Abstract
The clustered regularly interspaced short palindromic repeats (CRISPR) renaissance was catalysed by the discovery that RNA-guided prokaryotic CRISPR-associated (Cas) proteins can create targeted double-strand breaks in mammalian genomes. This finding led to the development of CRISPR systems that harness natural DNA repair mechanisms to repair deficient genes more easily and precisely than ever before. CRISPR has been used to knock out harmful mutant genes and to fix errors in coding sequences to rescue disease phenotypes in preclinical studies and in several clinical trials. However, most genetic disorders result from combinations of mutations, deletions and duplications in the coding and non-coding regions of the genome and therefore require sophisticated genome engineering strategies beyond simple gene knockout. To overcome this limitation, the toolbox of natural and engineered CRISPR–Cas systems has been dramatically expanded to include diverse tools that function in human cells for precise genome editing and epigenome engineering. The application of CRISPR technology to edit the non-coding genome, modulate gene regulation, make precise genetic changes and target infectious diseases has the potential to lead to curative therapies for many previously untreatable diseases.
Key points
-
CRISPR systems are RNA-guided ribonucleoproteins that function as both sequence-specific nucleic acid-targeting proteins and nucleases; these systems are being developed as therapies for simple Mendelian disorders.
-
Novel approaches using newly discovered CRISPR systems and Cas protein engineering have expanded the available genome engineering toolbox, enabling the development of potentially curative therapies for diseases with complex drivers.
-
Targeting and altering the non-coding genome with CRISPR could potentially ameliorate disease by changing the transcription or translation of target genes.
-
The use of CRISPR systems with nuclease-dead Cas proteins fused to transcriptional or epigenetic modulators enables targeted gene regulation without inducing DNA damage or altering the genetic code.
-
CRISPR base editors and prime editors can be used to create precise genome edits such as therapeutic mutations, insertions or deletions that are difficult to achieve using wild-type CRISPR–Cas nucleases.
-
In addition to genome engineering, the CRISPR toolbox could potentially be used to prevent and treat infectious diseases.
Similar content being viewed by others
Introduction
Adaptive immunity was once thought to be an exclusive feature of vertebrates1. However, the discovery that prokaryotes also possess a form of targeted immunity has led to the development of technologies that could lead to a radical change in the way that human diseases are treated. The breakthrough began in the early years of bacterial genome sequencing when researchers noticed ‘an unusual structure’ that contained short, repetitive DNA sequences in the Escherichia coli chromosome2. Subsequent studies identified more of these structural motifs in other prokaryotes3, and in 2005 the sequences between the repeats, termed clustered regularly interspaced short palindromic repeats (CRISPRs), were analysed and found to be exact matches to phage genomes4. Further analyses of the regions upstream and downstream of these repeat-spacer loci identified a group of coding genes that often co-localized at the CRISPR arrays. These coding genes were named CRISPR-associated (Cas) proteins5,6. A 2007 study reported that yogurt-fermenting bacteria (Streptococcus thermophilus) expressing Cas proteins and a CRISPR array containing spacers that matched a phage genome were protected from infection by the phage7. Notably, a single protein, CRISPR-associated protein 9 (Cas9), was identified as being solely responsible for RNA-mediated DNA cleavage in certain bacteria7.
The mechanism of this CRISPR-mediated phage protection was characterized through detailed biochemical work8. The CRISPR array is transcribed as a single RNA and then processed at the repeats into shorter CRISPR RNAs (crRNAs) that each contain a single spacer. The crRNAs hybridize with a small trans-activating CRISPR RNA (tracrRNA) and can then be recognized and bound by Cas9 to create a ribonucleoprotein (RNP) complex. The RNP complex associates with a phage genome, searching for sequences that match the spacer encoded on the crRNA. Once homology is found, Cas9 acts as a nuclease, creating a double-strand break (DSB) by cutting the DNA and thereby inhibiting the phage life cycle.
This simple mechanism was immediately recognized as a promising tool for editing DNA and curing disease. To simplify CRISPR–Cas9 and make it more amenable to gene editing, the crRNA and tracrRNA were fused into a single guide RNA (sgRNA) to create a two-component system: the Cas9 protein creates the DSB and the sgRNA guides the nuclease to a user-defined genomic site9 (Fig. 1). In mammalian cells, the system was first used to harness natural DNA repair mechanisms to perform gene editing via the more efficient non-homologous end joining (NHEJ) and less efficient homology-directed repair (HDR) processes10. NHEJ leads to error-prone indel (insertion and deletion) formation, whereas HDR is often a more desired therapeutic outcome owing to its precise manner of editing. The predominant existence of the NHEJ pathway for repairing CRISPR-induced DSBs led most early efforts to focus on knocking out mutant genes that have harmful effects in monogenic Mendelian diseases. However, many diseases cannot be treated with a simple gene knockout and require more nuanced genome engineering11,12,13,14.
a | CRISPR–Cas9 evolved as a prokaryotic adaptive immune system to protect against phages and other mobile genetic elements. The prokaryotic genome encodes a CRISPR array that contains spacers — short pieces of DNA that have exact homology to the genome of the invading pathogen — separated by repeats. Once transcribed, the array is processed into short CRISPR RNAs (crRNAs), each containing one spacer. The crRNAs duplex with trans-activating CRISPR RNAs (tracrRNAs) to create the secondary structure needed to interact with Cas9 and form a ribonucleoprotein (RNP) complex. In prokaryotes, the Cas9 RNP surveys the cell and binds to the phage genome. Cas9 cuts the phage DNA, creating a double-strand break (DSB) and disrupting the pathogen’s life cycle. b | To import CRISPR–Cas9 into other organisms or cells, the crRNA and tracrRNA are fused into a single guide RNA (sgRNA) that encodes a spacer targeting the genome at a defined site. The sgRNA together with Cas9 can be delivered as DNA via a viral vector or as RNA or protein via a lipid nanoparticle. In mammalian cells, Cas9 RNP creates a DSB and induces DNA repair pathways to generate nucleotide insertions and deletions (indels), leading to a gene edit that can potentially be used to treat disease. AAV, adeno-associated virus.
In this Review, we focus on the applications of CRISPR to potentially treat diseases that cannot be overcome by inducing frameshifts or premature stops in coding genes. We provide an overview of Cas protein engineering and CRISPR systems beyond Cas9 that create a toolbox to engineer the human genome. We then detail how each of these tools might be uniquely leveraged to create new therapies for diseases that have yet to be cured by other forms of medicine.
CRISPR gene editing
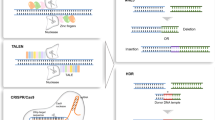
Mammalian cells have evolved a pathway to repair DSBs by ligating damaged strands predominantly through NHEJ15,16. During this DNA repair process, nucleotides are inserted or deleted (indels), leading to nearly random mutations in the ligated DNA sequence and creating a permanent edit to the genome. Decades of research developing zinc finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs) demonstrated the ability to harness the NHEJ mechanism to overcome genetic diseases in mammalian cells17,18. A disease-driving locus is targeted by engineered nucleases to create indels that result in gene knockout via frameshift mutations or gene repair via a random indel correcting the mutation. Generating ZFNs and TALENs to target a precise locus requires laborious design, build and test cycles to identify amino acid substitutions that selectively bind to a desired genomic sequence. As DNA binding is difficult to accurately predict from amino acid changes, targeting a precise genomic location can be challenging19.
The simplicity and predictability of the CRISPR targeting mechanism transformed gene editing from a complicated protein engineering problem to a RNA coding problem, instantly making CRISPR an attractive tool for basic research and clinical application. This advance resulted in an explosion of CRISPR-related publications and quickly led to CRISPR-based therapies being used in preclinical studies and early clinical trials directed towards a multitude of well-defined Mendelian disorders11,12,13,14.
A few CRISPR-based therapies directed towards monogenic disorders have reached clinical trials. For example, hereditary transthyretin amyloidosis (hATTR), a rare, fatal neuropathy that affects 50,000 people worldwide, is characterized by a point mutation in the coding sequence of the transthyretin (TTR) gene that leads to destruction of the peripheral nervous system20. This nonsense mutation induces protein misfolding, leading to oligomerization of transthyretin into fibrils that accumulate in the extracellular matrix and disrupt normal cell functions. A nanoparticle-based therapy, NTLA-2001, that encapsulates mRNA that encodes Cas9 and an sgRNA that targets TTR was developed to treat this disease21. When delivered to patients, the Cas9–sgRNA RNP creates a DSB within the coding sequence of TTR, resulting in a frameshift mutation that silences the mutant gene. The early results of a phase I clinical trial suggest that NTLA-2001 can dramatically reduce the expression of TTR, which could be highly beneficial for reducing symptom progression in patients with hATTR22.
In preclinical models, CRISPR-based therapies have been used to leverage gene knockout for many indications including the treatment of cancer, metabolic disorders and neurological disorders11,12,13,14. However, most diseases are more complex than monogenic disorders and cannot be fixed by simply editing a mutated allele. To treat these diseases, point mutations need to be precisely corrected, transcription must be carefully tuned to rescue gene dosage or more nuanced editing of non-coding regions must be considered. This next generation of therapies will use novel CRISPR tools and methods, moving the field from gene editing to a broader concept of genome engineering.
The Cas toolbox
Since the initial discovery of Cas9 as a mammalian gene editor, two key advances have enabled expansion of CRISPR technology to diseases with complex drivers: importing CRISPR systems that use other Cas proteins into mammalian cells and engineering Cas molecules to enhance their functionality. Together, the available Cas molecules comprise a set of genome engineering tools that create opportunities to cure diseases beyond the limitations of wild-type Cas9. With this large toolbox, a suitable CRISPR tool can be chosen to meet the needs of the specific disease, instead of limiting potential therapies to conform to the existing capabilities of Cas9.
Naturally occurring Cas proteins
Two distinct classes of CRISPR system exist, class I and class II (Fig. 2). Class II Cas proteins, including Cas9, have their targeting and nuclease functions encoded in a single protein, spanning a large group of RNA-guided nucleases that have evolved in numerous prokaryotic species. The two most widely used Cas9 proteins were discovered in Streptococcus pyogenes (SpCas9) and Staphylococcus aureus (SaCas9). Evolution in these different environments endowed these proteins with unique traits that need to be considered when applying them as a therapy8. For example, all known DNA-targeting Cas nucleases require a targeted locus to be flanked by a specific sequence called a protospacer adjacent motif (PAM). PAMs are short segments of DNA not encoded on the crRNA that a Cas protein must recognize to begin the process of DNA melting and target binding. SpCas9 requires a flanking NGG PAM whereas SaCas9 recognizes an NNGRRT PAM. In addition to their efficiency and specificity for DNA cutting, PAM sequences are important factors that can determine the genomic space that can be targeted by a particular Cas protein. For example, the PAM of SpCas9 is simple and more frequently represented on the genome than the PAM of SaCas9, making it easier to define a target site for gene editing. The size of the Cas protein is another crucial factor that must be considered. The coding sequence of SaCas9 (3.2 kb) is significantly smaller than that of SpCas9 (4.1 kb), making SaCas9 more amenable for packaging in gene delivery vectors such as adeno-associated viruses (AAVs) that have an ~4.7 kb packaging limit.
