Abstract
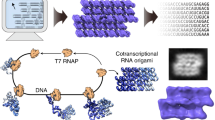
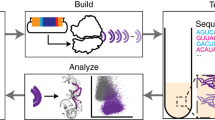
RNA nanotechnology seeks to create nanoscale machines by repurposing natural RNA modules. The field is slowed by the current need for human intuition during three-dimensional structural design. Here, we demonstrate that three distinct problems in RNA nanotechnology can be reduced to a pathfinding problem and automatically solved through an algorithm called RNAMake. First, RNAMake discovers highly stable single-chain solutions to the classic problem of aligning a tetraloop and its sequence-distal receptor, with experimental validation from chemical mapping, gel electrophoresis, solution X-ray scattering and crystallography with 2.55 Å resolution. Second, RNAMake automatically generates structured tethers that integrate 16S and 23S ribosomal RNAs into single-chain ribosomal RNAs that remain uncleaved by ribonucleases and assemble onto messenger RNA. Third, RNAMake enables the automated stabilization of small-molecule binding RNAs, with designed tertiary contacts that improve the binding affinity of the ATP aptamer and improve the fluorescence and stability of the Spinach RNA in cell extracts and in living Escherichia coli cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from the corresponding author upon reasonable request. Furthermore, all of our chemical mapping data are available on https://rmdb.stanford.edu, and a detailed table of the accession identifications is given in the Supplementary Information.
References
Guo, P. The emerging field of RNA nanotechnology. Nat. Nanotechnol. 5, 833–842 (2010).
Grabow, W. W. & Jaeger, L. RNA self-assembly and RNA nanotechnology. Acc. Chem. Res. 47, 1871–1880 (2014).
Leontis, N. B., Lescoute, A. & Westhof, E. The building blocks and motifs of RNA architecture. Curr. Opin. Struct. Biol. 16, 279–287 (2006).
Jaeger, L. & Chworos, A. The architectonics of programmable RNA and DNA nanostructures. Curr. Opin. Struct. Biol. 16, 531–543 (2006).
Jaeger, L. & Leontis, N. B. Tecto-RNA: one-dimensional self-assembly through tertiary interactions. Angew. Chem. Int. Ed. 39, 2521–2524 (2000).
Zhang, H. et al. Crystal structure of 3WJ core revealing divalent ion-promoted thermostability and assembly of the Phi29 hexameric motor pRNA. RNA 19, 1226–1237 (2013).
Weizmann, Y. & Andersen, E. S. RNA nanotechnology—the knots and folds of RNA nanoparticle engineering. MRS Bull. 42, 930–935 (2017).
Jasinski, D., Haque, F., Binzel, D. W. & Guo, P. Advancement of the emerging field of RNA nanotechnology. ACS Nano 11, 1142–1164 (2017).
Afonin, K. A. et al. Computational and experimental characterization of RNA cubic nanoscaffolds. Methods 67, 256–265 (2014).
Jossinet, F., Ludwig, T. E. & Westhof, E. Assemble: an interactive graphical tool to analyze and build RNA architectures at the 2D and 3D levels. Bioinformatics 26, 2057–2059 (2010).
Wimberly, B. T. et al. Structure of the 30S ribosomal subunit. Nature 407, 327–339 (2000).
Nguyen, T. H. D. et al. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 523, 47–52 (2015).
Miao, Z. et al. RNA-Puzzles Round III: 3D RNA structure prediction of five riboswitches and one ribozyme. RNA 23, 655–672 (2017).
Nasalean, L., Baudrey, S., Leontis, N. B. & Jaeger, L. Controlling RNA self-assembly to form filaments. Nucleic Acids Res. 34, 1381–1392 (2006).
Fried, S. D., Schmied, W. H., Uttamapinant, C. & Chin, J. W. Ribosome subunit stapling for orthogonal translation in E. coli. Angew. Chem. Int. Ed. 54, 12791–12794 (2015).
Orelle, C. et al. Protein synthesis by ribosomes with tethered subunits. Nature 524, 119–124 (2015).
Carlson, E. D. Creating Ribo-T: (design, build, test)n. ACS Synth. Biol. 4, 1173–1175 (2015).
Schmied, W. H. et al. Controlling orthogonal ribosome subunit interactions enables evolution of new function. Nature 564, 444–448 (2018).
Famulok, M. Oligonucleotide aptamers that recognize small molecules. Curr. Opin. Struct. Biol. 9, 324–329 (1999).
Porter, E. B., Polaski, J. T., Morck, M. M. & Batey, R. T. Recurrent RNA motifs as scaffolds for genetically encodable small-molecule biosensors. Nat. Chem. Biol. 13, 295–301 (2017).
Gotrik, M. et al. Direct selection of fluorescence-enhancing RNA aptamers. J. Am. Chem. Soc. 140, 3583–3591 (2018).
