Abstract
Co-phase separation of RNAs and RNA-binding proteins drives the biogenesis of ribonucleoprotein granules. RNAs can also undergo phase transitions in the absence of proteins. However, the physicochemical driving forces of protein-free, RNA-driven phase transitions remain unclear. Here we report that various types of RNA undergo phase separation with system-specific lower critical solution temperatures. This entropically driven phase separation is an intrinsic feature of the phosphate backbone that requires Mg2+ ions and is modulated by RNA bases. RNA-only condensates can additionally undergo enthalpically favourable percolation transitions within dense phases. This is enabled by a combination of Mg2+-dependent bridging interactions between phosphate groups and RNA-specific base stacking and base pairing. Phase separation coupled to percolation can cause dynamic arrest of RNAs within condensates and suppress the catalytic activity of an RNase P ribozyme. Our work highlights the need to incorporate RNA-driven phase transitions into models for ribonucleoprotein granule biogenesis.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data supporting this study are included in the Article and/or the Supplementary Information. Source data are provided with this paper.
References
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Li, P. et al. Phase transitions in the assembly of multivalent signalling proteins. Nature 483, 336–340 (2012).
Berry, J., Weber, S. C., Vaidya, N., Haataja, M. & Brangwynne, C. P. RNA transcription modulates phase transition-driven nuclear body assembly. Proc. Natl Acad. Sci. USA 112, E5237–E5245 (2015).
Woodruff, J. B. et al. The centrosome is a selective condensate that nucleates microtubules by concentrating tubulin. Cell 169, 1066–1077 (2017).
Zeng, M. et al. Phase transition in postsynaptic densities underlies formation of synaptic complexes and synaptic plasticity. Cell 166, 1163–1175 (2016).
Jiang, H. et al. Phase transition of spindle-associated protein regulate spindle apparatus assembly. Cell 163, 108–122 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Martin, E. W. et al. Valence and patterning of aromatic residues determine the phase behavior of prion-like domains. Science 367, 694–699 (2020).
Dzuricky, M., Rogers, B. A., Shahid, A., Cremer, P. S. & Chilkoti, A. De novo engineering of intracellular condensates using artificial disordered proteins. Nat. Chem. 12, 814–825 (2020).
Shin, Y. et al. Spatiotemporal control of intracellular phase transitions using light-activated optoDroplets. Cell 168, 159–171 (2017).
Nakamura, H. et al. Intracellular production of hydrogels and synthetic RNA granules by multivalent molecular interactions. Nat. Mater. 17, 79–89 (2018).
Zhao, E. M. et al. Light-based control of metabolic flux through assembly of synthetic organelles. Nat. Chem. Biol. 15, 589–597 (2019).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699 (2018).
Lin, Y.-H., Forman-Kay, J. D. & Chan, H. S. Sequence-specific polyampholyte phase separation in membraneless organelles. Phys. Rev. Lett. 117, 178101 (2016).
Alshareedah, I., Moosa, M. M., Raju, M., Potoyan, D. A. & Banerjee, P. R. Phase transition of RNA−protein complexes into ordered hollow condensates. Proc. Natl Acad. Sci. USA 117, 15650–15658 (2020).
Garcia-Jove Navarro, M. et al. RNA is a critical element for the sizing and the composition of phase-separated RNA–protein condensates. Nat. Commun. 10, 3230 (2019).
Roden, C. & Gladfelter, A. S. RNA contributions to the form and function of biomolecular condensates. Nat. Rev. Mol. Cell Biol. 22, 183–195 (2021).
Tauber, D., Tauber, G. & Parker, R. Mechanisms and regulation of RNA condensation in RNP granule formation. Trends Biochem. Sci. 45, 764–778 (2020).
Banerjee, P. R., Milin, A. N., Moosa, M. M., Onuchic, P. L. & Deniz, A. A. Reentrant phase transition drives dynamic substructure formation in ribonucleoprotein droplets. Angew. Chem. 129, 11512–11517 (2017).
Eisenberg, H. & Felsenfeld, G. Studies of the temperature-dependent conformation and phase separation of polyriboadenylic acid solutions at neutral pH. J. Mol. Biol. 30, 17–37 (1967).
Pullara, P., Alshareedah, I. & Banerjee, P. R. Temperature-dependent reentrant phase transition of RNA–polycation mixtures. Soft Matter 18, 1342–1349 (2022).
Van Treeck, B. et al. RNA self-assembly contributes to stress granule formation and defining the stress granule transcriptome. Proc. Natl Acad. Sci. USA 115, 2734–2739 (2018).