Mining the prokaryotic metagenome has uncovered numerous Cas systems, each with their own unique capabilities that can be leveraged for therapeutic genome engineering. These systems are categorized into two classes: class I systems, which perform their nucleic acid-targeting and nuclease activity as multiple proteins, and class II systems, which have both functions encoded on one protein. a | Type I-E, also known as Cascade, is a class I double-stranded DNA (dsDNA) nuclease. The ability of type I-E to handle longer guide RNAs (gRNAs) means that it can target genomes with higher specificity than other Cas systems35. b | Type I-F can interact with TniQ, part of a transposase complex, and is capable of targeted transposition, inserting entirely new DNA sequences into the genome37. c | Type II-A, also known as Cas9, is the first CRISPR–Cas system to be used therapeutically in humans. Cas9 can use single guide RNA (sgRNA) and typically has high GC protospacer adjacent motifs (PAMs)10. d | Type V-A, also known as Cas12a, typically has high AT PAMs and therefore has access to different genomic regions than Cas9. Cas12a can process multiple CRISPR RNAs (crRNAs) from a single transcript, enabling facile multiplexed DNA targeting25. e | Type V-F, also known as Cas12f, is an extremely small nuclease (1.4–1.6 kb), which makes it more amenable to viral packaging than other Cas systems. Cas12f has high AT PAMs29. f | Type VI-D, also known as Cas13d, targets single-stranded RNA (ssRNA), enabling transcriptome modification, and has no PAM requirement34. ssDNA, single-stranded DNA.
The observation that class II systems can possess unique features prompted researchers to search the metagenome space to find new CRISPR proteins. The repetitive, palindromic feature of natural prokaryotic CRISPR loci was used to identify Cas genes from sequencing data of prokaryotes and establish a landscape of potential gene-editing proteins23,24. This work led to the discovery of Cas12a (originally called Cpf1), which can generate DSBs in the human genome25. The PAM of Cas12a (TTTV) differs from those of SpCas9 and SaCas9, enabling targeting of new genomic locations, and is smaller than that of SpCas9 (Lachnospiraceae bacterium Cas12a is ~3.7 kb). Most importantly, Cas12a can process a CRISPR array into individual crRNAs, enabling facile multiplexed targeting through expression of multiple crRNAs from a single transcript. By contrast, Cas9 multiplexing requires each sgRNA to have its own promoter, making expression of multiple sgRNAs difficult. This finding catalysed the discovery and characterization of new Cas12 proteins including Cas12b26, Cas12c27, Cas12d (previously CasY)28, Cas12e (previously CasX)28, the hypercompact Cas12f (previously Cas14, approximately 1.4–1.6 kb)29, Cas12g27, Cas12h27, Cas12i27 and Cas12j (previously CasΦ)30. Some of these systems (Cas12b31, Cas12e32, Cas12f33 and Cas12j30) have shown promise as gene editors in human cells.
In 2018, a group of class II proteins, known as Cas13, with the ability to bind and cleave single-stranded RNA were discovered34. The Cas13 mechanism results in transcript knockdown akin to RNA interference (RNAi), enabling the destruction of specific mRNA and enabling targeted changes to the transcriptome. Furthermore, many Cas13 proteins can flexibly target the transcript without being restricted by the need for a specific flanking sequence. This feature makes Cas13 a versatile tool for changing the phenotype of a cell without creating heritable changes to the genome.
Class I CRISPR–Cas systems segregate their targeting and nuclease functions into multiple proteins35. For example, the Cascade complex contains multiple subunits of Cas5, Cas6, Cas7, Cas8 and Cas11 that bind crRNA and direct the complex to target DNA. The complex further recruits Cas3 to perform the nuclease function. The large, multi-component nature of class I systems imposes a challenge for delivery and expression that reduces their utility in human cells. However, the longer crRNA (and therefore potential for increased specificity) and the vast diversity of class I systems (more than 80% of known CRISPR systems belong to class I) make them an attractive option for gene editing. For example, Cascade has been used to create long-range genomic deletions in human embryonic stem cells36.
Multiple Cas proteins that are capable of targeted genomic insertions have also been discovered. For example, a class I system (type I-F) and a class II system (Cas12k) can knock in DNA fragments to a specific site by CRISPR-mediated recruitment of transposition machinery. This ability to create targeted insertions has been demonstrated both in vitro and in prokaryotic hosts37,38. Although Cas-mediated transposons have yet to be imported into human cells, these and the other Cas systems described demonstrate the vast diversity and biological functions of CRISPR systems.
Engineered Cas proteins
As CRISPR evolved in prokaryotes24, most Cas systems do not perform optimally when expressed in a more complex genomic environment in human cells, resulting in poor editing efficiency or specificity. Advances in protein engineering enabled the development of enhanced Cas proteins using techniques such as structure-guided mutations, directed evolution and phage-assisted evolution39,40. For example, structure-guided engineering of the DNA-binding pocket of Cas12a and Cas12f increased indel frequency, resulting in more efficient human genome editors33,41,42. Notably, Cas12f has been engineered to generate a hypercompact class of Cas effectors (~1.4–1.6 kb) that are more amenable than other Cas proteins for in vivo delivery and expression33.
To expand Cas applications beyond genome editing, we removed the catalytic activity of SpCas9 to generate a nuclease-dead version of the protein termed dCas9 (ref.43). This engineering converts Cas9 from an RNA-guided nuclease into an RNA-guided binding protein. In E. coli, targeting dCas9 to a coding sequence or its promoter did not cut the gene but inhibited transcription by blocking RNA polymerase. This approach, termed CRISPR interference (CRISPRi), has profoundly changed the way that CRISPR is used and catalysed a series of CRISPR technologies for gene editing and gene regulation43.
Translating CRISPRi into mammalian cells by targeting dCas9 to a coding sequence is typically not sufficient to block transcription. To achieve gene knockdown in mammalian cells, dCas9 is fused with a repressor domain, for example, the Krüppel-associated box (KRAB), to induce local gene repression when brought in proximity to a specific locus44 (Fig. 3a). Here, dCas9 targets a precise genomic location, localizing the fused KRAB domain to silence gene expression only where the dCas9–KRAB fusion protein is bound. Since this discovery, numerous fusions have been generated, creating an expanded toolbox for epigenome engineering. Nuclease-dead Cas proteins have been fused to transcriptional activators to upregulate specific genes in a method termed CRISPR activation (CRISPRa). The first example of CRISPRa fused VP64, four tandem repeats of the herpes simplex virus VP16 domain that induces transcription, to dCas9 and targeted regions proximal to promoters to upregulate specific genes in mammalian cells44. Subsequently, dCas9 was fused with a myriad of transcriptional activator domains including RTA, VP64, HSF1 and p65, to enable highly specific gene upregulation in multiple cell types44,45,46,47 (Fig. 3b). Epigenetic DNA-modifying domains including the DNA methylation domains DNMT3A and DNMT3L, as well as the DNA demethylation domain TET can also be fused to dCas9. In addition, histone modifiers that write H3K27 acetylation or methylation, H3K4 methylation, H3K9 methylation or H3K79 methylation, can be fused individually or in combination to write or erase changes in the histone epigenome. These CRISPR-mediated epigenetic modifications can be used to reprogramme the transcriptome and achieve novel functions such as prolonged targeted gene silencing or activation compared with traditional CRISPRi or CRISPRa48 (Fig. 3c–e).
Catalytic residues in Cas proteins can be mutated to render the protein nuclease dead (dCas). Fusing dCas to other protein domains endows novel functionality that can be targeted to precise locations on the human genome. a | dCas fused to transcriptional repressors such as the Krüppel-associated box (KRAB) generates CRISPR interference (CRISPRi), which is capable of targeted gene downregulation lasting for as long as the fusion is present44. b | Fusion of dCas to transcriptional activators such as VP64 generates CRISPR activation (CRISPRa), which enables precise gene upregulation while the fusion protein is bound45. c | Targeted DNA methylation enabled by fusion of DNA methyltransferases such as DNMT3 to dCas results in long-term and even heritable gene repression that is independent of the fusion protein being bound to the genome48. d | Natural or engineered DNA methylation can be removed by fusing methylcytosine dioxygenases such as TET to dCas48. e | Other targeted epigenetic marks can be generated by fusing histone methyl or acetyl transferases such as p300 to dCas to modify histone residues leading to long-term, stable upregulation or downregulation of targeted genes48. f | Fusing dCas or nickase Cas (nCas), which creates single-stranded DNA breaks, to the cytosine deaminase APOBEC1 converts local cytosines into uracil, which is later converted into thymine52,53,54,55,56. g | Fusion of adenosine deaminases such as TadA to dCas or nCas leads to the local conversion of adenosine into inosine, which is resolved as guanine57,58,59,60. h | Prime editing uses a long prime editing guide RNA (pegRNA) that binds to a nicked DNA strand. This creates the starting conditions for the reverse transcriptase (RT) fused to nCas9 to write the genetic information encoded on the pegRNA directly on the genome. The pegRNA can be designed to enable use of prime editing to create large insertions or deletions63.
Other CRISPR–Cas systems can also be mutated into dCas systems to take advantage of their unique properties. For example, the crRNA processing feature of dCas12a enables highly multiplexed CRISPRa or inducible and logic-gated gene regulation49,50. dCas13 fused to RNA-modifying domains such as the ADAR deaminase domain that converts A into I enables targeted coding or epitranscriptome changes to study their effects on cellular phenotype51.
Beyond transcriptional regulation or epigenetic engineering, Cas proteins can be fused to nucleotide modifiers to enable precise gene editing. Although DSB-mediated indel formation is useful for gene knockout, the random nature of indels makes this mechanism difficult to harness for precise mutation correction. To overcome this limitation, DNA base-editing enzymes have been fused to dCas and to a mutated nickase version of Cas (nCas) that generates a single-stranded break. For example, when fused to dCas9, the cytidine deaminase enzyme APOBEC1 makes a targeted C-to-U conversion on the DNA strand that is not bound by sgRNA. This U is read as a T upon DNA replication, creating a precise C-to-T mutation52 (Fig. 3f). Cytosine base editor (CBE) systems have been greatly improved by switching dCas9 for nCas9, fusing uracil DNA glycosylase (UGI), mutating or homologue swapping APOBEC1 and linker optimization52,53,54,55,56.
