Abstract
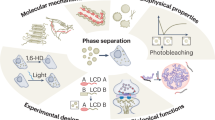
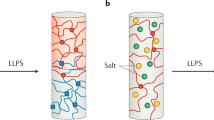
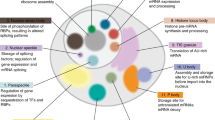
Phase separation, in which macromolecules partition into a concentrated phase that is immiscible with a dilute phase, is involved with fundamental cellular processes across the tree of life. We review the principles of phase separation and highlight how it impacts diverse processes in the fungal kingdom. These include the regulation of autophagy, cell signalling pathways, transcriptional circuits and the establishment of asymmetry in fungal cells. We describe examples of stable, phase-separated assemblies including membraneless organelles such as the nucleolus as well as transient condensates that also arise through phase separation and enable cells to rapidly and reversibly respond to important environmental cues. We showcase how research into phase separation in model yeasts, such as Saccharomyces cerevisiae and Schizosaccharomyces pombe, in conjunction with that in plant and human fungal pathogens, such as Ashbya gossypii and Candida albicans, is continuing to enrich our understanding of fundamental molecular processes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Gomes, E. & Shorter, J. The molecular language of membraneless organelles. J. Biol. Chem. 294, 7115–7127 (2019).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science https://doi.org/10.1126/science.aaf4382 (2017).
Mitrea, D. M. & Kriwacki, R. W. Phase separation in biology; functional organization of a higher order. Cell Commun. Signal. 14, 1 (2016).
Riback, J. A. et al. Stress-triggered phase separation is an adaptive, evolutionarily tuned response. Cell 168, 1028–1040 (2017).
Ivanov, P., Kedersha, N. & Anderson, P. Stress granules and processing bodies in translational control. Cold Spring Harb. Perspect. Biol. https://doi.org/10.1101/cshperspect.a032813 (2019).
Moon, S. L. et al. Multicolour single-molecule tracking of mRNA interactions with RNP granules. Nat. Cell Biol. 21, 162–168 (2019).
Jain, S. et al. ATPase-modulated stress granules contain a diverse proteome and substructure. Cell 164, 487–498 (2016).
Kroschwald, S. et al. Promiscuous interactions and protein disaggregases determine the material state of stress-inducible RNP granules. eLife 4, e06807 (2015).
Anderson, P. & Kedersha, N. RNA granules. J. Cell Biol. 172, 803–808 (2006).
Kroschwald, S. et al. Different material states of Pub1 condensates define distinct modes of stress adaptation and recovery. Cell Rep. 23, 3327–3339 (2018).
Yoo, H., Bard, J. A. M., Pilipenko, E. V. & Drummond, D. A. Chaperones directly and efficiently disperse stress-triggered biomolecular condensates. Mol. Cell 82, 741–755 (2022).
Luo, Y., Na, Z. & Slavoff, S. A. P-bodies: composition, properties, and functions. Biochemistry 57, 2424–2431 (2018).
Schutz, S., Noldeke, E. R. & Sprangers, R. A synergistic network of interactions promotes the formation of in vitro processing bodies and protects mRNA against decapping. Nucleic Acids Res. https://doi.org/10.1093/nar/gkx353 (2017).
Fromm, S. A. et al. In vitro reconstitution of a cellular phase-transition process that involves the mRNA decapping machinery. Angew. Chem. Int. Ed. Engl. 53, 7354–7359 (2014).
Xing, W., Muhlrad, D., Parker, R. & Rosen, M. K. A quantitative inventory of yeast P body proteins reveals principles of composition and specificity. eLife https://doi.org/10.7554/eLife.56525 (2020).
Fuller, G. G. et al. RNA promotes phase separation of glycolysis enzymes into yeast G bodies in hypoxia. eLife https://doi.org/10.7554/eLife.48480 (2020).
Jin, M. et al. Glycolytic enzymes coalesce in G bodies under hypoxic stress. Cell Rep. 20, 895–908 (2017).
Munder, M. C. et al. A pH-driven transition of the cytoplasm from a fluid- to a solid-like state promotes entry into dormancy. eLife https://doi.org/10.7554/eLife.09347 (2016).
Joyner, R. P. et al. A glucose-starvation response regulates the diffusion of macromolecules. eLife https://doi.org/10.7554/eLife.09376 (2016).
Dechant, R. et al. Cytosolic pH is a second messenger for glucose and regulates the PKA pathway through V-ATPase. EMBO J. 29, 2515–2526 (2010).