Boeynaems, S. et al. Spontaneous driving forces give rise to protein−RNA condensates with coexisting phases and complex material properties. Proc. Natl Acad. Sci. USA 116, 7889–7898 (2019).
Jain, A. & Vale, R. D. RNA phase transitions in repeat expansion disorders. Nature 546, 243–247 (2017).
Fay, M. M. & Anderson, P. J. The role of RNA in biological phase separations. J. Mol. Biol. 430, 4685–4701 (2018).
Guo, Q., Shi, X. & Wang, X. RNA and liquid–liquid phase separation. Noncoding RNA Res. 6, 92–99 (2021).
Polymenidou, M. The RNA face of phase separation. Science 360, 859–860 (2018).
Saha, S. & Hyman, A. A. RNA gets in phase. J. Cell Biol. 216, 2235–2237 (2017).
Weber, S. C. & Brangwynne, C. P. Getting RNA and protein in phase. Cell 149, 1188–1191 (2012).
Ma, Y. et al. Nucleobase clustering contributes to the formation and hollowing of repeat-expansion RNA condensate. J. Am. Chem. Soc. 144, 4716–4720 (2022).
Poudyal, R. R., Sieg, J. P., Portz, B., Keating, C. D. & Bevilacqua, P. C. RNA sequence and structure control assembly and function of RNA condensates. RNA 27, 1589–1601 (2021).
Tolokh, I. S. et al. Why double-stranded RNA resists condensation. Nucleic Acids Res. 42, 10823–10831 (2014).
Fay, M. M., Anderson, P. J. & Ivanov, P. ALS/FTD-associated C9ORF72 repeat RNA promotes phase transitions in vitro and in cells. Cell Reports 21, 3573–3584 (2017).
Gatchel, J. R. & Zoghbi, H. Y. Diseases of unstable repeat expansion: mechanisms and common principles. Nat. Rev. Genet. 6, 743–755 (2005).
Zhang, Y. et al. G-quadruplex structures trigger RNA phase separation. Nucleic Acids Res. 47, 11746–11754 (2019).
Choi, J.-M., Hyman, A. A. & Pappu, R. V. Generalized models for bond percolation transitions of associative polymers. Phys. Rev. E 102, 042403 (2020).
Choi, J.-M., Holehouse, A. S. & Pappu, R. V. Physical principles underlying the complex biology of intracellular phase transitions. Annu. Rev. Biophys. 49, 107–133 (2020).
Futscher, M. H., Philipp, M., Müller-Buschbaum, P. & Schulte, A. The role of backbone hydration of poly(N-isopropyl acrylamide) across the volume phase transition compared to its monomer. Sci. Rep. 7, 17012 (2017).
Halperin, A., Kröger, M. & Winnik, F. M. Poly(N-isopropylacrylamide) phase diagrams: fifty years of research. Angew. Chem. Int. Ed. 54, 15342–15367 (2015).
Tanaka, F. Theoretical study of molecular association and thermoreversible gelation in polymers. Polym. J. 34, 479–509 (2002).
Tanaka, F. In Molecular Gels: Materials with Self-Assembled Fibrillar Networks (eds. R.G. Weiss and P. Terech) 17–78 (Springer, 2006).
Tanaka, F. Polymer Physics: Applications to Molecular Association and Thermoreversible Gelation (Cambridge Univ. Press, 2011).
Rubinstein, M. & Dobrynin, A. V. Solutions of associative polymers. Trends in Polymer Science 5, 181–186 (1997).
Mittag, T. & Pappu, R. V. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol. Cell 82, 2201–2214 (2022).
Su, X. et al. Phase separation of signaling molecules promotes T cell receptor signal transduction. Science 352, 595–599 (2016).
Bhandari, K., Cotten, M. A., Kim, J., Rosen, M. K. & Schmit, J. D. Structure–function properties in disordered condensates. J. Phys. Chem. B 125, 467–476 (2021).
Bevilacqua, P. C., Williams, A. M., Chou, H.-L. & Assmann, S. M. RNA multimerization as an organizing force for liquid–liquid phase separation. RNA 28, 16–26 (2022).
Johansson, J. et al. An RNA thermosensor controls expression of virulence genes in Listeria monocytogenes. Cell 110, 551–561 (2002).
Puglisi, J. D. & Tinoco, I. Jr. In Methods in Enzymology (eds. J.E. Dahlberg and J.N. Abelson) Vol. 180, 304–325 (Elsevier, 1989).