Cas9 fused to an E. coli TadA that was optimized via protein evolution can deaminate A into I to create an A-to-G conversion57 (Fig. 3g). Adenosine base editors (ABEs) have been improved through subsequent rounds of TadA-directed and phage-assisted evolution, addition of extra TadA domains, and improved codon usage and nuclear localization57,58,59,60. To generate simultaneous C-to-T and A-to-G conversions at a single site, both cytosine and adenosine deaminases can be fused to nCas9 (refs.61,62).
Base editors can create specific mutations at regions close to the Cas binding region but are not sufficient to make multiple base pair insertions, deletions or mutations beyond A-to-G or C-to-T. Furthermore, if many As or Cs are present around the target site, they might inadvertently become mutated, creating off-target effects. To overcome this limitation, a method to insert longer stretches of DNA has been developed, termed prime editing63 (Fig. 3h). Here, nCas9 is fused to a reverse transcriptase and the sgRNA is elongated at the 3′ end to encode both the desired insertion or deletion sequence and a priming region complementary to the nicked DNA (collectively called prime editing guide RNA (pegRNA)). The pegRNA primes the nicked strand and the reverse transcriptase converts the desired edit into DNA directly on the genome, creating a targeted insertion or deletion in place. Prime editing was originally used to make all 12 base-to-base conversions, insertions of up to 44 bp and deletions from 1 to 80 bp. Using two pegRNAs, larger gene replacement or excision strategies can be created, enabling targeted insertion of sites such as Bxb1 recombinase sites for large genomic insertions (up to several kilobase pairs)64.
Targeting the non-coding genome
In most cases, the ablation of genes through NHEJ-induced indels to cause a frameshift or to introduce a premature stop codon does not sufficiently overcome drivers of diseases. Many diseases are driven by mutations in the non-coding regions of the genome, such as promoters or enhancers that regulate transcription or introns that affect mRNA splicing and protein translation. These non-coding sites present new therapeutic opportunities to manipulate gene expression instead of changing the primary sequence of the gene. Most of the human genome is non-coding, and CRISPR can make genetic and epigenetic changes in this vast space to affect gene regulation.
Introns
mRNAs contain regulatory elements that can modulate translation but do not themselves code for protein. Mutations in intronic regions can result in improper splicing of pre-mRNA, leading to mistranslated proteins that can lead to disease. For example, a rare form of childhood blindness known as Leber congenital amaurosis type 10 (LCA10) is characterized by an intronic point mutation in CEP290 that results in dysfunctional photoreceptors and ultimately retinal degeneration65. This mutation leads to aberrant splicing in the CEP290 pre-mRNA that introduces a premature stop site, resulting in a truncated protein and loss of function. The addition of CEP290 complementary DNA (cDNA) as a gene therapy could correct gene dosage and in theory cure the disease; however, the large size of CEP290 protein (2,479 amino acids) makes it impossible to package into viral vectors such as AAVs and therefore prevents its use as a gene therapy. To overcome this problem, an AAV5-based therapy known as EDIT-101 that encapsulates SaCas9 and two sgRNAs targeting genomic locations upstream and downstream of the intronic CEP290 point mutation is being developed66. The two sgRNAs enable cutting around the mutation to induce its removal or inversion and thereby restore normal splicing of CEP290 pre-mRNA (Fig. 4a).
CRISPR technologies can be used to edit non-coding regions of the human genome. a | Proper intron splicing can be restored by creating targeted cuts around an intronic mutation (Mut). These cuts lead to deletion or inversion of the mutated region and thereby correct the mRNA66,68,69,70. b | Entire exons carrying deleterious mutations can be skipped by modifying the flanking introns72,73,74. c | The expression of a gene can be indirectly increased by knocking out the expression of its inhibitory transcription factor, for example, by indel formation in the enhancer region of the transcription factor82,83,84. d | Single or dual guide RNA (gRNA) strategies that target Cas proteins around a transcriptional start site cause the region to be deleted, leading to a reduction in gene expression86,87,88. e | Editing long non-coding RNA (lncRNA) can modulate gene dosage. For example, in settings in which a lncRNA silences a paternal allele, leaving only a mutant maternal allele, the generation of indels in the lncRNA can disrupt its function, leading to the therapeutic expression of the wild-type paternal allele91,92. f | To increase the concentration of a therapeutically relevant gene, Cas proteins can be used to form indels in the microRNA (miRNA) or the 3′ untranslated region, leading to a reduction in miRNA binding and amelioration of the resulting RNA interference101,102,103,104,105,106,107,108.
Intron targeting has also been used in preclinical studies to correct the genetic blood disorder β-thalassaemia, which is caused by a myriad of mutations in HBB. One of the most common disease-causing HBB mutations, particularly in Southeast Asian populations, is a point mutation in intron 2 (IVS2-654) that alters splicing67. Cas9 has been targeted to the aberrant intron to restore HBB gene expression in induced pluripotent stem cells (iPSCs) in vitro, creating a potential avenue for cell therapy through haemopoietic stem cell replacement68. Similarly, CRISPR–Cas9 targeted to intron 16 of LZTR1 can overcome the disease phenotype associated with Noonan syndrome-associated cardiomyopathy in iPSC-derived cardiomyocytes in vitro69. This type of intronic targeting has also been used in vitro to correct a rare mutation in CFTR (affecting about 2,000 patients worldwide) that leads to cystic fibrosis70.
CRISPR–Cas can be used to delete entire exons by taking advantage of a process known as exon skipping. Here, Cas9 is targeted to introns flanking a mutated exon to alter pre-mRNA processing and cause the aberrant exon to be spliced out, thus maintaining an intact open reading frame but removing the mutation (Fig. 4b). This approach has been applied in vitro to Duchenne muscular dystrophy, a muscle-wasting disease characterized by a mutation in the dystrophin gene (DMD) that results in the deletion of exon 50 (ref.71). To correct this deletion, exon skipping can be harnessed by targeting the splice acceptor site at exon 50, the introns flanking exon 51 or the introns surrounding exons 45–55 of DMD in human myoblasts72,73. In all cases, the loss of exon 51 restores DMD and overcomes the disease pathology. Exon skipping by dual targeting CRISPR–Cas9 can also be used to target fusion oncogenes. This approach was used to create a cancer-specific therapy that controlled tumour burden in a mouse xenograft model74.
Untranslated regions
mRNAs contain regulatory elements in the untranslated regions (UTRs) that flank their translational start and stop sites. These regions perform a myriad of regulatory functions such as initiating and terminating translation, altering RNA trafficking and stability, interacting with RNA-binding proteins or microRNAs (miRNAs), and controlling post-transcriptional modifications. Targeting of UTRs to overcome disease pathologies has been demonstrated in vitro. For example, expansion of CTG repeats within the 3′ UTR of DM1 protein kinase (DMPK) from 5–38 repeats in healthy cells to more than 50 repeats in mutated cells causes a neuromuscular disorder known as myotonic dystrophy type 1 (DM1)75. Targeted deletion of the CTG repeats by CRISPR–Cas led to loss of the aberrant mRNA transcripts in DM1 neural stem cells76.
In iPSCs from patients with Duchenne muscular dystrophy, upregulation of the dystrophin-related gene utrophin (UTRN) can circumvent the effects of DMD loss of function, but miRNAs that bind to UTRN destroy the transcript. Cas9 targeting of the miRNA binding sites led to upregulation of the utrophin protein and overcame the disease phenotype77 (Fig. 4c).
Expression of the huntingtin gene (HTT) leads to the neural degeneration that is associated with Huntington disease. To knock out this gene in a mutation-independent manner, Cas9 can be targeted to the HTT 5′ UTR, leading to improper maturation of the transcript and reducing the expression of the disease-causing allele78. Similarly, disruption of a 52 bp regulatory element in the 3′ UTR of amyloid precursor protein led to a substantial reduction in the disease-inducing amyloid-β peptide (Aβ) in a mouse model of Alzheimer disease79.
Cis-regulatory elements
Cis-regulatory elements, including promoters, enhancers and silencers, are important regulatory regions that modulate coding genes to control and alter their expression. Creating indels in these regions disrupts their function and can be used to correct gene dosages that drive disease. For example, sickle cell disease (SCD) and transfusion dependent β-thalassaemia (TDT) are monogenic diseases caused by mutations in HBB80,81 that result in malformation and loss of function of HBB protein. The γ-globin genes (HBG1 and HBG2) that encode fetal haemoglobin (HbF) have the same function as HBB but are silenced by the transcription factor BCL11A during maturation into adulthood. An ex vivo gene-editing technique, CTX001, has been developed that reduces expression of BCL11A to upregulate HBG in autologous haematopoietic stem and progenitor cells82. This technique uses CRISPR editing of the BCL11A enhancer to reduce gene expression rather than complete ablation of BCL11A, which would lead to other pathologies83,84 (Fig. 4a). This ‘one size fits all’ therapeutic strategy has the potential to benefit more patients than specific strategies that each correct one of the myriads of individual mutations in HBB.
The therapeutic potential of editing cis-regulatory elements is also being investigated in other disorders. For example, the mutated transcription factor FOXA1 is an oncogene with a role in the onset and progression of prostate cancer85. Targeting transcription factor binding elements in the FOXA1 promoter modulates the function of these elements, reducing expression of the gene and inhibiting prostate cancer cell growth in vitro86. In addition, a dual sgRNA approach has been used in vitro to excise a 44 kb promoter region upstream of a mutant HTT gene to silence its expression and thereby ablate expression of the Huntington disease-causing variant87 (Fig. 4d). Similarly, a dual targeting approach that binds once in the promoter and once in the first intron of HTT removes the transcriptional start site and first exon, inhibiting gene expression88.
Non-coding RNAs
Some non-coding RNAs affect gene expression by binding to mRNA through Watson–Crick base pairing, which creates another avenue to alter gene expression through gene editing89. For example, miRNAs can bind to UTRs and target them for destruction. Long non-coding RNAs (lncRNAs) can act through several mechanisms including activating or inhibiting transcription or translation, altering splicing or remodelling chromatin epigenetics. Altering the primary sequences of the non-coding elements ablates their function and downstream effects on gene expression.
Angelman syndrome is a neurodevelopmental disorder caused by a maternally inherited mutant UBE3A gene that could be rescued by expression of the paternal allele90. However, the paternal allele is silenced by the lncRNA UBE3A-ATS. CRISPR–Cas9 targeting of UBE3A-ATS ablated its function, leading to expression of the paternal UBE3A gene and rescuing the disease phenotype in cultured human neurons and in a mouse model of the disease91,92 (Fig. 4e).
CRISPR-mediated disruption of lncRNA through indel formation has been widely investigated in vitro in cancer to reduce cell growth93,94,95,96,97,98 and overcome metastasis96,99. CRISPR-mediated mRNA knockdown using the Cas13 system has been used to cleave lncRNA and inhibit bladder cancer proliferation in vitro and in vivo100.
