Abstract
Cooperation is commonly believed to be favourable in spatially structured environments, as these systems promote genetic relatedness that reduces the likelihood of exploitation by cheaters. Here we show that a Pseudomonas aeruginosa population that exhibited cooperative swarming was invaded by cheaters when subjected to experimental evolution through cycles of range expansion on solid media, but not in well-mixed liquid cultures. Our results suggest that cooperation is disfavoured in a more structured environment, which is the opposite of the prevailing view. We show that spatial expansion of the population prolongs cooperative swarming, which was vulnerable to cheating. Our findings reveal a mechanism by which spatial structures can suppress cooperation through modulation of the quantitative traits of cooperation, a process that leads to population divergence towards distinct colonization strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available on Zenodo (https://doi.org/10.5281/zenodo.10210659). Raw sequencing reads from whole-genome sequencing are deposited in the NCBI Sequence Read Archive (BioProject PRJNA875393).
Code availability
The MATLAB code used for data generation and/or analysis in the study is available on GitHub (https://github.com/youlab/MultistrainPatterns_NanLuo).
References
Celiker, H. & Gore, J. Cellular cooperation: insights from microbes. Trends Cell Biol. 23, 9–15 (2013).
Ghoul, M., Griffin, A. S. & West, S. A. Toward an evolutionary definition of cheating. Evolution 68, 318–331 (2014).
West, S. A., Cooper, G. A., Ghoul, M. B. & Griffin, A. S. Ten recent insights for our understanding of cooperation. Nat. Ecol. Evol. 5, 419–430 (2021).
Nowak, M. A. & May, R. M. Evolutionary games and spatial chaos. Nature 359, 826–829 (1992).
Nowak, M. A. & Sigmund, K. in The Geometry of Ecological Interactions: Simplifying Spatial Complexity (eds Metz J. A. J. et al.) 135–150 (Cambridge Univ. Press, 2000).
Ackermann, M. et al. Self-destructive cooperation mediated by phenotypic noise. Nature 454, 987–990 (2008).
Funk, F. & Hauert, C. Directed migration shapes cooperation in spatial ecological public goods games. PLoS Comput. Biol. 15, e1006948 (2019).
Xavier, J. B. & Foster, K. R. Cooperation and conflict in microbial biofilms. Proc. Natl Acad. Sci. USA 104, 876–881 (2007).
Momeni, B., Waite, A. J. & Shou, W. Spatial self-organization favors heterotypic cooperation over cheating. eLife 2, e00960 (2013).
Hol, F. J. et al. Spatial structure facilitates cooperation in a social dilemma: empirical evidence from a bacterial community. PLoS ONE 8, e77042 (2013).
Xu, S. & Van Dyken, J. D. Microbial expansion–collision dynamics promote cooperation and coexistence on surfaces. Evolution 72, 153–169 (2018).
Hamilton, W. D. The genetical evolution of social behaviour. I. J. Theor. Biol. 7, 1–16 (1964).
Hamilton, W. D. The genetical evolution of social behaviour. II. J. Theor. Biol. 7, 17–52 (1964).
Griffin, A. S., West, S. A. & Buckling, A. Cooperation and competition in pathogenic bacteria. Nature 430, 1024–1027 (2004).
Nowak, M. A. Five rules for the evolution of cooperation. Science 314, 1560–1563 (2006).
West, S. A., Griffin, A. S. & Gardner, A. Evolutionary explanations for cooperation. Curr. Biol. 17, R661–R672 (2007).
Cornwallis, C. K., West, S. A. & Griffin, A. S. Routes to indirect fitness in cooperatively breeding vertebrates: kin discrimination and limited dispersal. J. Evol. Biol. 22, 2445–2457 (2009).
Lion, S. & Baalen, M. Self-structuring in spatial evolutionary ecology. Ecol. Lett. 11, 277–295 (2008).
