., Hawley, B. R., Wilczynska, A. & Bushell, M. The roles of RNA in DNA double-strand break repair. Br. J. Cancer (2020).
Belotserkovskii, B. P., Tornaletti, S., D’Souza, A. D. & Hanawalt, P. C. R-loop generation during transcription: Formation, processing and cellular outcomes. DNA Repair 71, 69–81 (2018).
Huertas, P. & Aguilera, A. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12, 711–721 (2003).
Chen, X., Yang, J. R., Zhang, J. & Nascent, R. N. A. folding mitigates transcription-associated mutagenesis. Genome Res. 26, 50–59 (2016).
Ginno, P. A., Lim, Y. W., Lott, P. L., Korf, I. & Chedin, F. GC skew at the 5’ and 3’ ends of human genes links R-loop formation to epigenetic regulation and transcription termination. Genome Res. 23, 1590–1600 (2013).
Stolz, R. et al. Interplay between DNA sequence and negative superhelicity drives R-loop structures. Proc. Natl Acad. Sci. USA 116, 6260–6269 (2019).
Wahba, L., Costantino, L., Tan, F. J., Zimmer, A. & Koshland, D. S1-DRIP-seq identifies high expression and polyA tracts as major contributors to R-loop formation. Genes Dev. 30, 1327–1338 (2016).
Salas-Armenteros, I. et al. Human THO-Sin3A interaction reveals new mechanisms to prevent R-loops that cause genome instability. EMBO J. 36, 3532–3547 (2017).
Yang, X. et al. m6A promotes R-loop formation to facilitate transcription termination. Cell Res. 29, 1035–1038 (2019).
Kotsantis, P. et al. Increased global transcription activity as a mechanism of replication stress in cancer. Nat. Commun. 7, 13087 (2016).
Stork, C. T. et al. Co-transcriptional R-loops are the main cause of estrogen-induced DNA damage. Elife https://doi.org/10.7554/eLife.17548 (2016). This study reports that activation of transcription by oestrogen (oestradiol) can induce R-loop-dependent DNA damage at genomic loci that are enriched in rearrangements in breast cancer.
Hamperl, S., Bocek, M. J., Saldivar, J. C., Swigut, T. & Cimprich, K. A. Transcription-replication conflict orientation modulates R-loop levels and activates distinct DNA damage responses. Cell 170, 774–786 (2017).
Sordet, O. et al. Ataxia telangiectasia mutated activation by transcription- and topoisomerase I-induced DNA double-strand breaks. EMBO Rep. 10, 887–893 (2009).
Garcia-Benitez, F., Gaillard, H. & Aguilera, A. Physical proximity of chromatin to nuclear pores prevents harmful R loop accumulation contributing to maintain genome stability. Proc. Natl Acad. Sci. USA 114, 10942–10947 (2017).
Wahba, L., Amon, J. D., Koshland, D., Vuica-Ross, M. & RNase, H. and multiple RNA biogenesis factors cooperate to prevent RNA:DNA hybrids from generating genome instability. Mol. Cell 44, 978–988 (2011).
Nadel, J. et al. RNA:DNA hybrids in the human genome have distinctive nucleotide characteristics, chromatin composition, and transcriptional relationships. Epigenetics Chromatin 8, 46 (2015).
Wahba, L., Gore, S. K. & Koshland, D. The homologous recombination machinery modulates the formation of RNA-DNA hybrids and associated chromosome instability. Elife 2, e00505 (2013).
Chen, L. et al. R-ChIP using inactive RNase H reveals dynamic coupling of R-loops with transcriptional pausing at gene promoters. Mol. Cell 68, 745–757 (2017).
Dumelie, J. G. & Jaffrey, S. R. Defining the location of promoter-associated R-loops at near-nucleotide resolution using bisDRIP-seq. Elife https://doi.org/10.7554/eLife.28306 (2017).
Ginno, P. A., Lott, P. L., Christensen, H. C., Korf, I. & Chedin, F. R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol. Cell 45, 814–825 (2012).
Malig, M., Hartono, S. R., Giafaglione, J. M., Sanz, L. A. & Chedin, F. Ultra-deep coverage single-molecule R-loop footprinting reveals principles of R-loop formation. J. Mol. Biol. https://doi.org/10.1016/j.jmb.2020.02.014 (2020). This study demonstrates the feasibility of mapping R-loops genome-wide at high resolution by capturing and sequencing the single-stranded DNA of the displaced strand, providing additional insights into the principles underlying R-loop formation. It is a valuable complementary approach to DNA–RNA immunoprecipitation coupled with high-throughput sequencing (described in Sanz et al., 2016).
Sanz, L. A. et al. Prevalent, dynamic, and conserved r-loop structures associate with specific epigenomic signatures in mammals. Mol. Cell 63, 167–178 (2016). This study is among the first to report genome-wide mapping of R-loops in human cells using the S9.6 antibody, which specifically recognizes RNA–DNA hybrids. This approach is known as DNA–RNA immunoprecipitation coupled with high-throughput sequencing.
Yan, Q., Shields, E. J., Bonasio, R. & Sarma, K. Mapping native R-loops genome-wide using a targeted nuclease approach. Cell Rep. 29, 1369–1380 (2019). This study, along with the studies by Chen et al. (2017) and Ginno et al. (2012), demonstrates the feasibility of mapping R-loops genome-wide by capturing them with a catalytically inactive RNase H that is conjugated either to an epitope tag or to the micrococcal nuclease.
Chedin, F. Nascent connections: R-loops and chromatin patterning. Trends Genet. 32, 828–838 (2016).
Crossley, M. P., Bocek, M. & Cimprich, K. A. R-loops as cellular regulators and genomic threats. Mol. Cell 73, 398–411 (2019).
Garcia-Muse, T. & Aguilera, A. R loops: from physiological to pathological roles. Cell 179, 604–618 (2019).
Niehrs, C. & Luke, B. Regulatory R-loops as facilitators of gene expression and genome stability. Nat. Rev. Mol. Cell Biol. 21, 167–178 (2020).
Grunseich, C. et al. Senataxin mutation reveals how R-loops promote transcription by blocking DNA methylation at gene promoters. Mol. Cell 69, 426–437 (2018).
Skourti-Stathaki, K., Proudfoot, N. J. & Gromak, N. Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol. Cell 42, 794–805 (2011).
Sun, Q., Csorba, T., Skourti-Stathaki, K., Proudfoot, N. J. & Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 340, 619–621 (2013).
Tan-Wong, S. M., Dhir, S. & Proudfoot, N. J. R-loops promote antisense transcription across the mammalian genome. Mol. Cell 76, 600–616 (2019).
Holt, I. J. The mitochondrial R-loop. Nucleic Acids Res. 47, 5480–5489 (2019).
Posse, V. et al. RNase H1 directs origin-specific initiation of DNA replication in human mitochondria. PLoS Genet. 15, e1007781 (2019).
Wongsurawat, T., Jenjaroenpun, P., Kwoh, C. K. & Kuznetsov, V. Quantitative model of R-loop forming structures reveals a novel level of RNA-DNA interactome complexity. Nucleic Acids Res. 40, e16 (2012).
Belotserkovskii, B. P. & Hanawalt, P. C. Anchoring nascent RNA to the DNA template could interfere with transcription. Biophys. J. 100, 675–684 (2011).