Muscular atrophy is in part controlled by expression of miR-29b101. Delivery of Cas9 that targets miR-29b in multiple mouse models of muscular atrophy knocked out the miRNA and prevented muscle loss102 (Fig. 4f). In macrophage cell lines, CRISPR-mediated indel formation in miR-155 reduced pro-inflammatory cytokine expression in vitro, creating an avenue for treating the autoimmune disease rheumatoid arthritis103. miRNA targeting has also been extensively studied in cancer104; various approaches have been shown to lead to reductions in cancer cell proliferation105,106,107, inhibition of metastasis105,106 and death of cancer cells108 in preclinical studies.
Transcriptional and epigenetic modulation
Epigenetic changes and dysregulated expression that alters gene dosage drive many diseases, resulting in phenotypes that cannot be rescued simply by indel formation or microdeletion. Dysregulation can also be caused by an epigenetic change that is inaccessible to wild-type CRISPR systems. To fill this gap, dCas molecules can be creatively coupled to transcriptional or epigenetic modulators, to precisely target relevant therapeutic regulatory domains to specific regions of the genome without creating DNA damage or a DNA edit. This approach also mitigates the risks associated with DNA damage, p53-induced apoptosis, permanent off-target editing and abnormal chromosomal rearrangements.
CRISPR interference
dCas9–KRAB fusions can be targeted to protein-coding sequences to downregulate transcription, repressing the gene for as long as the CRISPR fusion protein is present without permanently editing DNA44,109. This feature is attractive for reducing expression of the voltage-gated sodium ion channel NaV1.7 (encoded by SCN9A) in the peripheral nervous system, which could reduce pain and thereby overcome the current reliance on opioids110. As developing small-molecule drugs is challenging and complete gene ablation would result in permanent undesirable pain insensitivity, CRISPRi is an attractive option to treat chronic pain111. CRISPRi targeted to NaV1.7 and delivered intrathecally reduced pain sensitivity and reversed chronic pain in mouse models of carrageenan-induced inflammatory pain, paclitaxel-induced neuropathic pain and BzATP-induced pain, demonstrating the therapeutic advantage of CRISPRi over traditional CRISPR editing in these settings111.
The regulatory effects of CRISPRi can be used in many other diseases in which complete CRISPR-mediated gene knockout is not therapeutically useful. In one form of long QT syndrome (LQTS) that can be caused by a myriad of mutations in CALM2, dCas9–KRAB was used to reduce expression of the mutant gene in vitro112. This intervention overcame the disease phenotype in iPSC-derived cardiomyocytes and creates a generalizable therapeutic approach that is independent of the location of the nonsense mutation. In a mouse model of retinitis pigmentosa, dCas9–KRAB targeted to Nrl rescued retinal function when delivered to postmitotic cells that normally have reduced capacity for the DNA repair mechanisms that are essential for indel formation113. Overexpression of DUX4 in myocytes leads to facioscapulohumeral muscular dystrophy (FSHD)114,115. DUX4 has many genomic copies that could lead to toxicity if numerous DSBs were created, and gene editing at such large repetitive regions can lead to unpredictable outcomes. CRISPRi has been leveraged in vitro and in vivo to reduce DUX4 expression without the risk of inducing apoptosis owing to DNA damage. In contrast to CRISPR gene editing, CRISPRi can be inducible and reversible, which further alleviates safety concerns when testing in the clinic.
CRISPR activation
Fusion of dCas proteins to activators provides a method for targeted gene upregulation that can overcome various types of disease including those caused by haploinsufficiencies44. Unlike ectopic transgene expression, CRISPRa can be used to precisely tune the magnitude of gene upregulation. In addition, this system can be packaged into viral vectors more easily than larger transgenes. For example, nuclease-dead SaCas9 (SadCas9) was fused to the VP64 domain and delivered to mouse models of obesity that had a haploinsufficiency of either Sim1 or Mc4r116. Targeting CRISPRa to the promoter region increased transcription of both genes, rescuing the obesity phenotype and demonstrating cell specificity by precisely targeting tissue-specific cis-regulatory elements.
CRISPRa can upregulate genes independently of mutations. Multiple LAMA2 mutations lead to congenital muscular dystrophy type 1A (MDC1A), which can be rescued by ectopic expression of LAMA1. AAV-based delivery of SadCas9 fused to VP64 was used to upregulate Lama1 in a mouse model of MDC1A, improving muscle fibrosis and preventing disease progression117. The ability to overcome muscle wasting in a mutation-independent manner has also been used to overcome the Duchenne muscular dystrophy phenotype in vitro through upregulation of Lama1 (ref.118) or a utrophin gene (UTRN)119. Cas9 expressed in mice with a sgRNA containing an aptamer that recruits p65 and HSF1 domains47 was able to upregulate genes to treat Duchenne muscular dystrophy (Klotho or Utrn), acute kidney injury (Il10 or Klotho) and type 1 diabetes (Pdx1)120. Importantly, the sgRNA had a spacer of 14 bp instead of 20 bp, which allowed Cas9 to bind to DNA but not to create DSBs, resulting in a nuclease-deficient system.
Use of CRISPRa to upregulate therapeutically useful coding genes has been demonstrated in vitro for autoimmune diseases121, neurodegenerative diseases122,123 and cancer124,125,126,127. However, the usefulness of this approach extends beyond upregulation of single proteins to endogenous non-coding RNA. For example, dCas9 fused to VP64, p65 and RTA (collectively known as VPR) has been used in vivo to increase expression of DANCR, a lncRNA that increases bone regeneration through chondrogenic differentiation128. CRISPRa can also be used to upregulate multiple gene targets by the addition of multiple sgRNAs. For example, CRISPRa was demonstrated to simultaneously upregulate Bdnf, Gdnf and Ngf in adipose-derived stem cells ex vivo to promote peripheral nerve regeneration in a rat model of nerve injury129.
Traditional CRISPRi and CRISPRa constructs are large and challenging to package into single AAVs; however, the development of hypercompact Cas molecules can overcome this issue. For example, structurally guided engineering of a natural Cas12f system reduced the size of Cas by almost 60% (2.6 kb) to produce a miniature Cas system (CasMINI, ~1.55 kb)33. CasMINI can be fused to many commonly used activating or repressive modulators to create proteins that are much smaller than the 4.7 kb packaging limit of AAV vectors for in vivo delivery. These hypercompact systems can also be encoded on mRNA for more efficient delivery and expression in human tissues and in vivo.
CRISPR epigenetic modification
CRISPRa and CRISPRi gene regulation methods result in transient gene modulation. In postmitotic cells or disease indications in which transient gene expression results in a therapeutic benefit, this transience does not present a challenge. However, some diseases require long-lasting and heritable changes to gene regulation. Epigenetic modifications via targeted addition of methyl groups to DNA or insertion of acetyl or methyl groups on histone residues locally modulate gene expression130. These modifications are often persistent and can be inherited by daughter cells, creating an opportunity for long-lasting gene expression modulation. Many epigenetic modifiers have been fused to CRISPR proteins to make chemical modifications at the DNA or chromatin level48. For example, CRISPRoff131 and CRISPR-KAL109 can lead to long-term (for example, several months) gene silencing by modifying H3K9me3 and DNA methylation. These approaches are potentially suitable for treating diseases that require persistent gene perturbation.
DNA methylation domains from the DNMT3 family have been fused with dCas9 to achieve long-term gene silencing132,133,134. For example, targeting the SNCA intron 1 with a dCas–DNMT3 fusion protein generated targeted DNA hypermethylation in human iPSC-derived dopaminergic neurons carrying a SNCA triplication and rescued the Parkinson disease-related phenotype in vitro135. To reverse the silencing effects of natural DNA methylation, ten-eleven translocation methylcytosine dioxygenase 1 (TET1) catalytic domain was fused with dCas9 to selectively remove DNA methyl groups and upregulate gene expression136,137. This approach has been investigated as a potential therapy for fragile X syndrome, which is an intellectual disability caused by a CGG expansion in FMR1 that results in extensive methylation and therefore reduces gene expression138. Targeting of dCas9–TET1 to FMR1 demethylated the CGG repeats, reactivated sustained gene expression and rescued the disease phenotype in iPSC-derived neurons in vivo139. Fusion proteins comprising dCas9 and TET enzyme catalytic domains have also been used to treat cancer in vitro (targeting BRCA1 (ref.140)) and in vivo (targeting SARI141) and attenuate renal fibrosis in vivo (targeting Rasal1 or Klotho)142. Importantly, the resulting DNA methylation changes are long-lasting, heritable and reversible143.
To site-specifically modify histones, the catalytic core of the p300 domain was fused to dCas9. When directed to enhancer regions on DNA, this fusion adds an acetyl group to lysine 27 of histone H3 (H3K27ac), resulting in activation of gene expression144. In mice, expression of dCas9–p300 was able to upregulate Foxp3 expression in T cells, converting them into regulatory T (Treg) cells with the potential to treat autoimmunity145,146. The H3K27ac mark can be removed using dCas9 fused to histone deacetylase 1 (HDAC1). This approach has been targeted to KRAS to inhibit cancer growth147. Additional suppressive CRISPR histone modifiers include decreasing H3K4 methylation, increasing H3K9 methylation and enhancing HP1α binding, which when targeted to GRN can reduce cell proliferation and invasion in hepatoma cells148.
Base and prime editing
The random process of indel formation is difficult to harness to correct precise mutations, as the number or identity of the added nucleotides cannot be controlled. With the exception of cell therapies in which engineered cells are clonally expanded, checked for proper mutation and then reintroduced into the body, wild-type CRISPR systems are often poor choices for precise mutation correction. To fill this gap, CRISPR fusions that make precise genetic changes have been generated and deployed in a myriad of diseases.
Base editing
Given the rapid improvement in the technology and ability to correct deleterious point mutations with unparalleled precision, base editors have been quickly adopted as potential approaches to treat well-understood diseases with known missense mutations. CBEs that create C-to-T mutations have been used in a wide variety of in vivo models. Both Cas9 and Cas12a CBEs have been used to correct a missense mutation in the Pah gene in a mouse model of the human autosomal recessive liver disease phenylketonuria (PKU)149. The ability to make a C-to-T conversion enables the generation of stop codons, which always begin with a thymine. In a mouse model of amyotrophic lateral sclerosis (ALS), SpCas9 CBE was used to create a premature stop codon in SOD1, reducing muscle atrophy and improving neuromuscular function150. The large size of the CBE CRISPR constructs necessitated the protein to be split into two AAV vectors and fused post-translationally in the cell using inteins149,150.
ABEs are highly relevant for therapeutics, as C•G to T•A transitions account for approximately half of all known pathogenic point mutations57. In a Duchenne muscular dystrophy mouse model, ABEs that were delivered to the muscles as two AAVs were able to correct a single mutation in Dmd and improve the disease phenotype151. ABEs have also been used to correct the LMNA mutation in a mouse model of Hutchinson–Gilford progeria syndrome, extending the median lifespan from 215 to 510 days152. Notably, use of an ABE to correct a nonsense mutation ex vivo in a mouse model of sickle cell disease led to approval of the BEACON-101 phase I/II trial of this therapy153,154. Other therapeutic uses of base editors have been reviewed elsewhere155,156,157.