Montange, R. K. & Batey, R. T. Riboswitches: emerging themes in RNA structure and function. Annu. Rev. Biophys. 37, 117–133 (2008).
Macke, T. J. & Case, D. A. in Molecular Modeling of Nucleic Acids (eds Leontis, N. B. & SantaLucia, J.) 379–393 (American Chemical Society, 1997).
Lee, J. et al. RNA design rules from a massive open laboratory. Proc. Natl Acad. Sci. USA 111, 2122–2127 (2014).
Dibrov, S. M., McLean, J., Parsons, J. & Hermann, T. Self-assembling RNA square. Proc. Natl Acad. Sci. USA 108, 6405–6408 (2011).
Afonin, K. A. et al. Multifunctional RNA nanoparticles. Nano Lett. 14, 5662–5671 (2014).
Khisamutdinov, E. F. et al. Fabrication of RNA 3D nanoprisms for loading and protection of small RNAs and model drugs. Adv. Mater. 28, 10079–10087 (2016).
Bindewald, E., Grunewald, C., Boyle, B., O’Connor, M. & Shapiro, B. A. Computational strategies for the automated design of RNA nanoscale structures from building blocks using NanoTiler. J. Mol. Graph Model 27, 299–308 (2008).
Huang, L. & Lilley, D. M. J. A quasi-cyclic RNA nano-scale molecular object constructed using kink turns. Nanoscale 8, 15189–15195 (2016).
Wu, L., Chai, D., Fraser, M. E. & Zimmerly, S. Structural variation and uniformity among tetraloop–receptor interactions and other loop–helix interactions in RNA crystal structures. PLoS ONE 7, e49225 (2012).
Frederiksen, J. K., Li, N.-S., Das, R., Herschlag, D. & Piccirilli, J. A. Metal-ion rescue revisited: biochemical detection of site-bound metal ions important for RNA folding. RNA 18, 1123–1141 (2012).
Rangan, P., Masquida, B., Westhof, E. & Woodson, S. A. Assembly of core helices and rapid tertiary folding of a small bacterial group I ribozyme. Proc. Natl Acad. Sci. USA 100, 1574–1579 (2003).
Fiore, J. L. & Nesbitt, D. J., An RNA folding motif: GNRA tetraloop–receptor interactions. Q Rev. Biophys. 46, 223–264 (2013).
Klein, D. J., Schmeing, T. M., Moore, P. B. & Steitz, T. A. The kink-turn: a new RNA secondary structure motif. EMBO J. 20, 4214–4221 (2001).
Jewett, M. C., Fritz, B. R., Timmerman, L. E. & Church, G. M. In vitro integration of ribosomal RNA synthesis, ribosome assembly, and translation. Mol. Syst. Biol. 9, 678 (2013).
Fritz, B. R., Jamil, O. K. & Jewett, M. C. Implications of macromolecular crowding and reducing conditions for in vitro ribosome construction. Nucleic Acids Res. 43, 4774–4784 (2015).
Underwood, K. A., Swartz, J. R. & Puglisi, J. D. Quantitative polysome analysis identifies limitations in bacterial cell-free protein synthesis. Biotechnol. Bioeng. 91, 425–435 (2005).
Carothers, J. M., Oestreich, S. C. & Szostak, J. W. Aptamers selected for higher-affinity binding are not more specific for the target ligand. J. Am. Chem. Soc. 128, 7929–7937 (2006).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Jiang, F., Kumar, R. A., Jones, R. A. & Patel, D. J. Structural basis of RNA folding and recognition in an AMP–RNA aptamer complex. Nature 382, 183–186 (1996).
Huang, Z. & Szostak, J. W. Evolution of aptamers with a new specificity and new secondary structures from an ATP aptamer. RNA 9, 1456–1463 (2003).
Sassanfar, M. & Szostak, J. W. An RNA motif that binds ATP. Nature 364, 550–553 (1993).
Sazani, P. L., Larralde, R. & Szostak, J. W. A small aptamer with strong and specific recognition of the triphosphate of ATP. J. Am. Chem. Soc. 126, 8370–8371 (2004).
Geary, C., Chworos, A., Verzemnieks, E., Voss, N. R. & Jaeger, L. Composing RNA nanostructures from a syntax of RNA structural modules. Nano Lett. 17, 7095–7101 (2017).
Kellenberger, C. A., Chen, C., Whiteley, A. T., Portnoy, D. A. & Hammond, M. C. RNA-based fluorescent biosensors for live cell imaging of second messenger cyclic di-AMP. J. Am. Chem. Soc. 137, 6432–6435 (2015).
Strack, R. L., Disney, M. D. & Jaffrey, S. R. A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat-containing RNA. Nat. Methods 10, 1219–1224 (2013).
Filonov, G. S., Moon, J. D., Svensen, N. & Jaffrey, S. R. Broccoli: rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 136, 16299–16308 (2014).