Tinoco, I. Jr & Bustamante, C. How RNA folds. J. Mol. Biol. 293, 271–281 (1999).
Vicens, Q. & Kieft, J. S. Thoughts on how to think (and talk) about RNA structure. Proc. Natl Acad. Sci. USA 119, e2112677119 (2022).
Wienken, C. J., Baaske, P., Duhr, S. & Braun, D. Thermophoretic melting curves quantify the conformation and stability of RNA and DNA. Nucleic Acids Res. 39, e52 (2011).
Flory, P. J. Thermodynamics of high polymer solutions. J. Chem. Phys. 10, 51–61 (1942).
Huggins, M. L. Solutions of long chain compounds. J. Chem. Phys. 9, 440 (1941).
Ranganathan, S. & Shakhnovich, E. I. Dynamic metastable long-living droplets formed by sticker-spacer proteins. Elife 9, e56159 (2020).
Roberts, S. et al. Injectable tissue integrating networks from recombinant polypeptides with tunable order. Nat. Mater. 17, 1154–1163 (2018).
Onuchic, P. L., Milin, A. N., Alshareedah, I., Deniz, A. A. & Banerjee, P. R. Divalent cations can control a switch-like behavior in heterotypic and homotypic RNA coacervates. Sci. Rep. 9, 12161 (2019).
Merindol, R., Loescher, S., Samanta, A. & Walther, A. Pathway-controlled formation of mesostructured all-DNA colloids and superstructures. Nat. Nanotechnol. 13, 730–738 (2018).
Ruff, K. M., Roberts, S., Chilkoti, A. & Pappu, R. V. Advances in understanding stimulus-responsive phase behavior of intrinsically disordered protein polymers. J. Mol. Biol. 430, 4619–4635 (2018).
Ellis, K. J. & Morrison, J. F. In Methods in Enzymology (ed. D.L. Purich) Vol. 87, 405–426 (Elsevier, 1982).
Zeng, X., Holehouse, A. S., Chilkoti, A., Mittag, T. & Pappu, R. V. Connecting coil-to-globule transitions to full phase diagrams for intrinsically disordered proteins. Biophys. J. 119, 402–418 (2020).
Zeng, X. et al. Design of intrinsically disordered proteins that undergo phase transitions with lower critical solution temperatures. APL Mater. 9, 021119 (2021).
Amin, A. N., Lin, Y.-H., Das, S. & Chan, H. S. Analytical theory for sequence-specific binary fuzzy complexes of charged intrinsically disordered proteins. J. Phys. Chem. B 124, 6709–6720 (2020).
Dignon, G. L., Zheng, W., Best, R. B., Kim, Y. C. & Mittal, J. Relation between single-molecule properties and phase behavior of intrinsically disordered proteins. Proc. Natl Acad. Sci. USA 115, 9929–9934 (2018).
Lin, Y.-H. & Chan, H. S. Phase separation and single-chain compactness of charged disordered proteins are strongly correlated. Biophys. J. 112, 2043–2046 (2017).
May, S., Iglič, A., Reščič, J., Maset, S. & Bohinc, K. Bridging like-charged macroions through long divalent rodlike ions. J. Phys. Chem. B 112, 1685–1692 (2008).
Fossat, M. J., Zeng, X. & Pappu, R. V. Uncovering differences in hydration free energies and structures for model compound mimics of charged side chains of amino acids. J. Phys. Chem. B 125, 4148–4161 (2021).
Zeng, X., Ruff, K. M. & Pappu, R. V. Competing interactions give rise to two-state behavior and switch-like transitions in charge-rich intrinsically disordered proteins. Proc. Natl Acad. Sci. USA 119, e2200559119 (2022).
Nguyen, H. T., Hori, N. & Thirumalai, D. Condensates in RNA repeat sequences are heterogeneously organized and exhibit reptation dynamics. Nat. Chem. 14, 775–785 (2022).
Harmon, T. S., Holehouse, A. S., Rosen, M. K. & Pappu, R. V. Intrinsically disordered linkers determine the interplay between phase separation and gelation in multivalent proteins. Elife 6, e30294 (2017).
Rubinstein, M. & Semenov, A. N. Thermoreversible gelation in solutions of associating polymers. 2. Linear dynamics. Macromolecules 31, 1386–1397 (1998).
Phan, H.-D., Lai, L. B., Zahurancik, W. J. & Gopalan, V. The many faces of RNA-based RNase P, an RNA-world relic. Trends Biochem. Sci. 46, 976–991 (2021).