Prime editing
Prime editing has the potential to create a wide array of therapeutic genome edits but has not yet been as widely investigated as other CRISPR systems. In the study that first described the tool, researchers corrected mutations in HBB that cause sickle cell disease, HEXA that cause Tay–Sachs disease and PRNP to protect against prion diseases63. Prime editing can be used to make precise mutations that are currently not possible using base editors. For example, in a mouse model of α1-antitrypsin deficiency (AATD), prime editors were effectively delivered to mice livers to remove a pathogenic E342K mutation in SERPINA1 by creating an A-to-G edit158. Prime editors have also been used to correct a mutation in Dnmt1 in mouse retinas by creating a G-to-T transversion, demonstrating the potential to correct eye disease. These precise edits could not be achieved using other CRISPR tools159.
In addition to base editing, prime editing can be used to insert oligonucleotides. In human iPSCs, prime editing was used to insert two nucleotides (AC) into exon 52 of DMD. This approach enabled exon reframing to rescue expression of DMD and the contractile function of iPSC-derived cardiomyocytes modelling Duchenne muscular dystrophy160. Although the therapeutic use of prime editors is still in its infancy, the flexibility of genomic edits that this methodology creates potentially enables correction of a myriad of diseases.
Infection prevention and treatment
In addition to modifying the human genome, CRISPR–Cas therapies can be used to target latent and chronic viral infections in human cells. For example, intrastromal injection of Cas9 as a non-integrating lentivirus prevented herpes simplex virus type 1 (HSV-1) infection and disease pathology and destroyed the viral reservoir in mouse models161. This system can also be used for other herpes viruses, such as Epstein–Barr virus (EBV), by targeting Cas9 to essential promoters162 or coding sequences163 in the viral genome. Cas9 and Cas12a have both been used to target the long terminal repeats and Gag–Pol polyprotein of HIV-1 (refs.164,165,166,167,168). Cas9 has also been used to target coding sequences and the covalently closed circular DNA in hepatitis B virus (HBV)169,170,171,172,173,174 and to cut the DNA genome of human papillomavirus (HPV)175,176,177. In addition, Cas9 with a modified sgRNA was used to destroy the RNA genome of hepatitis C virus (HCV)178. Notably, a CRISPR-based strategy to clear HIV infections (EBT-101) has entered a phase I/II clinical trial179. Furthermore, our laboratory has developed a strategy using Cas13d to target viral RNA genomes and demonstrated the utility of this approach as a prophylactic for both influenza A virus (IAV) and sever acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection180. Notably, this strategy works on a broad spectrum of coronaviruses and variants of SARS-CoV-2 owing to the ability to target highly evolutionarily conserved regions in the viral genome181.
CRISPR has also been used to target bacterial infections. Cas9 can be packaged into bacteriophages and delivered to antibiotic-resistant S. aureus to target bacterial resistance genes and re-sensitize the bacteria to treatment182. As DSBs in bacterial genomes result in cell death, plasmid or phage-delivered Cas9 is an effective antimicrobial strategy in both E. coli and S. aureus183,184,185. Cas3 has been harnessed to target and shred (that is, create long-range deletions) the genome of Clostridiodes difficile (also known as Clostridium difficile), which is one of the most harmful and antibiotic-resistant bacterial species in existence186. The vast therapeutic CRISPR toolbox is rapidly expanding beyond human genome engineering to treat a wide variety of infectious diseases.
Challenges of delivering CRISPR tools
CRISPRs are multi-component systems that require packaging of the large protein, the gRNA and all the elements that control their expression. Many established approaches exist for in vitro and ex vivo delivery of these components as DNA, RNA or RNP complexes187,188. Both integrating and non-integrating lentiviruses can be used to deliver CRISPR components but are limited owing to the potential for insertional mutagenesis and low efficiency, respectively. DNA, RNA or RNP can also be delivered using methods that physically introduce the components to cells, such as electroporation or microinjection. These approaches benefit from controllable dosing and efficient delivery but can be technically difficult and create viability issues.
Many diseases that could benefit from a CRISPR therapy cannot be treated ex vivo and therefore cannot be delivered using lentiviruses, microinjection or electroporation. Delivery of CRISPR molecules in vivo poses a major challenge that has limited their potential as therapeutics187,188,189,190. AAV is commonly used to deliver CRISPR components as DNA both in vivo and ex vivo. This approach can be used to deliver small CRISPR systems in a single vector or larger components split between multiple vectors. However, the limited packaging capacity and tropism of AAVs prevent them from being universally used. Lipid nanoparticles (LNPs) can deliver CRISPR tools as RNA, resulting in more transient effects than those obtained with viral AAV delivery and therefore reducing the risk of off-target editing. However, many LNPs almost exclusively traffic to the liver and cannot reach other therapeutically relevant tissues. Virus-like particles (VLPs) are exciting vehicles for the delivery of CRISPR components. An RNA-binding protein or CRISPR RNP is fused to a retroviral Gag–Pol, enabling CRISPR RNA or RNP to be encapsulated in a viral vector. Although the therapeutic use of VLPs is still in its infancy, this approach has been demonstrated to have low levels of off-target effects and flexible tropism191,192,193.
Current CRISPR therapeutics are limited by the small packaging capacities and tissue trafficking properties of the available delivery vectors, which restrict the use of these CRISPR tools and reduce their potential disease indications. Various approaches, such as directed evolution of AAV capsids, functionalization of LNPs and molecular engineering of CRISPR components, are being investigated with the aim of improving the efficacy, safety and specificity of in vivo delivery vehicles. To realize the full potential of CRISPR therapies, further efforts are required to get these tools to the relevant tissues with high efficiency, high specificity and minimal toxicity.
Conclusions
The ease with which CRISPR can create targeted DSBs in the human genome enabled quick adoption as a broad tool to overcome genetic disorders. As a first step, CRISPR was used to perform targeted gene knockouts, as Cas9 can be targeted anywhere on the coding sequence to induce a frameshift to silence a deleterious protein. However, most diseases are complex and cannot be cured by this simple coding sequence-targeting strategy. The use of CRISPR to target diseases with complex drivers has been catalysed by developing more nuanced strategies that target the non-coding genome and fix gene expression more indirectly (for example, by exon skipping or intron corrections). Beyond these approaches, the rapid discovery of natural CRISPR molecules with beneficial properties and further engineering of these proteins to create molecules that alter transcription, change the epigenome, make precise mutations or enable writing directly on the genome have dramatically increased the range of indications that can potentially be treated using CRISPR–Cas systems. However, further advances are needed to fully leverage these proteins.
As discussed above, current CRISPR tools are limited by the challenges of in vivo delivery to the relevant tissues. In addition, off-target events caused by CRISPR systems must be precisely controlled to create highly targeted therapies. Bioinformatic strategies to improve the specificity of gRNA, altering the chemical composition and length of gRNA, the discovery and engineering of new Cas variants, temporal restriction of CRISPR systems using transient delivery methods or anti-CRISPRs, and moving from DSBs to more targeted systems such as prime editors, base editors or epigenetic modulators could greatly reduce off-target effects194,195,196. However, more research is required to develop a maximally safe and effective CRISPR therapy.
The use of CRISPR tools to target more nuanced disease drivers requires a better understanding of how non-coding DNA and epigenetic states affect a disease pathology. Point mutations in coding sequences are much easier to link to a disease phenotype than mutations in non-coding sequences owing to a deep understanding of how a genetic change results in an amino acid change by looking at sequencing information. Using the right CRISPR tool that can link sequence, epigenome, transcriptome and phenotype information to the root cause of a pathology that is not driven by simple polymorphisms will be helpful to define new cures. However, the rapid advances in CRISPR tools, multi-omic methods and delivery mechanisms suggest that genome engineering techniques will soon be developed for a multitude of diseases, potentially resulting in curative therapies for many underserved patient populations.
References
Cooper, M. D. & Alder, M. N. The evolution of adaptive immune systems. Cell 124, 815–822 (2006).
Ishino, Y., Shinagawa, H., Makino, K., Amemura, M. & Nakata, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 169, 5429–5433 (1987).
Mojica, F. J. M., Díez-Villaseñor, C., Soria, E. & Juez, G. Biological significance of a family of regularly spaced repeats in the genomes of Archaea, Bacteria and mitochondria. Mol. Microbiol. 36, 244–246 (2000).
Bolotin, A., Quinquis, B., Sorokin, A. & Ehrlich, S. D. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151, 2551–2561 (2005).
Jansen, R., Embden, J. D. Av, Gaastra, W. & Schouls, L. M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 43, 1565–1575 (2002).
Haft, D. H., Selengut, J., Mongodin, E. F. & Nelson, K. E. A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLoS Comput. Biol. 1, e60 (2005).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Jiang, F. & Doudna, J. A. CRISPR-Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Konishi, C. T. & Long, C. Progress and challenges in CRISPR-mediated therapeutic genome editing for monogenic diseases. J. Biomed. Res. 35, 148–162 (2020).
Wu, S. S., Li, Q. C., Yin, C. Q., Xue, W. & Song, C. Q. Advances in CRISPR/Cas-based gene therapy in human genetic diseases. Theranostics 10, 4374–4382 (2020).
Kotagama, O. W., Jayasinghe, C. D. & Abeysinghe, T. Era of genomic medicine: a narrative review on CRISPR technology as a potential therapeutic tool for human diseases. BioMed. Res. Int. 2019, 1369682 (2019).
Hirakawa, M. P., Krishnakumar, R., Timlin, J. A., Carney, J. P. & Butler, K. S. Gene editing and CRISPR in the clinic: current and future perspectives. Biosci. Rep. 40, BSR20200127 (2020).
Chang, H. H. Y., Pannunzio, N. R., Adachi, N. & Lieber, M. R. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat. Rev. Mol. Cell Biol. 18, 495–506 (2017).
Davis, A. J. & Chen, D. J. DNA double strand break repair via non-homologous end-joining. Transl Cancer Res. 2, 130–143 (2013).
Gaj, T., Gersbach, C. A. & Barbas, C. F. 3rd ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013).
Gupta, R. M. & Musunuru, K. Expanding the genetic editing tool kit: ZFNs, TALENs, and CRISPR-Cas9. J. Clin. Invest. 124, 4154–4161 (2014).
Khan, S. H. Genome-editing technologies: concept, pros, and cons of various genome-editing techniques and bioethical concerns for clinical application. Mol. Ther. Nucleic Acids 16, 326–334 (2019).
Adams, D., Koike, H., Slama, M. & Coelho, T. Hereditary transthyretin amyloidosis: a model of medical progress for a fatal disease. Nat. Rev. Neurol. 15, 387–404 (2019).