Sexton, D. J. & Schuster, M. Nutrient limitation determines the fitness of cheaters in bacterial siderophore cooperation. Nat. Commun. 8, 230 (2017).
Boyle, K. E., Monaco, H., van Ditmarsch, D., Deforet, M. & Xavier, J. B. Integration of metabolic and quorum sensing signals governing the decision to cooperate in a bacterial social trait. PLoS Comput. Biol. 11, e1004279 (2015).
Xavier, J. B., Kim, W. & Foster, K. R. A molecular mechanism that stabilizes cooperative secretions in Pseudomonas aeruginosa. Mol. Microbiol. 79, 166–179 (2011).
Kummerli, R., Jiricny, N., Clarke, L. S., West, S. A. & Griffin, A. S. Phenotypic plasticity of a cooperative behaviour in bacteria. J. Evol. Biol. 22, 589–598 (2009).
Butaite, E., Kramer, J., Wyder, S. & Kummerli, R. Environmental determinants of pyoverdine production, exploitation and competition in natural Pseudomonas communities. Environ. Microbiol. 20, 3629–3642 (2018).
Diggle, S. P., Griffin, A. S., Campbell, G. S. & West, S. A. Cooperation and conflict in quorum-sensing bacterial populations. Nature 450, 411–414 (2007).
Sandoz, K. M., Mitzimberg, S. M. & Schuster, M. Social cheating in Pseudomonas aeruginosa quorum sensing. Proc. Natl Acad. Sci. USA 104, 15876–15881 (2007).
Monaco, H. et al. Spatial–temporal dynamics of a microbial cooperative behavior resistant to cheating. Nat. Commun. 13, 721 (2022).
van Ditmarsch, D. et al. Convergent evolution of hyperswarming leads to impaired biofilm formation in pathogenic bacteria. Cell Rep. 4, 697–708 (2013).
Latifi, A., Foglino, M., Tanaka, K., Williams, P. & Lazdunski, A. A hierarchical quorum-sensing cascade in Pseudomonas aeruginosa links the transcriptional activators LasR and RhIR (VsmR) to expression of the stationary-phase sigma factor RpoS. Mol. Microbiol. 21, 1137–1146 (1996).
Pesci, E. C., Pearson, J. P., Seed, P. C. & Iglewski, B. H. Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol. 179, 3127–3132 (1997).
Ochsner, U. A., Fiechter, A. & Reiser, J. Isolation, characterization, and expression in Escherichia coli of the Pseudomonas aeruginosa rhlAB genes encoding a rhamnosyltransferase involved in rhamnolipid biosurfactant synthesis. J. Biol. Chem. 269, 19787–19795 (1994).
de Kievit, T., Seed, P. C., Nezezon, J., Passador, L. & Iglewski, B. H. RsaL, a novel repressor of virulence gene expression in Pseudomonas aeruginosa. J. Bacteriol. 181, 2175–2184 (1999).
Rampioni, G. et al. The quorum-sensing negative regulator RsaL of Pseudomonas aeruginosa binds to the lasI promoter. J. Bacteriol. 188, 815–819 (2006).
Tremblay, J., Richardson, A. P., Lepine, F. & Deziel, E. Self-produced extracellular stimuli modulate the Pseudomonas aeruginosa swarming motility behaviour. Environ. Microbiol. 9, 2622–2630 (2007).
Kearns, D. B. A field guide to bacterial swarming motility. Nat. Rev. Microbiol. 8, 634–644 (2010).
Wadhwa, N. & Berg, H. C. Bacterial motility: machinery and mechanisms. Nat. Rev. Microbiol. 20, 161–173 (2022).
Caiazza, N. C., Shanks, R. M. & O’Toole, G. A. Rhamnolipids modulate swarming motility patterns of Pseudomonas aeruginosa. J. Bacteriol. 187, 7351–7361 (2005).