Gopalan, V., Vioque, A. & Altman, S. RNase P: variations and uses. J. Biol. Chem. 277, 6759–6762 (2002).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Cho, I.-M., Lai, L. B., Susanti, D., Mukhopadhyay, B. & Gopalan, V. Ribosomal protein L7Ae is a subunit of archaeal RNase P. Proc. Natl Acad. Sci. USA 107, 14573–14578 (2010).
Phan, H.-D. et al. Elucidation of structure–function relationships in Methanocaldococcus jannaschii RNase P, a multi-subunit catalytic ribonucleoprotein. Nucleic Acids Res. 50, 8154–8167 (2022).
Pulukkunat, D. K. & Gopalan, V. Studies on Methanocaldococcus jannaschii RNase P reveal insights into the roles of RNA and protein cofactors in RNase P catalysis. Nucleic Acids Res. 36, 4172–4180 (2008).
Tsai, H.-Y., Pulukkunat, D. K., Woznick, W. K. & Gopalan, V. Functional reconstitution and characterization of Pyrococcus furiosus RNase P. Proc. Natl Acad. Sci. USA 103, 16147–16152 (2006).
Wan, F. et al. Cryo-electron microscopy structure of an archaeal ribonuclease P holoenzyme. Nat. Commun. 10, 2617 (2019).
Marathe, I. A. et al. Protein cofactors and substrate influence Mg2+-dependent structural changes in the catalytic RNA of archaeal RNase P. Nucleic Acids Res. 49, 9444–9458 (2021).
Loughrey, D., Watters, K. E., Settle, A. H. & Lucks, J. B. SHAPE-Seq 2.0: systematic optimization and extension of high-throughput chemical probing of RNA secondary structure with next generation sequencing. Nucleic Acids Res. 42, e165 (2014).
Denesyuk, N. A. & Thirumalai, D. Coarse-grained model for predicting RNA folding thermodynamics. J. Phys. Chem. B 117, 4901–4911 (2013).
Buchmueller, K. L. & Weeks, K. M. Tris-borate is a poor counterion for RNA: a cautionary tale for RNA folding studies. Nucleic Acids Res. 32, e184 (2004).
Iglesias-Artola, J. M. et al. Charge-density reduction promotes ribozyme activity in RNA–peptide coacervates via RNA fluidization and magnesium partitioning. Nat. Chem. 14, 407–416 (2022).
Poudyal, R. R. et al. Template-directed RNA polymerization and enhanced ribozyme catalysis inside membraneless compartments formed by coacervates. Nat. Commun. 10, 490 (2019).
Higgs, P. G. & Lehman, N. The RNA World: molecular cooperation at the origins of life. Nat. Rev. Genet. 16, 7–17 (2015).
Drobot, B. et al. Compartmentalised RNA catalysis in membrane-free coacervate protocells. Nat. Commun. 9, 3643 (2018).
Wiedner, H. J. & Giudice, J. It’s not just a phase: function and characteristics of RNA-binding proteins in phase separation. Nat. Struct. Mol. Biol. 28, 465–473 (2021).
Van Treeck, B. & Parker, R. Emerging roles for intermolecular RNA–RNA interactions in RNP assemblies. Cell 174, 791–802 (2018).
Cheng, Y. et al. Increased Alu RNA processing in Alzheimer brains is linked to gene expression changes. EMBO Rep. 22, e52255 (2021).
Lin, C.-L. G. et al. Aberrant RNA processing in a neurodegenerative disease: the cause for absent EAAT2, a glutamate transporter, in amyotrophic lateral sclerosis. Neuron 20, 589–602 (1998).
Tank, E. M. et al. Abnormal RNA stability in amyotrophic lateral sclerosis. Nat. Commun. 9, 2845 (2018).
Tsai, H.-Y., Lai, L. B. & Gopalan, V. A modified pBluescript-based vector for facile cloning and transcription of RNAs. Anal. Biochem. 303, 214–217 (2002).
Taylor, N. O., Wei, M.-T., Stone, H. A. & Brangwynne, C. P. Quantifying dynamics in phase-separated condensates using fluorescence recovery after photobleaching. Biophys. J. 117, 1285–1300 (2019).
Sousa da Silva, A. W. & Vranken, W. F. ACPYPE—antechamber Python parser interface. BMC Res. Notes 5, 367 (2012).