Gillmore, J. D. et al. CRISPR-Cas9 in vivo gene editing for transthyretin amyloidosis. N. Engl. J. Med. 385, 493–502 (2021).
US National Library of Medicine. ClinicalTrials.gov https://ClinicalTrials.gov/show/NCT04601051 (2021).
Couvin, D. et al. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 46, W246–W251 (2018).
Makarova, K. S. et al. Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Shmakov, S. et al. Discovery and functional characterization of diverse class 2 CRISPR-Cas systems. Mol. Cell 60, 385–397 (2015).
Yan, W. X. et al. Functionally diverse type V CRISPR-Cas systems. Science 363, 88–91 (2019).
Burstein, D. et al. New CRISPR–Cas systems from uncultivated microbes. Nature 542, 237–241 (2017).
Harrington, L. B. et al. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362, 839–842 (2018).
Pausch, P. et al. CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369, 333–337 (2020).
Strecker, J. et al. Engineering of CRISPR-Cas12b for human genome editing. Nat. Commun. 10, 212 (2019).
Liu, J.-J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218–223 (2019).
Xu, X. et al. Engineered miniature CRISPR-Cas system for mammalian genome regulation and editing. Mol. Cell 81, 4333–4345 (2021).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676 (2018).
Zheng, Y. et al. Endogenous type I CRISPR-Cas: from foreign DNA defense to prokaryotic engineering. Front. Bioeng. Biotechnol. 8, 62 (2020).
Dolan, A. E. et al. Introducing a spectrum of long-range genomic deletions in human embryonic stem cells using type I CRISPR-Cas. Mol. Cell 74, 936–950 (2019).
Klompe, S. E., Vo, P. L. H., Halpin-Healy, T. S. & Sternberg, S. H. Transposon-encoded CRISPR-Cas systems direct RNA-guided DNA integration. Nature 571, 219–225 (2019).
Strecker, J. et al. RNA-guided DNA insertion with CRISPR-associated transposases. Science 365, 48–53 (2019).
Liu, R., Liang, L., Freed, E. F. & Gill, R. T. Directed evolution of CRISPR/Cas systems for precise gene editing. Trends Biotechnol. 39, 262–273 (2021).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
DeWeirdt, P. C. et al. Optimization of AsCas12a for combinatorial genetic screens in human cells. Nat. Biotechnol. 39, 94–104 (2021).
Guo, L. Y. et al. Multiplexed genome regulation in vivo with hyper-efficient Cas12a. Nat. Cell Biol. 24, 590–600 (2022).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Maeder, M. L. et al. CRISPR RNA-guided activation of endogenous human genes. Nat. Methods 10, 977–979 (2013).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Nakamura, M., Gao, Y., Dominguez, A. A. & Qi, L. S. CRISPR technologies for precise epigenome editing. Nat. Cell Biol. 23, 11–22 (2021).
Campa, C. C., Weisbach, N. R., Santinha, A. J., Incarnato, D. & Platt, R. J. Multiplexed genome engineering by Cas12a and CRISPR arrays encoded on single transcripts. Nat. Methods 16, 887–893 (2019).
Kempton, H. R., Goudy, L. E., Love, K. S. & Qi, L. S. Multiple input sensing and signal integration using a split Cas12a system. Mol. Cell 78, 184–191 (2020).
Tang, T., Han, Y., Wang, Y., Huang, H. & Qian, P. Programmable system of Cas13-mediated RNA modification and its biological and biomedical applications. Front. Cell Dev. Biol. 9, 677587 (2021).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9–cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Rees, H. A. et al. Improving the DNA specificity and applicability of base editing through protein engineering and protein delivery. Nat. Commun. 8, 15790 (2017).
Arbab, M. et al. Determinants of base editing outcomes from target library analysis and machine learning. Cell 182, 463–480 (2020).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Gaudelli, N. M. et al. Directed evolution of adenine base editors with increased activity and therapeutic application. Nat. Biotechnol. 38, 892–900 (2020).
Grünewald, J. et al. A dual-deaminase CRISPR base editor enables concurrent adenine and cytosine editing. Nat. Biotechnol. 38, 861–864 (2020).
Zhang, X. et al. Dual base editor catalyzes both cytosine and adenine base conversions in human cells. Nat. Biotechnol. 38, 856–860 (2020).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Anzalone, A. V. et al. Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing. Nat. Biotechnol. 40, 731–740 (2022).
Leroy, B. P. et al. Leber congenital amaurosis due to CEP290 mutations-severe vision impairment with a high unmet medical need: a review. Retina 41, 898–907 (2021).
Maeder, M. L. et al. Development of a gene-editing approach to restore vision loss in Leber congenital amaurosis type 10. Nat. Med. 25, 229–233 (2019).
Cheng, T. C. et al. β-Thalassemia in Chinese: use of in vivo RNA analysis and oligonucleotide hybridization in systematic characterization of molecular defects. Proc. Natl Acad. Sci. USA 81, 2821–2825 (1984).
Xu, P. et al. Both TALENs and CRISPR/Cas9 directly target the HBB IVS2–654 (C>T) mutation in β-thalassemia-derived iPSCs. Sci. Rep. 5, 12065 (2015).
Hanses, U. et al. Intronic CRISPR repair in a preclinical model of noonan syndrome-associated cardiomyopathy. Circulation 142, 1059–1076 (2020).
Sanz, D. J., Hollywood, J. A., Scallan, M. F. & Harrison, P. T. Cas9/gRNA targeted excision of cystic fibrosis-causing deep-intronic splicing mutations restores normal splicing of CFTR mRNA. PLoS ONE 12, e0184009 (2017).
Bladen, C. L. et al. The TREAT-NMD DMD global database: analysis of more than 7000 Duchenne muscular dystrophy mutations. Hum. Mutat. 36, 395–402 (2015).
Ousterout, D. G. et al. Multiplex CRISPR/Cas9-based genome editing for correction of dystrophin mutations that cause Duchenne muscular dystrophy. Nat. Commun. 6, 6244 (2015).
Amoasii, L. et al. Single-cut genome editing restores dystrophin expression in a new mouse model of muscular dystrophy. Sci. Transl Med. 9, eaan8081 (2017).
Martinez-Lage, M. et al. In vivo CRISPR/Cas9 targeting of fusion oncogenes for selective elimination of cancer cells. Nat. Commun. 11, 5060 (2020).
Ranum, L. P. W. & Cooper, T. A. RNA-mediated neuromuscular disorders. Annu. Rev. Neurosci. 29, 259–277 (2006).
Wang, Y. et al. Therapeutic genome editing for myotonic dystrophy type 1 using CRISPR/Cas9. Mol. Ther. 26, 2617–2630 (2018).
Sengupta, K. et al. Genome editing-mediated utrophin upregulation in Duchenne muscular dystrophy stem cells. Mol. Ther. Nucleic Acids 22, 500–509 (2020).
Kolli, N., Lu, M., Maiti, P., Rossignol, J. & Dunbar, G. L. CRISPR-Cas9 mediated gene-silencing of the mutant huntingtin gene in an in vitro model of Huntington’s disease. Int. J. Mol. Sci. 18, 754 (2017).
Nagata, K. et al. Generation of App knock-in mice reveals deletion mutations protective against Alzheimer’s disease-like pathology. Nat. Commun. 9, 1800 (2018).
Piel, F. B., Steinberg, M. H. & Rees, D. C. Sickle cell disease. N. Engl. J. Med. 376, 1561–1573 (2017).
Galanello, R. & Origa, R. Beta-thalassemia. Orphanet J. Rare Dis. 5, 11 (2010).
Frangoul, H. et al. CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia. N. Engl. J. Med. 384, 252–260 (2021).
Luc, S. et al. Bcl11a deficiency leads to hematopoietic stem cell defects with an aging-like phenotype. Cell Rep. 16, 3181–3194 (2016).
Yu, Y. et al. Bcl11a is essential for lymphoid development and negatively regulates p53. J. Exp. Med. 209, 2467–2483 (2012).
Robinson, J., Holmes, K. & Carroll, J. FOXA1 mutations in hormone-dependent cancers. Front. Oncol. 3, 20 (2013).
Zhou, S. et al. Noncoding mutations target cis-regulatory elements of the FOXA1 plexus in prostate cancer. Nat. Commun. 11, 441 (2020).
Shin, J. W. et al. Permanent inactivation of Huntington’s disease mutation by personalized allele-specific CRISPR/Cas9. Hum. Mol. Genet. 25, 4566–4576 (2016).
Monteys, A. M., Ebanks, S. A., Keiser, M. S. & Davidson, B. L. CRISPR/Cas9 editing of the mutant Huntingtin allele in vitro and in vivo. Mol. Ther. 25, 12–23 (2017).
Slack, F. J. & Chinnaiyan, A. M. The role of non-coding RNAs in oncology. Cell 179, 1033–1055 (2019).
Buiting, K., Williams, C. & Horsthemke, B. Angelman syndrome — insights into a rare neurogenetic disorder. Nat. Rev. Neurol. 12, 584–593 (2016).
Wolter, J. M. et al. Cas9 gene therapy for Angelman syndrome traps Ube3a-ATS long non-coding RNA. Nature 587, 281–284 (2020).
Schmid, R. S. et al. CRISPR/Cas9 directed to the Ube3a antisense transcript improves Angelman syndrome phenotype in mice. J. Clin. Invest. 131, e142574 (2021).
Singh, R. et al. Regulation of alternative splicing of Bcl-x by BC200 contributes to breast cancer pathogenesis. Cell Death Dis. 7, e2262 (2016).
Koirala, P. et al. LncRNA AK023948 is a positive regulator of AKT. Nat. Commun. 8, 14422 (2017).
Peng, W. X., Huang, J. G., Yang, L., Gong, A. H. & Mo, Y. Y. Linc-RoR promotes MAPK/ERK signaling and confers estrogen-independent growth of breast cancer. Mol. Cancer 16, 161 (2017).
Zhen, S. et al. Inhibition of long non-coding RNA UCA1 by CRISPR/Cas9 attenuated malignant phenotypes of bladder cancer. Oncotarget 8, 9634–9646 (2017).
Yu, Y., Nangia-Makker, P., Farhana, L. & Majumdar, A. P. N. A novel mechanism of lncRNA and miRNA interaction: CCAT2 regulates miR-145 expression by suppressing its maturation process in colon cancer cells. Mol. Cancer 16, 155 (2017).
Ye, X.-T., Huang, H., Huang, W.-P. & Hu, W.-L. LncRNA THOR promotes human renal cell carcinoma cell growth. Biochem. Biophys. Res. Commun. 501, 661–667 (2018).
Zhuo, W. et al. Long noncoding RNA GMAN, up-regulated in gastric cancer tissues, is associated with metastasis in patients and promotes translation of ephrin A1 by competitively binding GMAN-AS. Gastroenterology 156, 676–691 (2019).