Santamaria, G. et al. Evolution and regulation of microbial secondary metabolism. eLife 11, e76119 (2022).
Deng, P., de Vargas, R. L., van Ditmarsch, D. & Xavier, J. B. The ecological basis of morphogenesis: branching patterns in swarming colonies of bacteria. New J. Phys. 16, 015006–015006 (2014).
Gude, S. et al. Bacterial coexistence driven by motility and spatial competition. Nature 578, 588–592 (2020).
Celedón, R. S. & Díaz, L. B. Natural pigments of bacterial origin and their possible biomedical applications. Microorganisms 9, 739 (2021).
Luo, N., Wang, S., Lu, J., Ouyang, X. & You, L. Collective colony growth is optimized by branching pattern formation in Pseudomonas aeruginosa. Mol. Syst. Biol. 17, e10089 (2021).
Huberman, B. A. & Glance, N. S. Evolutionary games and computer simulations. Proc. Natl Acad. Sci. USA 90, 7716–7718 (1993).
Nowak, M. A. & Sigmund, K. The alternating prisoner’s dilemma. J. Theor. Biol. 168, 219–226 (1994).
Ohtsuki, H., Hauert, C., Lieberman, E. & Nowak, M. A. A simple rule for the evolution of cooperation on graphs and social networks. Nature 441, 502–505 (2006).
Lieberman, E., Hauert, C. & Nowak, M. A. Evolutionary dynamics on graphs. Nature 433, 312–316 (2005).
Darch, S. E., West, S. A., Winzer, K. & Diggle, S. P. Density-dependent fitness benefits in quorum-sensing bacterial populations. Proc. Natl Acad. Sci. USA 109, 8259–8263 (2012).
Denervaud, V. et al. Characterization of cell-to-cell signaling-deficient Pseudomonas aeruginosa strains colonizing intubated patients. J. Clin. Microbiol. 42, 554–562 (2004).
Schaber, J. A. et al. Analysis of quorum sensing-deficient clinical isolates of Pseudomonas aeruginosa. J. Med. Microbiol. 53, 841–853 (2004).
Andersen, S. B. et al. Privatisation rescues function following loss of cooperation. eLife 7, e38594 (2018).
Jin, Z. et al. Conditional privatization of a public siderophore enables Pseudomonas aeruginosa to resist cheater invasion. Nat. Commun. 9, 1383 (2018).
Steenackers, H. P., Parijs, I., Dubey, A., Foster, K. R. & Vanderleyden, J. Experimental evolution in biofilm populations. FEMS Microbiol. Rev. 40, 373–397 (2016).
Vega, N. M. & Gore, J. Collective antibiotic resistance: mechanisms and implications. Curr. Opin. Microbiol. 21, 28–34 (2014).
Muok, A. R. & Briegel, A. Intermicrobial hitchhiking: how nonmotile microbes leverage communal motility. Trends Microbiol. 29, 542–550 (2021).
Acknowledgements
We thank Duke Compute Cluster for assistance with high-throughput computation. We thank J. Xavier (Memorial Sloan Kettering Cancer Center) for sharing P. aeruginosa PA14 strains. This work was partially supported by grants from the Office of Naval Research (L.Y.: N00014-12-1-0631), National Science Foundation (L.Y.: MCB-1937259) and National Institutes of Health (L.Y.: R01GM098642).
Author information
Authors and Affiliations
Contributions
N.L. conceived the research, designed and performed the modelling and experiments, interpreted the results and wrote the paper. J.L. and E.Ş. designed and performed the experiments, interpreted the results and assisted with paper revisions. A.S. assisted with modelling, interpretation of results and paper revisions. Y.Y. assisted with the experiments. X.O. assisted with the modelling and experiments. S.A.W. assisted with the interpretation of results and paper revisions. L.Y. conceived the research, assisted in research design, interpreted the results and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Ákos T. Kovács and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental evolution of P. aeruginosa led to biomass decline.