Vanquelef, E. et al. R.E.D. Server: a web service for deriving RESP and ESP charges and building force field libraries for new molecules and molecular fragments. Nucleic Acids Res. 39, W511–W517 (2011).
Bayly, C. I., Cieplak, P., Cornell, W. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. J. Phys. Chem. 97, 10269–10280 (1993).
Duan, Y. et al. A point‐charge force field for molecular mechanics simulations of proteins based on condensed‐phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012 (2003).
Lee, C., Yang, W. & Parr, R. G. Development of the Colle–Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 37, 785 (1988).
Becke, A. D. Density‐functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
Kendall, R. A., Dunning, T. H. Jr & Harrison, R. J. Electron affinities of the first‐row atoms revisited. Systematic basis sets and wave functions. J. Chem. Phys. 96, 6796–6806 (1992).
Steinbrecher, T., Latzer, J. & Case, D. Revised AMBER parameters for bioorganic phosphates. J. Chem. Theory Comput. 8, 4405–4412 (2012).
Bergonzo, C. & Cheatham, T. E. III Improved force field parameters lead to a better description of RNA structure. J. Chem. Theory Comput. 11, 3969–3972 (2015).
Grotz, K. K., Cruz-León, S. & Schwierz, N. Optimized magnesium force field parameters for biomolecular simulations with accurate solvation, ion-binding, and water-exchange properties. J. Chem. Theory Comput. 17, 2530–2540 (2021).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Páll, S., Abraham, M. J., Kutzner, C., Hess, B. & Lindahl, E. Tackling exascale software challenges in molecular dynamics simulations with GROMACS. In International Conference on Exascale Applications and Software 2014 (eds. S. Markidis and E. Laure) 3–27 (Springer, 2014).
GROMACS 2021 manual. GROMACS development team https://doi.org/10.5281/zenodo.4457591 (2021).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Nosé, S. & Klein, M. Constant pressure molecular dynamics for molecular systems. Mol. Phys. 50, 1055–1076 (1983).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅ log (N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hess, B., Bekker, H., Berendsen, H. J. & Fraaije, J. G. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Zgarbová, M. et al. Refinement of the Cornell et al. nucleic acids force field based on reference quantum chemical calculations of glycosidic torsion profiles. J. Chem. Theory Comput. 7, 2886–2902 (2011).
Ferrenberg, A. M. & Swendsen, R. H. Optimized Monte Carlo data analysis. Comput. Phys. 3, 101–104 (1989).
Gallicchio, E., Andrec, M., Felts, A. K. & Levy, R. M. Temperature weighted histogram analysis method, replica exchange, and transition paths. J. Phys. Chem. B 109, 6722–6731 (2005).
Acknowledgements
This work was supported by the National Institute of General Medical Sciences of the National Institutes of Health (R35 GM138186 to P.R.B., GM120582 to V.G.), the Air Force Office of Scientific Research (FA9550-20-1-0241 to R.V.P.), the St. Jude Research Collaborative on Biophysics of RNP granules (to P.R.B. and R.V.P.) and the Henry M. Jackson Foundation for the Advancement of Military Medicine (USUHS subaward 5516 to V.G.). W.J.Z. gratefully acknowledges a Pelotonia postdoctoral fellowship from the OSU Comprehensive Cancer Center. The authors acknowledge members of the Banerjee, Gopalan and Pappu labs for valuable discussions during different stages of the manuscript preparation. X.Z. thanks A. A. Chen and F.-Y. Dupradeau for helpful discussions on the forcefield parameters used in this work, and S. Tahan for support in the use of the RIS cluster at Washington University in St. Louis.
Author information
Authors and Affiliations
Contributions
P.R.B. conceived the idea for this study. P.R.B. and G.M.W. designed the study with input from L.B.L., W.J.Z., V.G., X.Z. and R.V.P. G.M.W. performed the RNA phase separation experiments and data analysis with assistance from P.P. L.B.L, V.S. and W.J.Z. synthesized the different RNAs and modified them with fluorescent labels. W.J.Z. performed all the RNase P activity experiments. X.Z. and R.V.P. performed the all-atom molecular dynamics simulations, analysed the results and developed the framework to explain the observed phenomenology. All authors contributed to the writing and revision of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Hue Sun Chan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phase separation of homopolymeric RNAs.