Li, C. et al. CRISPR-CasRx targeting LncRNA LINC00341 inhibits tumor cell growth in vitro and in vivo. Front. Mol. Biosci. 8, 638995 (2021).
Li, J. et al. miR-29b contributes to multiple types of muscle atrophy. Nat. Commun. 8, 15201 (2017).
Li, J. et al. CRISPR/Cas9-mediated miR-29b editing as a treatment of different types of muscle atrophy in mice. Mol. Ther. 28, 1359–1372 (2020).
Jing, W. et al. CRISPR/CAS9-mediated genome editing of miRNA-155 inhibits proinflammatory cytokine production by RAW264.7 cells. BioMed. Res. Int. 2015, 326042 (2015).
Nguyen, D.-D. & Chang, S. Development of novel therapeutic agents by inhibition of oncogenic microRNAs. Int. J. Mol. Sci. 19, 65 (2018).
Huo, W. et al. Lentiviral CRISPR/Cas9 vector mediated miR-21 gene editing inhibits the epithelial to mesenchymal transition in ovarian cancer cells. J. Cancer 8, 57–64 (2017).
Yi, B. et al. CRISPR interference and activation of the microRNA-3662-HBP1 axis control progression of triple-negative breast cancer. Oncogene 41, 268–279 (2022).
Hannafon, B. N. et al. miR-23b and miR-27b are oncogenic microRNAs in breast cancer: evidence from a CRISPR/Cas9 deletion study. BMC Cancer 19, 642 (2019).
El Fatimy, R., Subramanian, S., Uhlmann, E. J. & Krichevsky, A. M. Genome editing reveals glioblastoma addiction to microRNA-10b. Mol. Ther. 25, 368–378 (2017).
Nakamura, M., Ivec, A. E., Gao, Y. & Qi, L. S. Durable CRISPR-based epigenetic silencing. BioDesign Res. 2021, 9815820 (2021).
Cox, J. J. et al. An SCN9A channelopathy causes congenital inability to experience pain. Nature 444, 894–898 (2006).
Moreno, A. M. et al. Long-lasting analgesia via targeted in situ repression of Na(V)1.7 in mice. Sci. Transl Med. 13, eaay9056 (2021).
Limpitikul, W. B. et al. A precision medicine approach to the rescue of function on malignant calmodulinopathic long-QT syndrome. Circ. Res. 120, 39–48 (2017).
Moreno, A. M. et al. In situ gene therapy via AAV-CRISPR-Cas9-mediated targeted gene regulation. Mol. Ther. 26, 1818–1827 (2018).
Himeda, C. L., Jones, T. I. & Jones, P. L. CRISPR/dCas9-mediated transcriptional inhibition ameliorates the epigenetic dysregulation at D4Z4 and represses DUX4-fl in FSH muscular dystrophy. Mol. Ther. 24, 527–535 (2016).
Himeda, C. L., Jones, T. I. & Jones, P. L. Targeted epigenetic repression by CRISPR/dSaCas9 suppresses pathogenic DUX4-fl expression in FSHD. Mol. Ther. Methods Clin. Dev. 20, 298–311 (2021).
Matharu, N. et al. CRISPR-mediated activation of a promoter or enhancer rescues obesity caused by haploinsufficiency. Science 363, eaau0629 (2019).
Kemaladewi, D. U. et al. A mutation-independent approach for muscular dystrophy via upregulation of a modifier gene. Nature 572, 125–130 (2019).
Perrin, A., Rousseau, J. & Tremblay, J. P. Increased expression of laminin subunit alpha 1 chain by dCas9-VP160. Mol. Ther. Nucleic Acids 6, 68–79 (2017).
Wojtal, D. et al. Spell checking nature: versatility of CRISPR/Cas9 for developing treatments for inherited disorders. Am. J. Hum. Genet. 98, 90–101 (2016).
Liao, H.-K. et al. In vivo target gene activation via CRISPR/Cas9-mediated trans-epigenetic modulation. Cell 171, 1495–1507.e1415 (2017).
Giménez, C. A. et al. CRISPR-on system for the activation of the endogenous human INS gene. Gene Ther. 23, 543–547 (2016).
Chen, C.-D., Zeldich, E., Li, Y., Yuste, A. & Abraham, C. R. Activation of the anti-aging and cognition-enhancing gene Klotho by CRISPR-dCas9 transcriptional effector complex. J. Mol. Neurosci. 64, 175–184 (2018).
Heman-Ackah, S. M., Bassett, A. R. & Wood, M. J. A. Precision modulation of neurodegenerative disease-related gene expression in human iPSC-derived neurons. Sci. Rep. 6, 28420 (2016).
Moses, C. et al. Activating PTEN tumor suppressor expression with the CRISPR/dCas9 system. Mol. Ther. Nucleic Acids 14, 287–300 (2019).
Xu, X. et al. CRISPR-ON-mediated KLF4 overexpression inhibits the proliferation, migration and invasion of urothelial bladder cancer in vitro and in vivo. Oncotarget 8, 102078–102087 (2017).
Poondla, N., Chandrasekaran, A. P., Heese, K., Kim, K. S. & Ramakrishna, S. CRISPR-mediated upregulation of DR5 and downregulation of cFLIP synergistically sensitize HeLa cells to TRAIL-mediated apoptosis. Biochem. Biophys. Res. Commun. 512, 60–65 (2019).
Kardooni, H. et al. CRISPR-mediated reactivation of DKK3 expression attenuates TGF-β signaling in prostate cancer. Cancers 10, 165 (2018).
Nguyen, N. T. K. et al. CRISPR activation of long non-coding RNA DANCR promotes bone regeneration. Biomaterials 275, 120965 (2021).
Hsu, M. N. et al. CRISPR-based activation of endogenous neurotrophic genes in adipose stem cell sheets to stimulate peripheral nerve regeneration. Theranostics 9, 6099–6111 (2019).
Lawrence, M., Daujat, S. & Schneider, R. Lateral thinking: how histone modifications regulate gene expression. Trends Genet. 32, 42–56 (2016).
Nuñez, J. K. et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184, 2503–2519 (2021).
Amabile, A. et al. Inheritable silencing of endogenous genes by hit-and-run targeted epigenetic editing. Cell 167, 219–232 (2016).
Vojta, A. et al. Repurposing the CRISPR-Cas9 system for targeted DNA methylation. Nucleic Acids Res. 44, 5615–5628 (2016).
Stepper, P. et al. Efficient targeted DNA methylation with chimeric dCas9–Dnmt3a–Dnmt3L methyltransferase. Nucleic Acids Res. 45, 1703–1713 (2017).
Kantor, B. et al. Downregulation of SNCA expression by targeted editing of DNA methylation: a potential strategy for precision therapy in PD. Mol. Ther. 26, 2638–2649 (2018).
Xu, X. et al. A CRISPR-based approach for targeted DNA demethylation. Cell Discov. 2, 16009 (2016).
Morita, S. et al. Targeted DNA demethylation in vivo using dCas9–peptide repeat and scFv–TET1 catalytic domain fusions. Nat. Biotechnol. 34, 1060–1065 (2016).
de Esch, C. E. F. et al. Epigenetic characterization of the FMR1 promoter in induced pluripotent stem cells from human fibroblasts carrying an unmethylated full mutation. Stem Cell Rep. 3, 548–555 (2014).
Liu, X. S. et al. Rescue of fragile X syndrome neurons by DNA methylation editing of the FMR1 gene. Cell 172, 979–992 (2018).
Choudhury, S. R., Cui, Y., Lubecka, K., Stefanska, B. & Irudayaraj, J. CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter. Oncotarget 7, 46545–46556 (2016).
Wang, Q. et al. Targeted demethylation of the SARI promotor impairs colon tumour growth. Cancer Lett. 448, 132–143 (2019).
Xu, X. et al. High-fidelity CRISPR/Cas9-based gene-specific hydroxymethylation rescues gene expression and attenuates renal fibrosis. Nat. Commun. 9, 3509 (2018).
Marx, N. et al. CRISPR-based targeted epigenetic editing enables gene expression modulation of the silenced beta-galactoside alpha-2,6-sialyltransferase 1 in CHO cells. Biotechnol. J. 13, 1700217 (2018).
Hilton, I. B. et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517 (2015).
Gemberling, M. P. et al. Transgenic mice for in vivo epigenome editing with CRISPR-based systems. Nat. Methods 18, 965–974 (2021).
Okada, M., Kanamori, M., Someya, K., Nakatsukasa, H. & Yoshimura, A. Stabilization of Foxp3 expression by CRISPR-dCas9-based epigenome editing in mouse primary T cells. Epigenetics Chromatin 10, 24 (2017).
Liu, J., Sun, M., Cho, K. B., Gao, X. & Guo, B. A CRISPR-Cas9 repressor for epigenetic silencing of KRAS. Pharmacol. Res. 164, 105304 (2021).
Wang, H. et al. Epigenetic targeting of granulin in hepatoma cells by synthetic CRISPR dCas9 Epi-suppressors. Mol. Ther. Nucleic Acids 11, 23–33 (2018).
Villiger, L. et al. Treatment of a metabolic liver disease by in vivo genome base editing in adult mice. Nat. Med. 24, 1519–1525 (2018).
Lim, C. K. W. et al. Treatment of a mouse model of ALS by in vivo base editing. Mol. Ther. 28, 1177–1189 (2020).
Ryu, S.-M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2018).
Koblan, L. W. et al. In vivo base editing rescues Hutchinson–Gilford progeria syndrome in mice. Nature 589, 608–614 (2021).
Newby, G. A. et al. Base editing of haematopoietic stem cells rescues sickle cell disease in mice. Nature 595, 295–302 (2021).
US National Library of Medicine. ClinicalTrials.gov https://ClinicalTrials.gov/show/NCT05456880 (2022).
Antoniou, P., Miccio, A. & Brusson, M. Base and prime editing technologies for blood disorders. Front. Genome Ed. 3, 618406 (2021).
Porto, E. M., Komor, A. C., Slaymaker, I. M. & Yeo, G. W. Base editing: advances and therapeutic opportunities. Nat. Rev. Drug Discov. 19, 839–859 (2020).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Liu, P. et al. Improved prime editors enable pathogenic allele correction and cancer modelling in adult mice. Nat. Commun. 12, 2121 (2021).
Zhi, S. et al. Dual-AAV delivering split prime editor system for in vivo genome editing. Mol. Ther. 30, 283–294 (2022).
Chemello, F. et al. Precise correction of Duchenne muscular dystrophy exon deletion mutations by base and prime editing. Sci. Adv. 7, eabg4910 (2021).
Yin, D. et al. Targeting herpes simplex virus with CRISPR–Cas9 cures herpetic stromal keratitis in mice. Nat. Biotechnol. 39, 567–577 (2021).