Orij, R., Postmus, J., Ter Beek, A., Brul, S. & Smits, G. J. In vivo measurement of cytosolic and mitochondrial pH using a pH-sensitive GFP derivative in Saccharomyces cerevisiae reveals a relation between intracellular pH and growth. Microbiology 155, 268–278 (2009).
Petrovska, I. et al. Filament formation by metabolic enzymes is a specific adaptation to an advanced state of cellular starvation. eLife https://doi.org/10.7554/eLife.02409 (2014).
Marini, G., Nuske, E., Leng, W., Alberti, S. & Pigino, G. Reorganization of budding yeast cytoplasm upon energy depletion. Mol. Biol. Cell 31, 1232–1245 (2020).
Parry, B. R. et al. The bacterial cytoplasm has glass-like properties and is fluidized by metabolic activity. Cell 156, 183–194 (2014).
Franzmann, T. M. et al. Phase separation of a yeast prion protein promotes cellular fitness. Science https://doi.org/10.1126/science.aao5654 (2018).
Franzmann, T. M. & Alberti, S. Prion-like low-complexity sequences: key regulators of protein solubility and phase behavior. J. Biol. Chem. 294, 7128–7136 (2019).
Lyke, D. R., Dorweiler, J. E. & Manogaran, A. L. The three faces of Sup35. Yeast 36, 465–472 (2019).
Andre, A. A. M. & Spruijt, E. Liquid–liquid phase separation in crowded environments. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21165908 (2020).
Delarue, M. et al. mTORC1 controls phase separation and the biophysical properties of the cytoplasm by tuning crowding. Cell 174, 338–349 (2018).
Shu, T. et al. nucGEMs probe the biophysical properties of the nucleoplasm. Preprint at bioRxiv https://doi.org/10.1101/2021.11.18.469159 (2022).
Alexandrov, A. I. et al. Analysis of novel hyperosmotic shock response suggests ‘beads in liquid’ cytosol structure. Biol. Open https://doi.org/10.1242/bio.044529 (2019).
Liu, B. et al. The polarisome is required for segregation and retrograde transport of protein aggregates. Cell 140, 257–267 (2010).
Xie, Y. et al. Polarisome scaffolder Spa2-mediated macromolecular condensation of Aip5 for actin polymerization. Nat. Commun. 10, 5078 (2019).
Xie, Y. & Miao, Y. Polarisome assembly mediates actin remodeling during polarized yeast and fungal growth. J Cell Sci. https://doi.org/10.1242/jcs.247916 (2021).
Zhang, H. et al. RNA controls polyQ protein phase transitions. Mol. Cell 60, 220–230 (2015).
Lee, C., Occhipinti, P. & Gladfelter, A. S. PolyQ-dependent RNA–protein assemblies control symmetry breaking. J. Cell Biol. 208, 533–544 (2015).
Lee, C. et al. Protein aggregation behavior regulates cyclin transcript localization and cell-cycle control. Dev. Cell 25, 572–584 (2013).
Roden, C. & Gladfelter, A. S. RNA contributions to the form and function of biomolecular condensates. Nat. Rev. Mol. Cell Biol. 22, 183–195 (2021).
Nakatogawa, H., Suzuki, K., Kamada, Y. & Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat. Rev. Mol. Cell Biol. 10, 458–467 (2009).
Noda, N. N., Wang, Z. & Zhang, H. Liquid–liquid phase separation in autophagy. J. Cell Biol. https://doi.org/10.1083/jcb.202004062 (2020).
Fujioka, Y. et al. Phase separation organizes the site of autophagosome formation. Nature 578, 301–305 (2020).
Yamamoto, H. et al. The intrinsically disordered protein Atg13 mediates supramolecular assembly of autophagy initiation complexes. Dev. Cell 38, 86–99 (2016).
Yamamoto, H. et al. Atg9 vesicles are an important membrane source during early steps of autophagosome formation. J. Cell Biol. 198, 219–233 (2012).
Suzuki, S. W. et al. Atg13 HORMA domain recruits Atg9 vesicles during autophagosome formation. Proc. Natl Acad. Sci. USA 112, 3350–3355 (2015).
Yamasaki, A. & Noda, N. N. Structural biology of the Cvt pathway. J. Mol. Biol. 429, 531–542 (2017).
Yamasaki, A. et al. Liquidity is a critical determinant for selective autophagy of protein condensates. Mol. Cell 77, 1163–1175 (2020).
Yamasaki, A. et al. Structural basis for receptor-mediated selective autophagy of aminopeptidase I aggregates. Cell Rep. 16, 19–27 (2016).