Gomez-Gonzalez, B. & Aguilera, A. Transcription-mediated replication hindrance: a major driver of genome instability. Genes Dev. 33, 1008–1026 (2019).
Gan, W. et al. R-loop-mediated genomic instability is caused by impairment of replication fork progression. Genes Dev. 25, 2041–2056 (2011).
Helmrich, A., Ballarino, M. & Tora, L. Collisions between replication and transcription complexes cause common fragile site instability at the longest human genes. Mol. Cell 44, 966–977 (2011).
Freudenreich, C. H. R-loops: targets for nuclease cleavage and repeat instability. Curr. Genet. 64, 789–794 (2018).
Sollier, J. et al. Transcription-coupled nucleotide excision repair factors promote R-loop-induced genome instability. Mol. Cell 56, 777–785 (2014).
Saini, N. & Gordenin, D. A. Hypermutation in single-stranded DNA. DNA Repair 91–92, 102868 (2020).
Yu, K. & Lieber, M. R. Current insights into the mechanism of mammalian immunoglobulin class switch recombination. Crit. Rev. Biochem. Mol. Biol. 54, 333–351 (2019).
Costantino, L. & Koshland, D. Genome-wide map of r-loop-induced damage reveals how a subset of R-loops contributes to genomic instability. Mol. Cell 71, 487–497 (2018).
Castellano-Pozo, M. et al. R loops are linked to histone H3 S10 phosphorylation and chromatin condensation. Mol. Cell 52, 583–590 (2013).
Garcia-Pichardo, D. et al. Histone mutants separate R loop formation from genome instability induction. Mol. Cell 66, 597–609 (2017). This study, along with that by Salas-Armenteros et al. (2017), suggests that specific chromatin contexts (such as histone modifications) may alter the properties of R-loops, making them more harmful.
Garcia-Rubio, M. et al. Yra1-bound RNA-DNA hybrids cause orientation-independent transcription-replication collisions and telomere instability. Genes Dev. 32, 965–977 (2018). This study, along with that by Hamperl et al. (2017), reports that co-directional transcription–replication conflicts are associated with enhanced R-loop clearance, providing a plausible explanation as to why transcription and replication tend to be co-oriented in mammalian cells.
Stuckey, R., Garcia-Rodriguez, N., Aguilera, A. & Wellinger, R. E. Role for RNA:DNA hybrids in origin-independent replication priming in a eukaryotic system. Proc. Natl Acad. Sci. USA 112, 5779–5784 (2015).
Gorthi, A. et al. EWS-FLI1 increases transcription to cause R-loops and block BRCA1 repair in Ewing sarcoma. Nature 555, 387–391 (2018).
Nergadze, S. G. et al. CpG-island promoters drive transcription of human telomeres. RNA 15, 2186–2194 (2009).
Azzalin, C. M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Luke, B. et al. The Rat1p 5’ to 3’ exonuclease degrades telomeric repeat-containing RNA and promotes telomere elongation in Saccharomyces cerevisiae. Mol. Cell 32, 465–477 (2008).
Schoeftner, S. & Blasco, M. A. Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nat. Cell Biol. 10, 228–236 (2008).
Pfeiffer, V. & Lingner, J. TERRA promotes telomere shortening through exonuclease 1-mediated resection of chromosome ends. PLoS Genet. 8, e1002747 (2012).
Herold, S. et al. Recruitment of BRCA1 limits MYCN-driven accumulation of stalled RNA polymerase. Nature 567, 545–549 (2019).
Shivji, M. K. K., Renaudin, X., Williams, C. H. & Venkitaraman, A. R. BRCA2 regulates transcription elongation by RNA polymerase II to prevent R-loop accumulation. Cell Rep. 22, 1031–1039 (2018).
Zhang, X. et al. Attenuation of RNA polymerase II pausing mitigates BRCA1-associated R-loop accumulation and tumorigenesis. Nat. Commun. 8, 15908 (2017).
Kim, N. & Jinks-Robertson, S. The Top1 paradox: friend and foe of the eukaryotic genome. DNA Repair 56, 33–41 (2017).
Nakazawa, Y. et al. Ubiquitination of DNA damage-stalled RNAPII promotes transcription-coupled repair. Cell 180, 1228–1244 (2020).
Tufegdzic Vidakovic, A. et al. Regulation of the RNAPII Pool Is Integral to the DNA damage response. Cell 180, e1221 (2020).
Cerritelli, S. M., Crouch, R. J. & Ribonuclease, H. the enzymes in eukaryotes. FEBS J. 276, 1494–1505 (2009).
Nguyen, H. D. et al. Functions of replication protein a as a sensor of R loops and a regulator of RNaseH1. Mol. Cell 65, 832–847 (2017).
Hatchi, E. et al. BRCA1 recruitment to transcriptional pause sites is required for R-loop-driven DNA damage repair. Mol. Cell 57, 636–647 (2015). This study, along with the studies by Herold et al. (2019) and Zhang et al. (2017), shows that BRCA1 promotes genome stability and tumour suppression in part by resolving R-loops at transcription pause sites.
Allison, D. F. & Wang, G. G. R-loops: formation, function, and relevance to cell stress. Cell Stress. 3, 38–46 (2019).
Petryk, N. et al. Replication landscape of the human genome. Nat. Commun. 7, 10208 (2016).
Barroso, S. et al. The DNA damage response acts as a safeguard against harmful DNA-RNA hybrids of different origins. EMBO Rep. 20, e47250 (2019).
Matos, D. A. et al. ATR protects the genome against R Loops through a MUS81-triggered feedback loop. Mol. Cell 77, 514–527 (2020).
Zhao, H., Zhu, M., Limbo, O. & Russell, P. RNase H eliminates R-loops that disrupt DNA replication but is nonessential for efficient DSB repair. EMBO Rep. https://doi.org/10.15252/embr.201745335 (2018).
Perego, M. G. L., Taiana, M., Bresolin, N., Comi, G. P. & Corti, S. R-loops in motor neuron diseases. Mol. Neurobiol. 56, 2579–2589 (2019).
Richard, P. & Manley, J. L. R loops and links to human disease. J. Mol. Biol. 429, 3168–3180 (2017).
Colak, D. et al. Promoter-bound trinucleotide repeat mRNA drives epigenetic silencing in fragile X syndrome. Science 343, 1002–1005 (2014).
Groh, M., Lufino, M. M., Wade-Martins, R. & Gromak, N. R-loops associated with triplet repeat expansions promote gene silencing in Friedreich ataxia and fragile X syndrome. PLoS Genet. 10, e1004318 (2014).
Sagie, S. et al. Telomeres in ICF syndrome cells are vulnerable to DNA damage due to elevated DNA:RNA hybrids. Nat. Commun. 8, 14015 (2017).
Zhu, Q. et al. Heterochromatin-encoded satellite RNAs induce breast cancer. Mol. Cell 70, 842–853 (2018).
Zhu, Q. et al. BRCA1 tumour suppression occurs via heterochromatin-mediated silencing. Nature 477, 179–184 (2011).
Bhatia, V. et al. BRCA2 prevents R-loop accumulation and associates with TREX-2 mRNA export factor PCID2. Nature 511, 362–365 (2014).