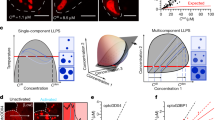
(a) A schematic of temperature-controlled microscopy assay to probe RNA phase separation. (b) Phase separation and arrest of poly(rA) upon heating. Brightfield images of 1.5 mg/ml−1 poly(rA) in 25 mM Tris-HCl (pH 7.5 at 25 °C), 5 mM MgCl2 (25T-5M buffer) during heating (red arrow) and cooling (cyan arrow) as indicated; observed LCPT of this sample is 39.3 ± 8.5 °C. Upon cooling, poly(rA) droplets did not dissolve. (c) Brightfield images of 1.5 mg/ml−1 poly(rG) in 25 mM Tris-HCl (pH 7.5 at 25 °C), 1 mM MgCl2 (25T-1M buffer) at two different temperatures. Extensive irreversible aggregation is evident. (d) LCST-type phase separation of poly(P). Brightfield images of 1.5 mg/ml−1 poly(P) in 25 mM Tris-HCl (pH 7.5 at 25 °C), 250 mM MgCl2 (25T-250M buffer) during heating (red arrow) and cooling (cyan arrow) as indicated; observed LCPT of this sample is 35.3 ± 5.0 °C. Buffer notation used: the number in front of ‘T’ indicates [Tris-HCl] and the number in front of ‘M’ indicates [Mg2+] in mM in each buffer. Error bars represent s.e.m. for n = 3 replicates.
Extended Data Fig. 2 An emerging model for RNA phase separation coupled percolation (PSCP) behaviour.
(a) The observed hierarchy of LCST-type phase transition propensity of RNA bases and the phosphate backbone. (b) An enthalpic model of RNA phase separation. Here, RNA in an ensemble of minimum free energy structures (left) is heated, thereby denaturing the RNA. Subsequent cooling below Tph should enable the RNA to undergo a UCST transition. This model implies that enthalpic interactions such as hydrogen bonding and base-stacking drive RNA phase separation. (c) PSCP model of RNA condensation. In this model desolvation entropy drives the self-association of RNA to form phase-separated condensates upon heating. During subsequent cooling, the rank order of the phase separation temperature (LCPT or Tph) and the percolation temperature (Tprc) determines refolding vs. condensate arrest. When Tph>Tprc, the system undergoes reversible phase separation whereas in cases of Tph< Tprc, the system shows hysteretic phase behaviour. Our experimental and computational results clearly show that RNA condensation proceeds via this pathway.
Supplementary information
Supplementary Information
Supplementary Figs. 1–21, Tables 1 and 2, legends for Videos 1–19 and references.
Supplementary Video 1
Phase separation of poly(rU). A sample of 1.5 mg ml−1 poly(rU) in 25 mM Tris-HCl (pH 7.5 at 25 °C) and 400 mM Mg2+ underwent thermal cycling via temperature-controlled microscopy. The poly(rU) phase separated reversibly with a UCPT of 24.2 ± 0.5 °C and an LCPT of 1.1 ± 0.3 °C (n = 3 replicates).
Supplementary Video 2
Phase separation of poly(rU). A sample of 1.5 mg ml−1 poly(rU) in 25 mM Tris-HCl (pH 7.5 at 25 °C) and 500 mM Mg2+ underwent thermal cycling via temperature-controlled microscopy. The poly(rU) phase separated reversibly with a UCPT of 25.2 ± 1.2 °C (n = 3 replicates) and no LCST was observed.
Supplementary Video 3
Phase separation of poly(rC). A sample of 1.5 mg ml−1 poly(rC) in 25 mM Tris-HCl (pH 7.5 at 25 °C) and 100 mM Mg2+ underwent thermal cycling via temperature-controlled microscopy. The poly(rC) phase separated reversibly with an LCPT of 57.6 ± 1.9 °C (n = 3 replicates).
Supplementary Video 4
Phase separation of poly(rA). A sample of 1.5 mg ml−1 poly(rA) in 25 mM Tris-HCl (pH 7.5 at 25 °C) and 5 mM Mg2+ underwent thermal cycling via temperature-controlled microscopy going to 80 °C. The poly(rA) phase separated irreversibly with an LCPT of 39.3 ± 8.5 °C (n = 3 replicates).
Supplementary Movie 5
Phase separation of poly(rA). A sample of 1.5 mg ml−1 poly(rA) in 25 mM Tris-HCl (pH 7.5 at 25 °C) and 5 mM Mg2+ underwent thermal cycling via temperature-controlled microscopy going to 34 °C. The poly(rA) phase separated irreversibly with an LCPT of 32.7 °C in this trial.