Zhang, G., Wang, Z., Du, Z. & Zhang, H. mTOR regulates phase separation of PGL granules to modulate their autophagic degradation. Cell 174, 1492–1506 (2018).
Wu, X. & Tu, B. P. Selective regulation of autophagy by the Iml1–Npr2–Npr3 complex in the absence of nitrogen starvation. Mol. Biol. Cell 22, 4124–4133 (2011).
Kato, M. et al. Redox state controls phase separation of the teast ataxin-2 protein via reversible oxidation of its methionine-rich low-complexity domain. Cell 177, 711–721 (2019).
Yang, Y. S. et al. Yeast ataxin-2 forms an intracellular condensate required for the inhibition of TORC1 signaling during respiratory growth. Cell 177, 697–710 (2019).
Prouteau, M. & Loewith, R. TOR signaling is going through a phase. Cell Metab. 29, 1019–1021 (2019).
Lin, Y. et al. Redox-mediated regulation of an evolutionarily conserved cross-β structure formed by the TDP43 low complexity domain. Proc. Natl Acad. Sci. USA 117, 28727–28734 (2020).
Martin, R., Pohlers, S., Muhlschlegel, F. A. & Kurzai, O. CO2 sensing in fungi: at the heart of metabolic signaling. Curr. Genet. 63, 965–972 (2017).
Zhang, M. et al. The intrinsically disordered region from PP2C phosphatases functions as a conserved CO2 sensor. Nat. Cell Biol. 24, 1029–1037 (2022).
Feric, M. et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell 165, 1686–1697 (2016).
Thiry, M. & Lafontaine, D. L. Birth of a nucleolus: the evolution of nucleolar compartments. Trends Cell Biol. 15, 194–199 (2005).
Hult, C. et al. Enrichment of dynamic chromosomal crosslinks drive phase separation of the nucleolus. Nucleic Acids Res. 45, 11159–11173 (2017).
Lawrimore, J. et al. The rDNA is biomolecular condensate formed by polymer–polymer phase separation and is sequestered in the nucleolus by transcription and R-loops. Nucleic Acids Res. 49, 4586–4598 (2021).
Hall, A. C., Ostrowski, L. A. & Mekhail, K. Phase separation as a melting pot for DNA repeats. Trends Genet. 35, 589–600 (2019).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Wang, L. et al. Histone modifications regulate chromatin compartmentalization by contributing to a phase separation mechanism. Mol. Cell 76, 646–659 (2019).
Keenen, M. M. et al. HP1 proteins compact DNA into mechanically and positionally stable phase separated domains. eLife https://doi.org/10.7554/eLife.64563 (2021).
Eeftens, J. M., Kapoor, M., Michieletto, D. & Brangwynne, C. P. Polycomb condensates can promote epigenetic marks but are not required for sustained chromatin compaction. Nat. Commun. 12, 5888 (2021).
Tatavosian, R. et al. Nuclear condensates of the Polycomb protein chromobox 2 (CBX2) assemble through phase separation. J. Biol. Chem. 294, 1451–1463 (2019).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855 (2018).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Shrinivas, K. et al. Enhancer features that drive formation of transcriptional condensates. Mol. Cell 75, 549–561 (2019).
Cho, W. K. et al. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 361, 412–415 (2018).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Mansour, M. R. et al. Oncogene regulation. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science 346, 1373–1377 (2014).
Reiter, F., Wienerroither, S. & Stark, A. Combinatorial function of transcription factors and cofactors. Curr. Opin. Genet. Dev. 43, 73–81 (2017).
Sorrells, T. R., Booth, L. N., Tuch, B. B. & Johnson, A. D. Intersecting transcription networks constrain gene regulatory evolution. Nature 523, 361–365 (2015).
Beyhan, S., Gutierrez, M., Voorhies, M. & Sil, A. A temperature-responsive network links cell shape and virulence traits in a primary fungal pathogen. PLoS Biol. 11, e1001614 (2013).
Borneman, A. R. et al. Divergence of transcription factor binding sites across related yeast species. Science 317, 815–819 (2007).
Nobile, C. J. et al. A recently evolved transcriptional network controls biofilm development in Candida albicans. Cell 148, 126–138 (2012).
Gomez-Pastor, R., Burchfiel, E. T. & Thiele, D. J. Regulation of heat shock transcription factors and their roles in physiology and disease. Nat. Rev. Mol. Cell Biol. 19, 4–19 (2018).
Hernday, A. D. et al. Structure of the transcriptional network controlling white–opaque switching in Candida albicans. Mol. Microbiol. 90, 22–35 (2013).