Garcia-Rubio, M. L. et al. The Fanconi anemia pathway protects genome integrity from R-loops. PLoS Genet. 11, e1005674 (2015).
Schwab, R. A. et al. The Fanconi anemia pathway maintains genome stability by coordinating replication and transcription. Mol. Cell 60, 351–361 (2015).
Bhatia, V., Herrera-Moyano, E., Aguilera, A. & Gomez-Gonzalez, B. The role of replication-associated repair factors on R-loops. Genes https://doi.org/10.3390/genes8070171 (2017).
Chang, E. Y. & Stirling, P. C. Replication fork protection factors controlling R-loop bypass and suppression. Genes https://doi.org/10.3390/genes8010033 (2017).
Okamoto, Y., Hejna, J. & Takata, M. Regulation of R-loops and genome instability in Fanconi anemia. J. Biochem. 165, 465–470 (2019).
Paull, T. T. RNA-DNA hybrids and the convergence with DNA repair. Crit. Rev. Biochem. Mol. Biol. 54, 371–384 (2019).
Wells, J. P., White, J. & Stirling, P. C. R loops and their composite cancer connections. Trends Cancer 5, 619–631 (2019).
Venkitaraman, A. R. How do mutations affecting the breast cancer genes BRCA1 and BRCA2 cause cancer susceptibility? DNA Repair 81, 102668 (2019).
Akman, G. et al. Pathological ribonuclease H1 causes R-loop depletion and aberrant DNA segregation in mitochondria. Proc. Natl Acad. Sci. USA 113, E4276–E4285 (2016).
Holt, I. J. The Jekyll and Hyde character of RNase H1 and its multiple roles in mitochondrial DNA metabolism. DNA Repair 84, 102630 (2019).
Kannan, A., Jiang, X., He, L., Ahmad, S. & Gangwani, L. ZPR1 prevents R-loop accumulation, upregulates SMN2 expression and rescues spinal muscular atrophy. Brain 143, 69–93 (2020).
Wang, I. X. et al. Human proteins that interact with RNA/DNA hybrids. Genome Res. 28, 1405–1414 (2018).
Cristini, A., Groh, M., Kristiansen, M. S. & Gromak, N. RNA/DNA hybrid interactome identifies DXH9 as a molecular player in transcriptional termination and R-loop-associated DNA damage. Cell Rep. 23, 1891–1905 (2018). This study, and that by Wang et al. (2018), reports the identification of human proteins that exhibit preferential interaction with RNA–DNA hybrids, providing a useful resource for future studies aimed at elucidating how RNA–DNA hybrids are regulated and their possible physiological functions.
Marnef, A., Cohen, S. & Legube, G. Transcription-coupled DNA double-strand break repair: active genes need special care. J. Mol. Biol. 429, 1277–1288 (2017).
Adelman, C. A. & Petrini, J. H. Division of labor: DNA repair and the cell cycle specific functions of the Mre11 complex. Cell Cycle 8, 1510–1514 (2009).
Keskin, H., Meers, C. & Storici, F. Transcript RNA supports precise repair of its own DNA gene. RNA Biol. 13, 157–165 (2016).
Aguilera, A. & Gomez-Gonzalez, B. DNA-RNA hybrids: the risks of DNA breakage during transcription. Nat. Struct. Mol. Biol. 24, 439–443 (2017).
Bader, A. S. & Bushell, M. DNA:RNA hybrids form at DNA double-strand breaks in transcriptionally active loci. Cell Death Dis. 11, 280 (2020).
Puget, N., Miller, K. M. & Legube, G. Non-canonical DNA/RNA structures during transcription-coupled double-strand break repair: roadblocks or bona fide repair intermediates? DNA Repair 81, 102661 (2019).
Derr, L. K., Strathern, J. N. & Garfinkel, D. J. RNA-mediated recombination in S. cerevisiae. Cell 67, 355–364 (1991).
Keskin, H. et al. Transcript-RNA-templated DNA recombination and repair. Nature 515, 436–439 (2014).
Lee, M. H. et al. Somatic APP gene recombination in Alzheimer’s disease and normal neurons. Nature 563, 639–645 (2018).
Li, S. et al. Precise gene replacement in rice by RNA transcript-templated homologous recombination. Nat. Biotechnol. 37, 445–450 (2019).
Nowacki, M. et al. RNA-mediated epigenetic programming of a genome-rearrangement pathway. Nature 451, 153–158 (2008).
Ono, R. et al. Double strand break repair by capture of retrotransposon sequences and reverse-transcribed spliced mRNA sequences in mouse zygotes. Sci. Rep. 5, 12281 (2015).
Onozawa, M. et al. Repair of DNA double-strand breaks by templated nucleotide sequence insertions derived from distant regions of the genome. Proc. Natl Acad. Sci. USA 111, 7729–7734 (2014).
Shen, Y. et al. RNA-driven genetic changes in bacteria and in human cells. Mutat. Res. 717, 91–98 (2011).
Storici, F. et al. Repair. Nature 447, 338–341 (2007). This study and the study by Keskin et al. (2014) demonstrate that yeast can use RNA transcript as a template to repair a DNA DSB.
Mazina, O. M., Keskin, H., Hanamshet, K., Storici, F. & Mazin, A. V. Rad52 inverse strand exchange drives RNA-templated DNA double-strand break repair. Mol. Cell 67, 19–29 (2017).
McDevitt, S., Rusanov, T., Kent, T., Chandramouly, G. & Pomerantz, R. T. How RNA transcripts coordinate DNA recombination and repair. Nat. Commun. 9, 1091 (2018).
Keskin, H. & Storici, F. Defects in RNase H2 stimulate DNA break repair by RNA reverse transcribed into cDNA. Microrna 4, 109–116 (2015).
Chakraborty, A. et al. Classical non-homologous end-joining pathway utilizes nascent RNA for error-free double-strand break repair of transcribed genes. Nat. Commun. 7, 13049 (2016).
Jang, Y. et al. Intrinsically disordered protein RBM14 plays a role in generation of RNA:DNA hybrids at double-strand break sites. Proc. Natl Acad. Sci. USA 117, 5329–5338 (2020).
Gout, J. F. et al. The landscape of transcription errors in eukaryotic cells. Sci. Adv. 3, e1701484 (2017).
Amon, J. D. & Koshland, D. RNase H enables efficient repair of R-loop induced DNA damage. Elife https://doi.org/10.7554/eLife.20533 (2016).
D’Alessandro, G. et al. BRCA2 controls DNA:RNA hybrid level at DSBs by mediating RNase H2 recruitment. Nat. Commun. 9, 5376 (2018).
Domingo-Prim, J. et al. EXOSC10 is required for RPA assembly and controlled DNA end resection at DNA double-strand breaks. Nat. Commun. 10, 2135 (2019).
Li, L. et al. DEAD box 1 facilitates removal of RNA and homologous recombination at DNA double-strand breaks. Mol. Cell Biol. 36, 2794–2810 (2016).
Ohle, C. et al. Transient RNA-DNA hybrids are required for efficient double-strand break repair. Cell 167, 1001–1013 (2016).
Yasuhara, T. et al. Human Rad52 promotes XPG-mediated R-loop processing to initiate transcription-associated homologous recombination repair. Cell 175, 558–570 (2018).