Yuen, K.-S. et al. CRISPR/Cas9-mediated genome editing of Epstein–Barr virus in human cells. J. Gen. Virol. 96, 626–636 (2015).
Wang, J. & Quake, S. R. RNA-guided endonuclease provides a therapeutic strategy to cure latent herpesviridae infection. Proc. Natl Acad. Sci. USA 111, 13157–13162 (2014).
Ebina, H., Misawa, N., Kanemura, Y. & Koyanagi, Y. Harnessing the CRISPR/Cas9 system to disrupt latent HIV-1 provirus. Sci. Rep. 3, 2510 (2013).
Kaminski, R. et al. Elimination of HIV-1 genomes from human T-lymphoid cells by CRISPR/Cas9 gene editing. Sci. Rep. 6, 22555 (2016).
Gao, Z., Fan, M., Das, A. T., Herrera-Carrillo, E. & Berkhout, B. Extinction of all infectious HIV in cell culture by the CRISPR-Cas12a system with only a single crRNA. Nucleic Acids Res. 48, 5527–5539 (2020).
Hu, W. et al. RNA-directed gene editing specifically eradicates latent and prevents new HIV-1 infection. Proc. Natl Acad. Sci. USA 111, 11461–11466 (2014).
Zhu, W. et al. The CRISPR/Cas9 system inactivates latent HIV-1 proviral DNA. Retrovirology 12, 22 (2015).
Zhen, S. et al. Harnessing the clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated Cas9 system to disrupt the hepatitis B virus. Gene Ther. 22, 404–412 (2015).
Song, J. et al. CRISPR/Cas9-mediated knockout of HBsAg inhibits proliferation and tumorigenicity of HBV-positive hepatocellular carcinoma cells. J. Cell Biochem. 119, 8419–8431 (2018).
Dong, C. et al. Targeting hepatitis B virus cccDNA by CRISPR/Cas9 nuclease efficiently inhibits viral replication. Antivir. Res. 118, 110–117 (2015).
Ramanan, V. et al. CRISPR/Cas9 cleavage of viral DNA efficiently suppresses hepatitis B virus. Sci. Rep. 5, 10833 (2015).
Seeger, C. & Sohn, J. A. Targeting hepatitis B virus with CRISPR/Cas9. Mol. Ther. Nucleic Acids 3, e216 (2014).
Seeger, C. & Sohn, J. A. Complete spectrum of CRISPR/Cas9-induced mutations on HBV cccDNA. Mol. Ther. 24, 1258–1266 (2016).
Kennedy, E. M. et al. Inactivation of the human papillomavirus E6 or E7 gene in cervical carcinoma cells by using a bacterial CRISPR/Cas RNA-guided endonuclease. J. Virol. 88, 11965–11972 (2014).
Hu, Z. et al. Disruption of HPV16-E7 by CRISPR/Cas system induces apoptosis and growth inhibition in HPV16 positive human cervical cancer cells. BioMed. Res. Int. 2014, 612823 (2014).
Zhen, S. et al. In vitro and in vivo growth suppression of human papillomavirus 16-positive cervical cancer cells by CRISPR/Cas9. Biochem. Biophys. Res. Commun. 450, 1422–1426 (2014).
Price, A. A., Sampson, T. R., Ratner, H. K., Grakoui, A. & Weiss, D. S. Cas9-mediated targeting of viral RNA in eukaryotic cells. Proc. Natl Acad. Sci. USA 112, 6164–6169 (2015).
Mancuso, P. et al. CRISPR based editing of SIV proviral DNA in ART treated non-human primates. Nat. Commun. 11, 6065 (2020).
Abbott, T. R. et al. Development of CRISPR as an antiviral strategy to combat SARS-CoV-2 and influenza. Cell 181, 865–876 (2020).
Zeng, L. et al. Broad-spectrum CRISPR-mediated inhibition of SARS-CoV-2 variants and endemic coronaviruses in vitro. Nat. Commun. 13, 2766 (2022).
Bikard, D. et al. Exploiting CRISPR-Cas nucleases to produce sequence-specific antimicrobials. Nat. Biotechnol. 32, 1146–1150 (2014).
Park, J. Y. et al. Genetic engineering of a temperate phage-based delivery system for CRISPR/Cas9 antimicrobials against Staphylococcus aureus. Sci. Rep. 7, 44929 (2017).
Citorik, R. J., Mimee, M. & Lu, T. K. Sequence-specific antimicrobials using efficiently delivered RNA-guided nucleases. Nat. Biotechnol. 32, 1141–1145 (2014).
Cobb, L. H. et al. CRISPR-Cas9 modified bacteriophage for treatment of Staphylococcus aureus induced osteomyelitis and soft tissue infection. PLoS ONE 14, e0220421 (2019).
Selle, K. et al. In vivo targeting of clostridioides difficile using phage-delivered CRISPR-Cas3 antimicrobials. mBio 11, e00019–e00020 (2020).
Yip, B. H. Recent advances in CRISPR/Cas9 delivery strategies. Biomolecules 10, 839 (2020).
Lino, C. A., Harper, J. C., Carney, J. P. & Timlin, J. A. Delivering CRISPR: a review of the challenges and approaches. Drug Deliv. 25, 1234–1257 (2018).
Raguram, A., Banskota, S. & Liu, D. R. Therapeutic in vivo delivery of gene editing agents. Cell 185, 2806–2827 (2022).
Wei, T. et al. Delivery of tissue-targeted scalpels: opportunities and challenges for in vivo CRISPR/Cas-based genome editing. ACS Nano 14, 9243–9262 (2020).
Hamilton, J. R. et al. Targeted delivery of CRISPR-Cas9 and transgenes enables complex immune cell engineering. Cell Rep. 35, 109207 (2021).
Ling, S. et al. Lentiviral delivery of co-packaged Cas9 mRNA and a VEGFA-targeting guide RNA prevents wet age-related macular degeneration in mice. Nat. Biomed. Eng. 5, 144–156 (2021).
Banskota, S. et al. Engineered virus-like particles for efficient in vivo delivery of therapeutic proteins. Cell 185, 250–265.e216 (2022).
Naeem, M., Majeed, S., Hoque, M. Z. & Ahmad, I. Latest developed strategies to minimize the off-target effects in CRISPR-Cas-mediated genome editing. Cells 9, 1608 (2020).
Manghwar, H. et al. CRISPR/Cas systems in genome editing: methodologies and tools for sgRNA design, off-target evaluation, and strategies to mitigate off-target effects. Adv. Sci. 7, 1902312 (2020).
Kimberland, M. L. et al. Strategies for controlling CRISPR/Cas9 off-target effects and biological variations in mammalian genome editing experiments. J. Biotechnol. 284, 91–101 (2018).
Acknowledgements
M.C. acknowledges support from the National Science Foundation Graduate Research Fellowship Program, the ARCS Foundation Scholarship, the Ruth L. Kirschstein National Research Service Awards for Individual Predoctoral Fellowship (F31AI164936), Siebel Scholar award, and Stanford Maternal and Child Health Research Institute (MCHRI). X.C. acknowledges support from the Stanford Bio-X SIGF Fellowship Program. P.B.F. acknowledges support from California Institute of Regenerative Medicine (CIRM). L.S.Q. acknowledges support from National Science Foundation CAREER award (Award #2046650), NIH (Grant # 1U01DK127405, 1R01CA266470, 1R21CA270609), Stanford MCHRI through the Uytengsu-Hamilton 22q11 Neuropsychiatry Research Award Program, and CIRM (DISC2-12669). L.S.Q. is a Chan Zuckerberg Biohub investigator.
Author information
Authors and Affiliations
Contributions
L.S.Q., M.C. and X.C. researched the data for the article and wrote the text. All authors made substantial contributions to discussion of the content and reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
L.S.Q. is a founder and scientific adviser of Epic Bio, and a scientific adviser of Laboratory of Genomics Research (LGR). The other authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Nephrology thanks Aron Geurts and Susanna Tomasoni, who co-reviewed with Piera Trionfini, for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Trans-activating CRISPR RNA
-
(tracrRNA). RNA that partially base pairs with a CRISPR RNA (crRNA) to form a crRNA–tracrRNA hybrid that binds to and acts as a guide for Cas protein to cleave the targeted DNA sequence.
- Non-homologous end joining
-
(NHEJ). A pathway that repairs DNA double-strand breaks by ligating the break ends without the need for a homologous template.
- Homology-directed repair
-
(HDR). A pathway that repairs DNA double-strand breaks guided by a homologous DNA template.
- Mendelian diseases
-
Genetic disorders caused by mutation in single genes.
- Zinc finger nucleases
-
(ZFNs). Artificial endonucleases that can target and alter specific DNA sequences in the genome. They are generated by fusing a zinc finger DNA-binding domain to a DNA-cleavage domain.
- Transcription activator-like effector nucleases
-
(TALENs). Artificial endonucleases that can target and alter specific DNA sequences in the genome. They are generated by fusing a TAL effector DNA-binding domain to a DNA-cleavage domain.
- Logic-gated gene regulation
-
A process used to control the timing and intensity of gene expression by performing a Boolean operation on multiple biological signals and executing a functional response when appropriate.
- Off-target effects
-
Effects that can occur when CRISPR molecules bind to and alter genomic sites that they were not intended to target, usually as a result of the presence of a similar sequence to the desired target site.
- Complementary DNA
-
(cDNA). DNA synthesized from a single-stranded RNA template that is usually used to express a protein in a cell that does not normally express the protein.
- MicroRNAs
-
(miRNAs). A class of single-stranded non-coding RNAs that can silence RNA and regulate gene expression by base pairing with complementary sequences in mRNA molecules.
- Long non-coding RNAs
-
(lncRNAs). A class of RNAs longer than 200 nucleotides that are not translated into protein and have roles in regulation of gene expression.
- Inteins
-
Protein segments that are capable of excising themselves and joining the flanking protein portions together through a peptide bond during protein splicing.
- Anti-CRISPRs
-
Proteins found in phages that inactivate CRISPR systems, enabling control over CRISPR tools.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chavez, M., Chen, X., Finn, P.B. et al. Advances in CRISPR therapeutics. Nat Rev Nephrol 19, 9–22 (2023). https://doi.org/10.1038/s41581-022-00636-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41581-022-00636-2
This article is cited by
-
Comprehensive review of CRISPR-based gene editing: mechanisms, challenges, and applications in cancer therapy
Molecular Cancer (2024)
-
High-throughput cell optoporation system based on Au nanoparticle layers mediated by resonant irradiation for precise and controllable gene delivery
Scientific Reports (2024)
-
Recent advances in CRISPR-based functional genomics for the study of disease-associated genetic variants
Experimental & Molecular Medicine (2024)
-
Interpreting non-coding disease-associated human variants using single-cell epigenomics
Nature Reviews Genetics (2023)
-
Methylation across the central dogma in health and diseases: new therapeutic strategies
Signal Transduction and Targeted Therapy (2023)