The total accumulation of biomass of P. aeruginosa PA14 colonies declined during experimental evolution. Cells of a colony were flushed off the plate after colony growth for 20 h and resuspended, and the biomass was determined by measuring OD600. The relative biomass of a colony was normalized by the average of wildtype colonies.
Extended Data Fig. 2 Nonswarmer or hyperswarmer did not emerge during experimental evolution of P. aeruginosa in liquid cultures.
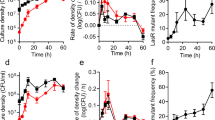
a, The growth rate of P. aeruginosa increased during experimental evolution within liquid cultures. Growth curves were measured using cell cultures of the ancestral strain (Day 1) and the populations on Day 5 and Day 7 of the experimental evolution (n = 3 for each group; all replicates were shown). Both experimental evolution and the growth curve measurements were carried out using liquid swarming media with 8 g/L casamino acids. b, Experimental evolution in liquid cultures did not alter the colony phenotype of P. aeruginosa or cause biomass decline. Shown are colonies of wildtype and the evolved populations of three independent lineages. c, Comparison of the phenotypic characterizations of wildtype and isolates from populations evolved in liquid or on plates. Strains with nonswarming phenotype (low colony area with low branching index) or hyperswarming phenotype (high colony area with low branching index) did not emerge during evolution in liquid cultures.
Extended Data Fig. 3 Nonswarmers are defective in surfactant production.
Cells were grown in liquid swarming media with 8 g/L casamino acids for 20 h. The amount of surfactant in the supernatant was determined using the drop collapse assay. The droplet area (normalized by the negative control, blank medium) reflects the surfactant concentration. N = 36 for each group. Data are presented as mean values +/- SEM. Unpaired, two-sided t test with Welch’s correction and a 95% confidence interval was used to compare between wildtype and nonswarmer. **** P < 0.0001.
Extended Data Fig. 4 Intermediate nutrient level causes P. aeruginosa wildtype and hyperswarmers to produce the highest amount of surfactant.
Cells were grown in liquid swarming media with varying initial concentrations of casamino acids for 20 h. The amount of surfactant in the supernatant was determined using the drop collapse assay. The droplet area (normalized by the negative control, blank medium) reflects the surfactant concentration. N = 36 for each group. Data are presented as mean values +/- SEM. Brown-Forsythe and Welch ANOVA test (which does not assume equal variances) were used to compare between groups. **** P < 0.0001, ** P = 0.0014. Data in Extended Data Fig. 3 were reused for comparison.
Extended Data Fig. 5 Cheater reduced the biomass accumulation of multi-strain colonies.
The total biomass of colonies with different compositions (normalized by wildtype) was determined by measuring OD600. All data points were shown: n = 3-5 for each colony type. Data are presented as mean values +/- SEM. One-way ANOVA followed by Tukey’s multiple comparisons test (95% confidence intervals) was used to compare between groups. Significantly different groups are indicated by letters: groups with at least one common letter are not significantly different; otherwise, they are significantly different.
Extended Data Fig. 6 Model recaptured the competition dynamics between strains.
Pairwise competition between the three types of strains (a, cheater vs. wild type; b, hyperswarmer vs. wildtype; c, cheater vs. hyperswarmer) were carried out either in liquid cultures or on solid swarming plates. Competing strains with fluorescent labels were mixed in varying ratios and grown for 16 h. The final population compositions were determined by CFU counting. In simulations, identical parameters were used when comparing liquid and solid phases. Data in panel a was presented in Fig. 4a in a different format.
Supplementary information
Supplementary Information
Supplementary Note 1, Figs. 1–12 and Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Luo, N., Lu, J., Şimşek, E. et al. The collapse of cooperation during range expansion of Pseudomonas aeruginosa. Nat Microbiol (2024). https://doi.org/10.1038/s41564-024-01627-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41564-024-01627-8