Lu, W. T. et al. Drosha drives the formation of DNA:RNA hybrids around DNA break sites to facilitate DNA repair. Nat. Commun. 9, 532 (2018).
Wei, W. et al. A role for small RNAs in DNA double-strand break repair. Cell 149, 101–112 (2012).
Wei, L. et al. DNA damage during the G0/G1 phase triggers RNA-templated, Cockayne syndrome B-dependent homologous recombination. Proc. Natl Acad. Sci. USA 112, E3495–E3504 (2015).
Welty, S. et al. RAD52 is required for RNA-templated recombination repair in post-mitotic neurons. J. Biol. Chem. 293, 1353–1362 (2018).
Hanamshet, K., Mazina, O. M. & Mazin, A. V. Reappearance from obscurity: mammalian Rad52 in homologous recombination. Genes https://doi.org/10.3390/genes7090063 (2016).
Jalan, M., Olsen, K. S. & Powell, S. N. Emerging roles of RAD52 in genome maintenance. Cancers https://doi.org/10.3390/cancers11071038 (2019).
Malacaria, E., Honda, M., Franchitto, A., Spies, M. & Pichierri, P. Physiological and pathological roles of RAD52 at DNA replication Forks. Cancers https://doi.org/10.3390/cancers12020402 (2020).
Nogueira, A., Fernandes, M., Catarino, R. & Medeiros, R. RAD52 functions in homologous recombination and its importance on genomic integrity maintenance and cancer therapy. Cancers https://doi.org/10.3390/cancers11111622 (2019).
Caron, P., van der Linden, J. & van Attikum, H. Bon voyage: a transcriptional journey around DNA breaks. DNA Repair 82, 102686 (2019).
Machour, F. E. & Ayoub, N. Transcriptional regulation at DSBs: mechanisms and consequences. Trends Genet. https://doi.org/10.1016/j.tig.2020.01.001 (2020).
Kim, J., Sturgill, D., Tran, A. D., Sinclair, D. A. & Oberdoerffer, P. Controlled DNA double-strand break induction in mice reveals post-damage transcriptome stability. Nucleic Acids Res. 44, e64 (2016).
Kruhlak, M. et al. The ATM repair pathway inhibits RNA polymerase I transcription in response to chromosome breaks. Nature 447, 730–734 (2007).
Purman, C. E. et al. Regional gene repression by DNA double-strand breaks in G1 phase cells. Mol. Cell Biol. https://doi.org/10.1128/MCB.00181-19 (2019).
Gutbrod, M. J. & Martienssen, R. A. Conserved chromosomal functions of RNA interference. Nat. Rev. Genet. 21, 311–331 (2020).
Michelini, F. et al. From “cellular” RNA to “smart” RNA: multiple roles of RNA in genome stability and beyond. Chem. Rev. 118, 4365–4403 (2018).
Bonath, F., Domingo-Prim, J., Tarbier, M., Friedlander, M. R. & Visa, N. Next-generation sequencing reveals two populations of damage-induced small RNAs at endogenous DNA double-strand breaks. Nucleic Acids Res. 46, 11869–11882 (2018).
Francia, S. et al. Site-specific DICER and DROSHA RNA products control the DNA-damage response. Nature 488, 231–235 (2012).
Michalik, K. M., Bottcher, R. & Forstemann, K. A small RNA response at DNA ends in Drosophila. Nucleic Acids Res. 40, 9596–9603 (2012).
Miki, D. et al. Efficient generation of diRNAs requires components in the posttranscriptional gene silencing pathway. Sci. Rep. 7, 301 (2017).
Pessina, F. et al. Functional transcription promoters at DNA double-strand breaks mediate RNA-driven phase separation of damage-response factors. Nat. Cell Biol. 21, 1286–1299 (2019).
Vitor, A. C. et al. Single-molecule imaging of transcription at damaged chromatin. Sci. Adv. 5, eaau1249 (2019).
Michelini, F. et al. Damage-induced lncRNAs control the DNA damage response through interaction with DDRNAs at individual double-strand breaks. Nat. Cell Biol. 19, 1400–1411 (2017).
Rossiello, F. et al. DNA damage response inhibition at dysfunctional telomeres by modulation of telomeric DNA damage response RNAs. Nat. Commun. 8, 13980 (2017).
Francia, S., Cabrini, M., Matti, V., Oldani, A. & d’Adda di Fagagna, F. DICER, DROSHA and DNA damage response RNAs are necessary for the secondary recruitment of DNA damage response factors. J. Cell Sci. 129, 1468–1476 (2016).
Wang, Q. & Goldstein, M. Small RNAs Recruit chromatin-modifying enzymes MMSET and Tip60 to reconfigure damaged DNA upon double-strand break and facilitate repair. Cancer Res. 76, 1904–1915 (2016).
Gao, M. et al. Ago2 facilitates Rad51 recruitment and DNA double-strand break repair by homologous recombination. Cell Res. 24, 532–541 (2014).
Mirman, Z. & de Lange, T. 53BP1: a DSB escort. Genes Dev. 34, 7–23 (2020).
Tarsounas, M. & Sung, P. The antitumorigenic roles of BRCA1-BARD1 in DNA repair and replication. Nat. Rev. Mol. Cell Biol. 21, 284–299 (2020).
Kilic, S. et al. Phase separation of 53BP1 determines liquid-like behavior of DNA repair compartments. EMBO J. 38, e101379 (2019). This study and the study by Pessina et al. (2019) demonstrate that focal assembly of 53BP1 at DNA damage sites is driven by phase separation, with Pessina et al. suggesting a role for diRNA in this process.
Botuyan, M. V. et al. Mechanism of 53BP1 activity regulation by RNA-binding TIRR and a designer protein. Nat. Struct. Mol. Biol. 25, 591–600 (2018).
Dai, Y., Zhang, A., Shan, S., Gong, Z. & Zhou, Z. Structural basis for recognition of 53BP1 tandem Tudor domain by TIRR. Nat. Commun. 9, 2123 (2018).
Drane, P. et al. TIRR regulates 53BP1 by masking its histone methyl-lysine binding function. Nature 543, 211–216 (2017).
Wang, J. et al. Molecular basis for the inhibition of the methyl-lysine binding function of 53BP1 by TIRR. Nat. Commun. 9, 2689 (2018).
Jacquet, K. et al. The TIP60 complex regulates bivalent chromatin recognition by 53BP1 through direct H4K20me binding and H2AK15 acetylation. Mol. Cell 62, 409–421 (2016).
Tang, J. et al. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nat. Struct. Mol. Biol. 20, 317–325 (2013).
Lenzken, S. C. et al. FUS-dependent phase separation initiates double-strand break repair. Preprint at bioRxiv https://doi.org/10.1101/798884 (2019).
Wang, W. Y. et al. Interaction of FUS and HDAC1 regulates DNA damage response and repair in neurons. Nat. Neurosci. 16, 1383–1391 (2013).
Altmeyer, M. et al. Liquid demixing of intrinsically disordered proteins is seeded by poly(ADP-ribose). Nat. Commun. 6, 8088 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Naumann, M. et al. Impaired DNA damage response signaling by FUS-NLS mutations leads to neurodegeneration and FUS aggregate formation. Nat. Commun. 9, 335 (2018).