Zordan, R. E., Miller, M. G., Galgoczy, D. J., Tuch, B. B. & Johnson, A. D. Interlocking transcriptional feedback loops control white–opaque switching in Candida albicans. PLoS Biol. 5, e256 (2007).
Frazer, C. et al. Epigenetic cell fate in Candida albicans is controlled by transcription factor condensates acting at super-enhancer-like elements. Nat. Microbiol. https://doi.org/10.1038/s41564-020-0760-7 (2020).
Chowdhary, S., Kainth, A. S., Pincus, D. & Gross, D. S. Heat shock factor 1 drives intergenic association of its target gene loci upon heat shock. Cell Rep. 26, 18–28 (2019).
Kainth, A. S., Chowdhary, S., Pincus, D. & Gross, D. S. Primordial super-enhancers: heat shock-induced chromatin organization in yeast. Trends Cell Biol. 31, 801–813 (2021).
McSwiggen, D. T. et al. Evidence for DNA-mediated nuclear compartmentalization distinct from phase separation. eLife https://doi.org/10.7554/eLife.47098 (2019).
Blobel, G. A., Higgs, D. R., Mitchell, J. A., Notani, D. & Young, R. A. Testing the super-enhancer concept. Nat. Rev. Genet. 22, 749–755 (2021).
Chong, S. et al. Tuning levels of low-complexity domain interactions to modulate endogenous oncogenic transcription. Mol. Cell https://doi.org/10.1016/j.molcel.2022.04.007 (2022).
Boehning, M. et al. RNA polymerase II clustering through carboxy-terminal domain phase separation. Nat. Struct. Mol. Biol. 25, 833–840 (2018).
Lu, H. et al. Phase-separation mechanism for C-terminal hyperphosphorylation of RNA polymerase II. Nature 558, 318–323 (2018).
Burke, K. A., Janke, A. M., Rhine, C. L. & Fawzi, N. L. Residue-by-residue view of in vitro FUS granules that bind the C-terminal domain of RNA polymerase II. Mol. Cell 60, 231–241 (2015).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature 572, 543–548 (2019).
Portz, B. & Shorter, J. Switching condensates: the CTD code goes liquid. Trends Biochem. Sci. 45, 1–3 (2020).
Quintero-Cadena, P., Lenstra, T. L. & Sternberg, P. W. RNA Pol II length and disorder enable cooperative scaling of transcriptional bursting. Mol. Cell 79, 207–220 (2020).
Henninger, J. E. et al. RNA-mediated feedback control of transcriptional condensates. Cell 184, 207–225 (2021).
Shao, W. et al. Phase separation of RNA-binding protein promotes polymerase binding and transcription. Nat. Chem. Biol. 18, 70–80 (2022).
Bi, X. et al. RNA targets ribogenesis factor WDR43 to chromatin for transcription and pluripotency control. Mol. Cell 75, 102–116 (2019).
Chen, G. et al. Taf14 recognizes a common motif in transcriptional machineries and facilitates their clustering by phase separation. Nat. Commun. 11, 4206 (2020).
Rencus-Lazar, S., DeRowe, Y., Adsi, H., Gazit, E. & Laor, D. Yeast models for the study of amyloid-associated disorders and development of future therapy. Front. Mol. Biosci. 6, 15 (2019).
Tuite, M. F. Yeast models of neurodegenerative diseases. Prog. Mol. Biol. Transl Sci.168, 351–379 (2019).
Zbinden, A., Perez-Berlanga, M., De Rossi, P. & Polymenidou, M. Phase separation and neurodegenerative diseases: a disturbance in the force. Dev. Cell 55, 45–68 (2020).
Darling, A. L. & Shorter, J. Combating deleterious phase transitions in neurodegenerative disease. Biochim. Biophys. Acta Mol. Cell. Res. 1868, 118984 (2021).
Sun, Z. et al. Molecular determinants and genetic modifiers of aggregation and toxicity for the ALS disease protein FUS/TLS. PLoS Biol. 9, e1000614 (2011).
Elden, A. C. et al. Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature 466, 1069–1075 (2010).
Shorter, J. Designer protein disaggregases to counter neurodegenerative disease. Curr. Opin. Genet. Dev. 44, 1–8 (2017).
Tariq, A. et al. Mining disaggregase sequence space to safely counter TDP-43, FUS, and α-synuclein proteotoxicity. Cell Rep. 28, 2080–2095 (2019).