Ketterer, S., Fuchs, D., Weber, W. & Meier, M. Systematic reconstruction of binding and stability landscapes of the fluorogenic aptamer Spinach. Nucleic Acids Res. 43, 9564–9572 (2015).
Song, W., Strack, R. L., Svensen, N. & Jaffrey, S. R. Plug-and-play fluorophores extend the spectral properties of Spinach. J. Am. Chem. Soc. 136, 1198–1201 (2014).
Shi, X., Huang, L., Lilley, D. M. J., Harbury, P. B. & Herschlag, D. The solution structural ensembles of RNA kink-turn motifs and their protein complexes. Nat. Chem. Biol. 12, 146–152 (2016).
Buenrostro, J. D. et al. Quantitative analysis of RNA–protein interactions on a massively parallel array reveals biophysical and evolutionary landscapes. Nat. Biotechnol. 32, 562–568 (2014).
Petrov, A. I., Zirbel, C. L. & Leontis, N. B. Automated classification of RNA 3D motifs and the RNA 3D Motif Atlas. RNA 19, 1327–1340 (2013).
Lu, X.-J., Bussemaker, H. J. & Olson, W. K. DSSR: an integrated software tool for dissecting the spatial structure of RNA. Nucleic Acids Res. 43, e142 (2015).
Huynh, D. Q. Metrics for 3D rotations: comparison and analysis. J. Math. Imaging Vis. 35, 155–164 (2009).
Karney, C. F. F. Quaternions in molecular modeling. J. Mol. Graph Model 25, 595–604 (2007).
Finkelstein, A. V., Badretdinov, A. Ya & Gutin, A. M. Why do protein architectures have Boltzmann-like statistics? Proteins 23, 142–150 (1995).
Acknowledgements
We thank S. Bonilla for assistance in performing the native gel assays and A. Watkins for discussions about ribosome tether design. We thank the Straight lab for graciously providing X. laevis whole cell lysate. This work was supported by the National Institutes of Health, through NIGMS MIRA R35 GM122579 (R.D.), R01 GM121487 (R.D.), New Innovator Award 1DP2GM110838 (J.B.L.), Ruth L. Kirschstein National Research Service Award Postdoctoral Fellowships GM112294 (J.D.Y.) and GM100953 (D.E.), P01 GM066275 (D.H.) and R35 GM118070 (J.S.K.), a Stanford School of Medicine Discovery Innovation Award (R.D.), Army Research Office W911NF-16-1-0372 (M.C.J.), the National Science Foundation through MCB-1716766 (M.C.J.), Career Award 1452441 (J.B.L.) and Graduate Research Fellowship DGE-1324585 (A.E.D.), the David and Lucile Packard Foundation (M.C.J.) and the Camille Dreyfus Teacher-Scholar Program (M.C.J.).
Author information
Authors and Affiliations
Contributions
R.D. and J.D.Y. conceived the study. J.D.Y. developed RNAMake and generated the models and sequences used throughout the study. A.N.O., W.K. and J.D.Y. performed the chemical mapping, titrations and native gel assays for miniTTR constructs. X.S. and D.H. designed and performed the SAXS on miniTTR 2 and 6. D.E. solved the miniTTR crystal structure assisted by D.A.C. and J.S.K. in preparing the RNA and analysis. E.D.C., A.E.D. and M.C.J. made and tested the RNAMake-designed ribosomes P.D.C. and J.B.L. carried out the SHAPE-seq and in vivo tests of Spinach-TTRs. M.R.G. performed fluorescence and lysate experiments on Spinach-TTRs. J.D.Y. and R.D. wrote the paper, with participation by all the authors.
Corresponding author
Ethics declarations
Competing interests
Stanford University is filing a patent on aptamer stabilization with RNAMake.
Additional information
Peer review information: Nature Nanotechnology thanks Peixuan Guo, Nils (G) Walter and other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Results, Supplementary Discussion, Supplementary Methods, Supplementary Tables 1–10, Supplementary Figs. 1–14 and Supplementary refs. 1–46.
Rights and permissions
About this article
Cite this article
Yesselman, J.D., Eiler, D., Carlson, E.D. et al. Computational design of three-dimensional RNA structure and function. Nat. Nanotechnol. 14, 866–873 (2019). https://doi.org/10.1038/s41565-019-0517-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-019-0517-8
This article is cited by
-
Community science designed ribosomes with beneficial phenotypes
Nature Communications (2023)
-
Cell-free Biosynthesis of Peptidomimetics
Biotechnology and Bioprocess Engineering (2023)
-
Nanodelivery of nucleic acids
Nature Reviews Methods Primers (2022)
-
Engineering synthetic RNA devices for cell control
Nature Reviews Genetics (2022)
-
Engineering DNA quadruplexes in DNA nanostructures for biosensor construction
Nano Research (2022)