Chen, L. et al. The augmented R-loop is a unifying mechanism for myelodysplastic syndromes induced by high-risk splicing factor mutations. Mol. Cell 69, 412–425 (2018).
Li, X. & Manley, J. L. Inactivation of the SR protein splicing factor ASF/SF2 results in genomic instability. Cell 122, 365–378 (2005).
Bonnet, A. et al. Introns protect eukaryotic genomes from transcription-associated genetic instability. Mol. Cell 67, 608–621 (2017). This study and the study by Li and Manley (2005) provide evidence that splicing factors, rather than splicing per se, inhibit co-transcriptional R-loop formation by binding to pre-mRNA, which prevents stable RNA–DNA interaction.
Gupta, S. K., Luo, L. & Yen, L. RNA-mediated gene fusion in mammalian cells. Proc. Natl Acad. Sci. USA 115, E12295–E12304 (2018). This study provides evidence that an RNA transcript containing sufficient sequence homologies to two disparate genomic loci can drive gene fusion.
Wu, H., Li, X. & Li, H. Gene fusions and chimeric RNAs, and their implications in cancer. Genes. Dis. 6, 385–390 (2019).
Kandel, E. S. & Nudler, E. Template switching by RNA polymerase II in vivo. Evidence and implications from a retroviral system. Mol. Cell 10, 1495–1502 (2002).
Yan, Z. et al. Genome-wide colocalization of RNA-DNA interactions and fusion RNA pairs. Proc. Natl Acad. Sci. USA 116, 3328–3337 (2019). This study maps RNA–DNA interactions genome-wide in normal cells and reports that spatial proximity between RNA and genomic DNA is associated with the creation of chimeric fusion transcripts.
Huang, H., Weng, H. & Chen, J. The biogenesis and precise control of RNA m6A methylation. Trends Genet. 36, 44–52 (2020).
Xiang, Y. et al. RNA m6A methylation regulates the ultraviolet-induced DNA damage response. Nature 543, 573–576 (2017).
Stern, H. R., Sefcikova, J., Chaparro, V. E. & Beuning, P. J. Mammalian DNA polymerase kappa activity and specificity. Molecules https://doi.org/10.3390/molecules24152805 (2019).
Rastogi, R. P., Richa, Kumar, A., Tyagi, M. B. & Sinha, R. P. Molecular mechanisms of ultraviolet radiation-induced DNA damage and repair. J. Nucleic Acids 2010, 592980 (2010).
Fu, Y. & Zhuang, X. m6A-binding YTHDF proteins promote stress granule formation. Nat. Chem. Biol. https://doi.org/10.1038/s41589-020-0524-y (2020).
Gao, Y. et al. Multivalent m6A motifs promote phase separation of YTHDF proteins. Cell Res. 29, 767–769 (2019).
Ries, R. J. et al. m6A enhances the phase separation potential of mRNA. Nature 571, 424–428 (2019).
Abakir, A. et al. N6-methyladenosine regulates the stability of RNA:DNA hybrids in human cells. Nat. Genet. https://doi.org/10.1038/s41588-019-0549-x (2019).
Tresini, M. et al. The core spliceosome as target and effector of non-canonical ATM signalling. Nature 523, 53–58 (2015).
Bell, J. C. et al. Chromatin-associated RNA sequencing (ChAR-seq) maps genome-wide RNA-to-DNA contacts. Elife https://doi.org/10.7554/eLife.27024 (2018).
Li, X. et al. GRID-seq reveals the global RNA-chromatin interactome. Nat. Biotechnol. 35, 940–950 (2017).
Mondal, T., Rasmussen, M., Pandey, G. K., Isaksson, A. & Kanduri, C. Characterization of the RNA content of chromatin. Genome Res. 20, 899–907 (2010). This study and the studies by Bell et al. (2018) and Li et al. (2017) describe different approaches for mapping the global RNA–chromatin interactome in D. melanogaster, mouse and human cells.
Rodriguez-Campos, A. & Azorin, F. RNA is an integral component of chromatin that contributes to its structural organization. PLoS ONE 2, e1182 (2007).
Caudron-Herger, M. & Rippe, K. Nuclear architecture by RNA. Curr. Opin. Genet. Dev. 22, 179–187 (2012).
Johnson, W. L. & Straight, A. F. RNA-mediated regulation of heterochromatin. Curr. Opin. Cell Biol. 46, 102–109 (2017).
Li, X. & Fu, X. D. Chromatin-associated RNAs as facilitators of functional genomic interactions. Nat. Rev. Genet. 20, 503–519 (2019).
Mishra, K. & Kanduri, C. Understanding long noncoding RNA and chromatin interactions: what we know so far. Noncoding RNA https://doi.org/10.3390/ncrna5040054 (2019).
Wang, Y. & Feigon, J. Structural biology of telomerase and its interaction at telomeres. Curr. Opin. Struct. Biol. 47, 77–87 (2017).
Mangaonkar, A. A. & Patnaik, M. M. Short telomere syndromes in clinical practice: bridging bench and bedside. Mayo Clin. Proc. 93, 904–916 (2018).
Toubiana, S. & Selig, S. DNA:RNA hybrids at telomeres - when it is better to be out of the (R) loop. FEBS J. 285, 2552–2566 (2018).
Kabeche, L., Nguyen, H. D., Buisson, R. & Zou, L. A mitosis-specific and R loop-driven ATR pathway promotes faithful chromosome segregation. Science 359, 108–114 (2018).
Montero, J. J., Lopez de Silanes, I., Grana, O. & Blasco, M. A. Telomeric RNAs are essential to maintain telomeres. Nat. Commun. 7, 12534 (2016).
Caceres-Gutierrez, R. & Herrera, L. A. Centromeric non-coding transcription: opening the black box of chromosomal instability? Curr. Genomics 18, 227–235 (2017).
Maicher, A., Kastner, L. & Luke, B. Telomeres and disease: enter TERRA. RNA Biol. 9, 843–849 (2012).
Arora, R. et al. RNaseH1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat. Commun. 5, 5220 (2014).
Hansen, A. S. et al. Distinct classes of chromatin loops revealed by deletion of an RNA-binding region in CTCF. Mol. Cell 76, 395–411 (2019).
Saldana-Meyer, R. et al. RNA interactions are essential for CTCF-mediated genome organization. Mol. Cell 76, 412–422 (2019). This study and the study by Hansen et al. (2019) show that CTCF-dependent genome organization is at least in part mediated by the interactions between CTCF and RNA.
Misteli, T. & Soutoglou, E. The emerging role of nuclear architecture in DNA repair and genome maintenance. Nat. Rev. Mol. Cell Biol. 10, 243–254 (2009).
Li, Y., Syed, J. & Sugiyama, H. RNA-DNA triplex formation by long noncoding RNAs. Cell Chem. Biol. 23, 1325–1333 (2016).
Klein, S. J. & O’Neill, R. J. Transposable elements: genome innovation, chromosome diversity, and centromere conflict. Chromosome Res. 26, 5–23 (2018).
Platt, R. N. II, Vandewege, M. W. & Ray, D. A. Mammalian transposable elements and their impacts on genome evolution. Chromosome Res. 26, 25–43 (2018).