Supplementary Video 6
Thermal cycling of poly(rG). A sample of 1.5 mg ml−1 poly(rG) in 25 mM Tris-HCl (pH 7.5 at 25 °C) and 1 mM Mg2+ underwent thermal cycling via temperature-controlled microscopy going to 80 °C. The poly(rG) remained aggregated at all temperatures.
Supplementary Video 7
Movie 7. Phase separation of poly(P). A sample of 1.5 mg ml−1 poly(P) in 25 mM Tris-HCl (pH 7.5 at 25 °C) and 250 mM Mg2+ underwent thermal cycling via temperature-controlled microscopy. The poly(P) phase separated with an LCPT of 35.3 ± 5.0 °C (n = 3 replicates).
Supplementary Video 8
Phase separation of (CAG)31 RNA. A sample of 10 µM (CAG)31 in 25 mM Tris-HCl (pH 7.5 at 25 °C), 10 mM Mg2+ and 10 mM Na+ underwent thermal cycling via temperature-controlled microscopy. Observed LCPT = 66.8 ± 3.9 °C (n = 3 replicates).
Supplementary Video 9
Dynamic arrest via percolation of (CAG)31 RNA upon phase separation. A sample of 100 µM (CAG)31 in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C), 50 mM Mg2+ and 25 mM Na+ underwent phase separation at 38.0 ± 3.5 °C (n = 3 replicates) followed by the formation of an extensive percolated network of aspherical droplets. These droplets relaxed and merged into spherical droplets as the temperature was increased to 80 °C.
Supplementary Video 10
Rapid fusion of (CAG)31 droplets. A sample of 50 µM (CAG)31 phase separated at 41.1 ± 1.7 °C in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C) and 50 mM Mg2+. The droplets underwent rapid shape relaxation through coalescence as the temperature was increased above 60 °C.
Supplementary Video 11
Dissolution of (CAG)31 droplets upon addition of EDTA. A sample of 10 µM (CAG)31 RNA underwent annealing in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C) and 50 mM Mg2+. The droplets dissolved upon the addition of a small volume of 500 mM EDTA from the left of the imaging area.
Supplementary Video 12
Phase separation and arrest of a scrambled (CAG)31 sequence. A sample of 50 µM scrambled (CAG)31 is shown in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C), 50 mM Mg2+ and 25 mM Na+. An irreversible phase transition was observed with an LCPT at 57.9 ± 4.2 °C (n = 3 replicates).
Supplementary Video 13
Phase separation and arrest of (CAG)20. A sample of 155 µM (CAG)20 was prepared in 10 mM Tris-HCl (pH 7.5 at 25 °C) and 200 mM Mg2+. Irreversible phase separation was observed at 59.9 ± 2.3 °C (n = 3 replicates).
Supplementary Video 14
Reversible phase separation of (CUG)31. A sample of 50 µM (CUG)31 RNA underwent thermal cycling with an LCPT of 72.7 ± 5.4 °C in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C), 50 mM Mg2+ and 25 mM Na+. The droplets dissolved after crossing the LCPT as the temperature decreased. The same conditions for (CAG)31 resulted in irreversible droplet formation.
Supplementary Video 15
Absence of phase separation for (CUU)31. A sample of 50 µM (CUU)31 RNA underwent thermal cycling in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C), 50 mM Mg2+ and 25 mM Na+. No phase separation was observed.
Supplementary Video 16
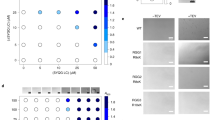
Phase separation and percolation of Pfu RNase P RNA. A sample of 10 µM Pfu RPR was imaged in a buffer containing 50 mM HEPES–KOH (pH 7.5 at 25 °C) and 50 mM Mg2+. Irreversible phase separation was observed at 68.2 ± 1.3 °C (n = 3 replicates).
Supplementary Video 17
Phase separation and percolation of Mja RNase P RNA. A sample of 10 µM Mja RPR was imaged in a buffer containing 50 mM Tris-HCl (pH 7.5 at 25 °C) and 50 mM Mg2+. Irreversible phase separation was observed at 65.9 ± 3.2 °C (n = 3 replicates).
Supplementary Video 18
Phase separation and percolation of Mma RNase P RNA. A sample of 10 µM Mma RPR was imaged in a buffer containing 50 mM Tris-HCl (pH 7.5 at 25 °C) and 50 mM Mg2+. Irreversible phase separation was observed at 54.2 ± 0.9 °C (n = 3 replicates).