Oldfield, C. J. & Dunker, A. K. Intrinsically disordered proteins and intrinsically disordered protein regions. Annu. Rev. Biochem. 83, 553–584 (2014).
Martin, E. W. et al. Valence and patterning of aromatic residues determine the phase behavior of prion-like domains. Science 367, 694–699 (2020).
Pak, C. W. et al. Sequence determinants of intracellular phase separation by complex coacervation of a disordered protein. Mol. Cell 63, 72–85 (2016).
Bremer, A. et al. Deciphering how naturally occurring sequence features impact the phase behaviours of disordered prion-like domains. Nat. Chem. 14, 196–207 (2022).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699 (2018).
Vernon, R. M. et al. Pi–Pi contacts are an overlooked protein feature relevant to phase separation. eLife https://doi.org/10.7554/eLife.31486 (2018).
Lin, Y., Currie, S. L. & Rosen, M. K. Intrinsically disordered sequences enable modulation of protein phase separation through distributed tyrosine motifs. J. Biol. Chem. 292, 19110–19120 (2017).
Murthy, A. C. et al. Molecular interactions underlying liquid–liquid phase separation of the FUS low-complexity domain. Nat. Struct. Mol. Biol. 26, 637–648 (2019).
Conicella, A. E. et al. TDP-43 α-helical structure tunes liquid–liquid phase separation and function. Proc. Natl Acad. Sci. USA 117, 5883–5894 (2020).
Conicella, A. E., Zerze, G. H., Mittal, J. & Fawzi, N. L. ALS mutations disrupt phase separation mediated by α-helical structure in the TDP-43 low-complexity C-terminal domain. Structure 24, 1537–1549 (2016).
Murray, D. T. et al. Structure of FUS protein fibrils and its relevance to self-assembly and phase separation of low-complexity domains. Cell 171, 615–627 (2017).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Kato, M., Zhou, X. & McKnight, S. L. How do protein domains of low sequence complexity work? RNA 28, 3–15 (2022).
Fawzi, N. L., Parekh, S. H. & Mittal, J. Biophysical studies of phase separation integrating experimental and computational methods. Curr. Opin. Struct. Biol. 70, 78–86 (2021).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Banani, S. F. et al. Compositional control of phase-separated cellular bodies. Cell 166, 651–663 (2016).
Mittag, T. & Pappu, R. V. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol. Cell 82, 2201–2214 (2022).
Brangwynne, C. P. Phase transitions and size scaling of membrane-less organelles. J. Cell Biol. 203, 875–881 (2013).
Dutagaci, B. et al. Charge-driven condensation of RNA and proteins suggests broad role of phase separation in cytoplasmic environments. eLife https://doi.org/10.7554/eLife.64004 (2021).
Dignon, G. L., Best, R. B. & Mittal, J. Biomolecular phase separation: from molecular driving forces to macroscopic properties. Annu. Rev. Phys. Chem. 71, 53–75 (2020).
Garcia-Jove Navarro, M. et al. RNA is a critical element for the sizing and the composition of phase-separated RNA–protein condensates. Nat. Commun. 10, 3230 (2019).
Guillen-Boixet, J. et al. RNA-induced conformational switching and clustering of G3BP drive stress granule assembly by condensation. Cell 181, 346–361 (2020).
Maharana, S. et al. RNA buffers the phase separation behavior of prion-like RNA binding proteins. Science 360, 918–921 (2018).
Ma, W., Zheng, G., Xie, W. & Mayr, C. In vivo reconstitution finds multivalent RNA–RNA interactions as drivers of mesh-like condensates. eLife https://doi.org/10.7554/eLife.64252 (2021).
Acknowledgements
We thank the members of the Bennett laboratory for useful discussions and B. Tu (UTSW) for feedback on sections of the review. Work in the Bennett laboratory is supported by NIAID grant nos AI141893, AI081704 and AI166869, and work in the Fawzi laboratory is supported by NSF BIO 1845734 and NINDS R01NS116176.
Author information
Authors and Affiliations
Contributions
M.I.S. wrote the initial draft, which was extensively revised by R.J.B. and C.F., with input from N.L.F.
Corresponding author
Ethics declarations
Competing interests
N.L.F. is a member of the scientific advisory board of Dewpoint Therapeutics.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Staples, M.I., Frazer, C., Fawzi, N.L. et al. Phase separation in fungi. Nat Microbiol 8, 375–386 (2023). https://doi.org/10.1038/s41564-022-01314-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01314-6
This article is cited by
-
Biomolecular condensates – extant relics or evolving microcompartments?
Communications Biology (2023)