Wimmer, K., Callens, T., Wernstedt, A. & Messiaen, L. The NF1 gene contains hotspots for L1 endonuclease-dependent de novo insertion. PLoS Genet. 7, e1002371 (2011).
Burns, K. H. Our conflict with transposable elements and its implications for human disease. Annu. Rev. Pathol. 15, 51–70 (2020).
Kazazian, H. H. Jr. & Moran, J. V. Mobile DNA in health and disease. N. Engl. J. Med. 377, 361–370 (2017).
Hall, A. C., Ostrowski, L. A., Pietrobon, V. & Mekhail, K. Repetitive DNA loci and their modulation by the non-canonical nucleic acid structures R-loops and G-quadruplexes. Nucleus 8, 162–181 (2017).
Ardeljan, D. et al. Cell fitness screens reveal a conflict between LINE-1 retrotransposition and DNA replication. Nat. Struct. Mol. Biol. 27, 168–178 (2020).
Jeon, J. et al. Retroelement insertion in a CRISPR/Cas9 editing site in the early embryo intensifies genetic mosaicism. Front. Cell Dev. Biol. 7, 273 (2019).
Morrish, T. A. et al. DNA repair mediated by endonuclease-independent LINE-1 retrotransposition. Nat. Genet. 31, 159–165 (2002).
Maizels, N. & Davis, L. Initiation of homologous recombination at DNA nicks. Nucleic Acids Res. 46, 6962–6973 (2018).
Wang, Y. et al. BRCA1 intronic Alu elements drive gene rearrangements and PARP inhibitor resistance. Nat. Commun. 10, 5661 (2019). This study demonstrates that Alu-mediated genomic rearrangements upstream of a nonsense mutation in the BRCA1 gene can restore the correct reading frame, leading to the production of a stable BRCA1 protein and the development of chemoresistance.
Anwar, S. L., Wulaningsih, W. & Lehmann, U. Transposable elements in human cancer: causes and consequences of deregulation. Int. J. Mol. Sci. https://doi.org/10.3390/ijms18050974 (2017).
Pavlicek, A. et al. Evolution of the tumor suppressor BRCA1 locus in primates: implications for cancer predisposition. Hum. Mol. Genet. 13, 2737–2751 (2004).
Mita, P. et al. BRCA1 and S phase DNA repair pathways restrict LINE-1 retrotransposition in human cells. Nat. Struct. Mol. Biol. 27, 179–191 (2020).
Goodier, J. L. Restricting retrotransposons: a review. Mob. DNA 7, 16 (2016).
Czech, B. et al. piRNA-guided genome defense: from biogenesis to silencing. Annu. Rev. Genet. 52, 131–157 (2018).
Ernst, C., Odom, D. T. & Kutter, C. The emergence of piRNAs against transposon invasion to preserve mammalian genome integrity. Nat. Commun. 8, 1411 (2017).
Iwasaki, Y. W., Siomi, M. C. & Siomi, H. PIWI-interacting RNA: its biogenesis and functions. Annu. Rev. Biochem. 84, 405–433 (2015).
Ozata, D. M., Gainetdinov, I., Zoch, A., O’Carroll, D. & Zamore, P. D. PIWI-interacting RNAs: small RNAs with big functions. Nat. Rev. Genet. 20, 89–108 (2019).
Schmid, M. & Jensen, T. H. Controlling nuclear RNA levels. Nat. Rev. Genet. 19, 518–529 (2018).
Dos Santos, R. F. et al. Major 3’-5’ exoribonucleases in the metabolism of coding and non-coding RNA. Prog. Mol. Biol. Transl. Sci. 159, 101–155 (2018).
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009).
Kilchert, C., Wittmann, S. & Vasiljeva, L. The regulation and functions of the nuclear RNA exosome complex. Nat. Rev. Mol. Cell Biol. 17, 227–239 (2016).
Levy, S. & Schuster, G. Polyadenylation and degradation of RNA in the mitochondria. Biochem. Soc. Trans. 44, 1475–1482 (2016).
Nair, L., Chung, H. & Basu, U. Regulation of long non-coding RNAs and genome dynamics by the RNA surveillance machinery. Nat. Rev. Mol. Cell Biol. 21, 123–136 (2020).
Pefanis, E. et al. RNA exosome-regulated long non-coding RNA transcription controls super-enhancer activity. Cell 161, 774–789 (2015).
Silva, S., Camino, L. P. & Aguilera, A. Human mitochondrial degradosome prevents harmful mitochondrial R loops and mitochondrial genome instability. Proc. Natl Acad. Sci. USA 115, 11024–11029 (2018).
Morton, D. J. et al. The RNA exosome and RNA exosome-linked disease. RNA 24, 127–142 (2018).
Pashler, A. L., Towler, B. P., Jones, C. I. & Newbury, S. F. The roles of the exoribonucleases DIS3L2 and XRN1 in human disease. Biochem. Soc. Trans. 44, 1377–1384 (2016).
Dhir, A. et al. Mitochondrial double-stranded RNA triggers antiviral signalling in humans. Nature 560, 238–242 (2018). This study and the study by Silva et al. (2018) report that the loss of mitochondrial RNA degradosome activity leads to R-loop-associated genome instability as well as the formation of immunogenic double-stranded RNA, highlighting the importance of RNA turnover in mitochondria.
Bresson, S. & Tollervey, D. Surveillance-ready transcription: nuclear RNA decay as a default fate. Open Biol. https://doi.org/10.1098/rsob.170270 (2018).
Traut, T. W. Physiological concentrations of purines and pyrimidines. Mol. Cell Biochem. 140, 1–22 (1994).
Nava, G. M. et al. One, no one, and one hundred thousand: the many forms of ribonucleotides in DNA. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21051706 (2020).
Cerritelli, S. M. & Crouch, R. J. The balancing act of ribonucleotides in DNA. Trends Biochem. Sci. 41, 434–445 (2016).
Nick McElhinny, S. A. et al. Abundant ribonucleotide incorporation into DNA by yeast replicative polymerases. Proc. Natl Acad. Sci. USA 107, 4949–4954 (2010).
Clausen, A. R., Zhang, S., Burgers, P. M., Lee, M. Y. & Kunkel, T. A. Ribonucleotide incorporation, proofreading and bypass by human DNA polymerase delta. DNA Repair 12, 121–127 (2013).
Goksenin, A. Y. et al. Human DNA polymerase epsilon is able to efficiently extend from multiple consecutive ribonucleotides. J. Biol. Chem. 287, 42675–42684 (2012).
Kasiviswanathan, R. & Copeland, W. C. Ribonucleotide discrimination and reverse transcription by the human mitochondrial DNA polymerase. J. Biol. Chem. 286, 31490–31500 (2011).
Koh, K. D., Balachander, S., Hesselberth, J. R. & Storici, F. Ribose-seq: global mapping of ribonucleotides embedded in genomic DNA. Nat. Methods 12, 251–257 (2015). This study describes an elegant protocol for mapping embedded ribonucleotides genome-wide in yeast, which takes advantage of alkali-dependent cleavage of the DNA backbone at the site of embedded ribonucleotides.
Crespan, E. et al. Impact of ribonucleotide incorporation by DNA polymerases beta and lambda on oxidative base excision repair. Nat. Commun. 7, 10805 (2016).