Supplementary Video 19
Melting of arrested Pfu RPR droplets. Droplets were prepared via annealing in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C), 25 mM Mg2+ and 10 mM Na+. The arrested Pfu RPR droplets then underwent thermal cycling via temperature-controlled microscopy showing the subsequent relaxation of the Pfu RPR condensates into spherical droplets.
Supplementary Video 20
FRAP of Pfu RPR. A sample of 10 µM Pfu RPR with 10 mM Tris-HCl (pH 7.5 at 25 °C) and 50 mM Mg2+ was used for FRAP experiments (see Methods for additional details).
Supplementary Video 21
FRAP of Mja RPR. A sample of 10 µM Mja RPR with 10 mM Tris-HCl (p 7.5 at 25 °C) and 50 mM Mg2+ was used for FRAP experiments (see Methods for additional details).
Supplementary Video 22
FRAP of Mma RPR. A sample of 10 µM Mma RPR with 10 mM Tris-HCl (pH 7.5 at 25 °C) and 50 mM Mg2+ was used for FRAP experiments (see Methods for additional details).
Supplementary Video 23
Reversible phase separation of Mja RPR in a refolding buffer. A sample of 20 µM RNA was prepared via a refolding protocol and diluted into refolding buffer containing 50 mM HEPES–KOH (pH 8.0 at 25 °C), 10 mM Mg2+ and 800 mM AmAc upon reaching 37 °C. Reversible phase separation was observed with an LCPT of 86.3 ± 3.1 °C (n = 3 replicates).
Supplementary Video 24
Reversible phase separation of Mma RPR in a refolding buffer. A sample of 20 µM RNA was prepared via a refolding protocol and diluted into refolding buffer containing 50 mM Tris-HCl (pH 7.5 at 25 °C), 7.5 mM Mg2+ and 500 mM AmAc upon reaching 37 °C. Reversible phase separation was observed with an LCPT of 49.2 ± 3.2 °C (n = 3 replicates).
Supplementary Video 25
Absence of an apparent phase separation of Pfu RPR in a refolding buffer. A sample of 20 µM RNA was prepared via a refolding protocol and diluted into refolding buffer containing working concentrations of 50 mM HEPES–KOH (pH 8.4 at 25 °C), 10 mM Mg2+ and 800 mM AmAc upon reaching 37 °C.
Supplementary Video 26
Absence of an apparent phase separation of Pfu RPR in a refolding buffer. A sample of 2.5 µM Pfu RPR was prepared in a buffer containing 50 mM HEPES–KOH (pH 7.5 at 25 °C), 10 mM Mg2+ and 800 mM AmAc. The sample was heated to 80 °C.
Supplementary Video 27
Phase separation of Pfu RPR in the activity assay buffer. A sample of 0.625 µM RNA was diluted into buffer containing working concentrations of 50 mM HEPES–KOH (pH 7.5 at 55 °C) and 500 mM Mg2+. During thermal cycling via temperature-controlled microscopy, irreversible phase separation was observed with an LCPT of 70.6 ± 2.6 °C (n = 3 replicates).
Supplementary Video 28
Phase separation of Pfu RPR in the activity assay buffer. A sample of 0.625 µM Pfu RPR was diluted into buffer containing working concentrations of 50 mM HEPES–KOH (pH 7.5 at 55 °C), 500 mM Mg2+ and 2 M AmAc. No phase separation was observed during thermal cycling via temperature-controlled microscopy (n = 3 replicates).
Supplementary Video 29
Phase separation of (CAG)31 RNA in the presence of dimethylsulfoxide (DMSO). A sample of 10 µM (CAG)31 RNA underwent thermal cycling in a buffer containing 10 mM Tris-HCl (pH 7.5 at 25 °C) and 50 mM Mg2+, with the addition of 5% (v/v) DMSO.
Supplementary Code 1
P values and significance for Pfu turnover assay.
Supplementary Data 1
Supplementary Information source data simulations as delimited text files.
Supplementary Data 2
Source data for Supplementary Information figures, for example, statistical source data.
Supplementary Data 3
Raw gel data.
Supplementary Data 4
Raw gel data.
Supplementary Data 5
Source data simulations for Fig. 2.
Source data
Source Data Figs. 1–6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wadsworth, G.M., Zahurancik, W.J., Zeng, X. et al. RNAs undergo phase transitions with lower critical solution temperatures. Nat. Chem. 15, 1693–1704 (2023). https://doi.org/10.1038/s41557-023-01353-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-023-01353-4
This article is cited by
-
A charge-dependent long-ranged force drives tailored assembly of matter in solution
Nature Nanotechnology (2024)