Nick McElhinny, S. A. & Ramsden, D. A. Polymerase mu is a DNA-directed DNA/RNA polymerase. Mol. Cell Biol. 23, 2309–2315 (2003).
Pryor, J. M. et al. Ribonucleotide incorporation enables repair of chromosome breaks by nonhomologous end joining. Science 361, 1126–1129 (2018). This study reports the striking finding that the insertion of ribonucleotides by error-prone DNA polymerases stimulates DSB ligation.
Su, Y. et al. Human DNA polymerase eta has reverse transcriptase activity in cellular environments. J. Biol. Chem. 294, 6073–6081 (2019).
Kennedy, E. M., Amie, S. M., Bambara, R. A. & Kim, B. Frequent incorporation of ribonucleotides during HIV-1 reverse transcription and their attenuated repair in macrophages. J. Biol. Chem. 287, 14280–14288 (2012).
Williams, J. S., Lujan, S. A. & Kunkel, T. A. Processing ribonucleotides incorporated during eukaryotic DNA replication. Nat. Rev. Mol. Cell Biol. 17, 350–363 (2016).
Hyjek, M., Figiel, M., Nowotny, M. & RNases, H. Structure and mechanism. DNA Repair 84, 102672 (2019).
Cerritelli, S. M. et al. Failure to produce mitochondrial DNA results in embryonic lethality in Rnaseh1 null mice. Mol. Cell 11, 807–815 (2003).
Holmes, J. B. et al. Primer retention owing to the absence of RNase H1 is catastrophic for mitochondrial DNA replication. Proc. Natl Acad. Sci. USA 112, 9334–9339 (2015).
Lima, W. F. et al. Viable RNaseH1 knockout mice show RNaseH1 is essential for R loop processing, mitochondrial and liver function. Nucleic Acids Res. 44, 5299–5312 (2016).
Reijns, M. A. et al. Enzymatic removal of ribonucleotides from DNA is essential for mammalian genome integrity and development. Cell 149, 1008–1022 (2012).
Grossman, L. I., Watson, R. & Vinograd, J. The presence of ribonucleotides in mature closed-circular mitochondrial DNA. Proc. Natl Acad. Sci. USA 70, 3339–3343 (1973).
Wong-Staal, F., Mendelsohn, J. & Goulian, M. Ribonucleotides in closed circular mitochondrial DNA from HeLa cells. Biochem. Biophys. Res. Commun. 53, 140–148 (1973).
Kellner, V. & Luke, B. Molecular and physiological consequences of faulty eukaryotic ribonucleotide excision repair. EMBO J https://doi.org/10.15252/embj.2019102309 (2019).
Klein, H. L. Genome instabilities arising from ribonucleotides in DNA. DNA Repair 56, 26–32 (2017).
Chiu, H. C. et al. RNA intrusions change DNA elastic properties and structure. Nanoscale 6, 10009–10017 (2014).
Lujan, S. A., Williams, J. S., Clausen, A. R., Clark, A. B. & Kunkel, T. A. Ribonucleotides are signals for mismatch repair of leading-strand replication errors. Mol. Cell 50, 437–443 (2013).
Hiller, B. et al. Mammalian RNase H2 removes ribonucleotides from DNA to maintain genome integrity. J. Exp. Med. 209, 1419–1426 (2012). This study and the studies by Cerritelli et al. (2003) and Reijns et al. (2012) are among the first to describe the deleterious phenotypes associated with a complete loss of RNase H1 or RNase H2 in mice.
Uehara, R. et al. Two RNase H2 mutants with differential rNMP processing activity reveal a threshold of ribonucleotide tolerance for embryonic development. Cell Rep. 25, 1135–1145 (2018).
Cilli, P., Minoprio, A., Bossa, C., Bignami, M. & Mazzei, F. Formation and repair of mismatches containing ribonucleotides and oxidized bases at repeated DNA sequences. J. Biol. Chem. 290, 26259–26269 (2015).
Mentegari, E. et al. Ribonucleotide incorporation by human DNA polymerase eta impacts translesion synthesis and RNase H2 activity. Nucleic Acids Res. 45, 2600–2614 (2017).
Malfatti, M. C. et al. Abasic and oxidized ribonucleotides embedded in DNA are processed by human APE1 and not by RNase H2. Nucleic Acids Res. 45, 11193–11212 (2017).
Cai, Y., Geacintov, N. E. & Broyde, S. Ribonucleotides as nucleotide excision repair substrates. DNA Repair 13, 55–60 (2014).
Vaisman, A. & Woodgate, R. Redundancy in ribonucleotide excision repair: competition, compensation, and cooperation. DNA Repair 29, 74–82 (2015).
Lindsey-Boltz, L. A., Kemp, M. G., Hu, J. & Sancar, A. Analysis of ribonucleotide removal from DNA by human nucleotide excision repair. J. Biol. Chem. 290, 29801–29807 (2015).
Sassa, A. et al. Processing of a single ribonucleotide embedded into DNA by human nucleotide excision repair and DNA polymerase eta. Sci. Rep. 9, 13910 (2019).
Lazzaro, F. et al. RNase H and postreplication repair protect cells from ribonucleotides incorporated in DNA. Mol. Cell 45, 99–110 (2012).
Nick McElhinny, S. A. et al. Genome instability due to ribonucleotide incorporation into DNA. Nat. Chem. Biol. 6, 774–781 (2010).
Sparks, J. L. & Burgers, P. M. Error-free and mutagenic processing of topoisomerase 1-provoked damage at genomic ribonucleotides. EMBO J. 34, 1259–1269 (2015).
Epshtein, A., Potenski, C. J. & Klein, H. L. Increased spontaneous recombination in RNase H2-deficient cells arises from multiple contiguous rNMPs and not from single rNMP residues incorporated by DNA polymerase epsilon. Microb. Cell 3, 248–254 (2016).
Zimmermann, M. et al. CRISPR screens identify genomic ribonucleotides as a source of PARP-trapping lesions. Nature 559, 285–289 (2018).
Huang, S. N., Williams, J. S., Arana, M. E., Kunkel, T. A. & Pommier, Y. Topoisomerase I-mediated cleavage at unrepaired ribonucleotides generates DNA double-strand breaks. EMBO J. 36, 361–373 (2017). This study and the studies by Nick McElhinny et al. (Nat. Chem. Biol., 2010) and Sparks and Burgers (2015) demonstrate how unexcised genomic ribonucleotides can lead to genome instability through a TOP1-mediated repair mechanism.
Sciascia, N. et al. Suppressing proteasome mediated processing of topoisomerase II DNA-protein complexes preserves genome integrity. Elife https://doi.org/10.7554/eLife.53447 (2020).
Berglund, A. K. et al. Nucleotide pools dictate the identity and frequency of ribonucleotide incorporation in mitochondrial DNA. PLoS Genet. 13, e1006628 (2017).
Moss, C. F. et al. Aberrant ribonucleotide incorporation and multiple deletions in mitochondrial DNA of the murine MPV17 disease model. Nucleic Acids Res. 45, 12808–12815 (2017).
Reyes, A. et al. RNASEH1 mutations impair mtDNA Replication and cause adult-onset mitochondrial encephalomyopathy. Am. J. Hum. Genet. 97, 186–193 (2015).
Gunther, C. et al. Defective removal of ribonucleotides from DNA promotes systemic autoimmunity. J. Clin. Invest. 125, 413–424 (2015).
Pizzi, S. et al. Reduction of hRNase H2 activity in Aicardi-Goutieres syndrome cells leads to replication stress and genome instability. Hum. Mol. Genet. 24, 649–658 (2015).
Crow, Y. J. et al. Mutations in genes encoding ribonuclease H2 subunits cause Aicardi-Goutieres syndrome and mimic congenital viral brain infection. Nat. Genet. 38, 910–916 (2006).
Mackenzie, K. J. et al. Ribonuclease H2 mutations induce a cGAS/STING-dependent innate immune response. EMBO J. 35, 831–844 (2016).
Pokatayev, V. et al. RNase H2 catalytic core Aicardi-Goutieres syndrome-related mutant invokes cGAS-STING innate immune-sensing pathway in mice. J. Exp. Med. 213, 329–336 (2016). This study and the study by Mackenzie et al. (2016) model disease-associated Rnaseh2 mutations in mice and show that deficient ribonucleotide excision leads to the activation of inflammatory immune responses.
Lim, Y. W. et al. hypomethylation and RNA:DNA hybrid accumulation in Aicardi-Goutieres syndrome. Elife https://doi.org/10.7554/eLife.08007 (2015).
Chon, H. et al. RNase H2 roles in genome integrity revealed by unlinking its activities. Nucleic Acids Res. 41, 3130–3143 (2013).
Cornelio, D. A., Sedam, H. N., Ferrarezi, J. A., Sampaio, N. M. & Argueso, J. L. Both R-loop removal and ribonucleotide excision repair activities of RNase H2 contribute substantially to chromosome stability. DNA Repair 52, 110–114 (2017).
Storci, G. et al. Genomic stability, anti-inflammatory phenotype, and up-regulation of the RNAseH2 in cells from centenarians. Cell Death Differ. 26, 1845–1858 (2019). This study reports that cells of centenarians overexpress RNase H2, while various diseased tissues show reduced RNase H2 expression, potentially linking RNase H2 function to human longevity.
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Acknowledgements
The authors thank S. John and N. Johnson for critical reading of the manuscript. The authors acknowledge and apologize to peers whose work could not be cited owing to space limitation.
Author information
Authors and Affiliations
Contributions
D.Z. researched data for article. D.Z., and A.N. wrote the article. All authors contributed substantially to discussion of the content and reviewed/edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks A. Aguilera, who co-reviewed with B. Gómez-González, F. Storici, who co-reviewed with C. Meers, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Replisome
-
A large ensemble of proteins that together create a replication fork and initiate DNA synthesis from a replication origin.
- Nucleolytic cleavage
-
A chemical or enzymatic process that breaks phosphodiester bonds in the DNA backbone.
- Structure-specific nucleases
-
Enzymes that recognize and cleave specific types of secondary structure within DNA.
- Nucleotide excision repair
-
A mode of DNA repair wherein a string of nucleotides containing the damaged bases are excised and the resultant gap is filled in by new DNA synthesis.
- Transition mutations
-
A type of point mutation that changes a purine nucleotide to another purine or a pyrimidine nucleotide to another pyrimidine.
- Immunoglobulin class switch recombination
-
A physiological gene rearrangement event in B lymphocytes that replaces the heavy-chain constant region of the expressed immunoglobulin isotype with a different heavy-chain constant region, thereby altering the effector function of the antibody.
- CpG islands
-
Distinct regions of the genome that contain a large number of predominantly unmethylated CG dinucleotide repeats, which are frequently found near mammalian gene promoters.
- Promoter-proximal pausing sites
-
Regions near the promoter of genes where RNA polymerase pauses until it is stimulated to resume transcriptional elongation.
- Gene gating
-
The process of tethering genes to nuclear pore complexes, which enables rapid RNA maturation and export.
- Gene conversion
-
A homology-directed DNA recombination event wherein a resected double-strand break invades the intact sister chromatid and uses it as a template to restore sequences around the break site.
- Single-strand annealing
-
A mutagenic form of DNA repair wherein repetitive regions on either side of a double-strand break are converted into single-stranded DNA by end resection and annealed to one another, resulting in the deletion of the intervening sequences.
- Break-induced replication
-
A non-reciprocal DNA recombination event wherein a resected one-ended double-strand break invades an intact genomic region and initiates long-range unidirectional templated DNA synthesis that typically proceeds to the end of the invaded chromosome.
- Synthetic lethality
-
A condition where combined deficiencies in two or more genes result in cell death, while perturbation of any one of these genes is tolerated.
- Phase separation
-
The separation of a single homogeneous mixture into two distinct phases.
- Ribonucleoprotein
-
A macromolecular complex consisting of both RNA and protein.
- Spliceosome
-
A large multisubunit ribonucleoprotein complex that removes introns from a precursor mRNA transcript.
- trans splicing
-
An RNA splicing event in which exons of two disparate precursor mRNAs are joined together into a single chimeric transcript.
- cis splicing of adjacent genes
-
An event in which two neighbouring genes are read as a single message due to readthrough transcription followed by intergenic splicing.
- Epitranscriptomic
-
Pertaining to RNA modifications that potentially regulate gene expression but that do not involve alterations in the primary ribonucleotide sequence.
- Translesion synthesis
-
A DNA damage tolerance mechanism that enables DNA synthesis past replication-blocking DNA lesions.
- Base excision repair
-
A mode of repair that removes a small, non-helix-distorting base lesion from the genome, leaving an abasic site that is subsequently cleaved and filled in by either non-displacement (short-patch) or strand-displacement (long-patch) DNA synthesis.
- Topologically associated domain
-
(TAD). A genomic domain created by chromosome folding that favours chromatin interactions within its borders.
- CTCF-mediated chromatin looping
-
A process by which CTCF binding to DNA facilitates long-range chromatin interactions between regions that are far apart on the linear chromosome.
- RNA–DNA triplex
-
A form of RNA–DNA interaction in which the RNA strand inserts itself into the major groove of the duplex DNA in a sequence-specific manner.
- RNA exosome
-
A multiprotein ribonuclease complex capable of degrading various types of RNA molecule.
- Degradosome
-
A multicomponent ribonuclease complex that degrades mitochondrial RNA.
- Slipped-strand mispairing
-
Erroneous base pairing in regions of repetitive DNA due to sequence misalignment, resulting in a short stretch of unpaired nucleotides that form a bulge in the DNA.
- Non-allelic recombination
-
A DNA recombination event between any two regions in the genome that have a high degree of sequence homology but are not alleles.
Rights and permissions
About this article
Cite this article
Zong, D., Oberdoerffer, P., Batista, P.J. et al. RNA: a double-edged sword in genome maintenance. Nat Rev Genet 21, 651–670 (2020). https://doi.org/10.1038/s41576-020-0263-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-020-0263-7
This article is cited by
-
RNA processing mechanisms contribute to genome organization and stability in B cells
Oncogene (2024)
-
R-loops as Janus-faced modulators of DNA repair
Nature Cell Biology (2021)
-
BRCA1 and RNAi factors promote repair mediated by small RNAs and PALB2–RAD52
Nature (2021)