Abstract
Breast cancer brain metastasis (BCBM) is a lethal disease with no effective treatments. Prior work has shown that brain cancers and metastases are densely infiltrated with anti-inflammatory, protumourigenic tumour-associated macrophages, but the role of brain-resident microglia remains controversial because they are challenging to discriminate from other tumour-associated macrophages. Using single-cell RNA sequencing, genetic and humanized mouse models, we specifically identify microglia and find that they play a distinct pro-inflammatory and tumour-suppressive role in BCBM. Animals lacking microglia show increased metastasis, decreased survival and reduced natural killer and T cell responses, showing that microglia are critical to promote anti-tumour immunity to suppress BCBM. We find that the pro-inflammatory response is conserved in human microglia, and markers of their response are associated with better prognosis in patients with BCBM. These findings establish an important role for microglia in anti-tumour immunity and highlight them as a potential immunotherapy target for brain metastasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession codes GSE147949 and GSE237386. Reference genome GRCh38/mm10 are available from Ensembl. MULTI-seq reads were processed according to the MULTI-seq protocol64 and available on GitHub (https://github.com/chris-mcginnis-ucsf/MULTIseq). Qptiff images were segmented in QuPath using StarDist and is available on GitHub (https://github.com/stardist/stardist). All other data are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Witzel, I., Oliveira-Ferrer, L., Pantel, K., Müller, V. & Wikman, H. Breast cancer brain metastases: biology and new clinical perspectives. Breast Cancer Res. 18, 1–9 (2016).
Ostrom, Q. T., Wright, C. H. & Barnholtz-Sloan, J. S. Brain Metastases: Epidemiology. Handbook of Clinical Neurology vol. 149 (Elsevier, 2018).
Niikura, N. et al. Treatment outcomes and prognostic factors for patients with brain metastases from breast cancer of each subtype: a multicenter retrospective analysis. Breast Cancer Res. Treat. 147, 103–112 (2014).
Brufsky, A. M. et al. Central nervous system metastases in patients with HER2-positive metastatic breast cancer: incidence, treatment, and survival in patients from registHER. Clin. Cancer Res. 17, 4834–4843 (2011).
Rostami, R., Mittal, S., Rostami, P., Tavassoli, F. & Jabbari, B. Brain metastasis in breast cancer: a comprehensive literature review. J. Neuro-Oncol. https://doi.org/10.1007/s11060-016-2075-3 (2016).
Martin, A. M. et al. Immunotherapy and symptomatic radiation necrosis in patients with brain metastases treated with stereotactic radiation. JAMA Oncol. 4, 1123–1124 (2018).
Deeken, J. F. & Löscher, W. The blood–brain barrier and cancer: transporters, treatment, and Trojan horses. Clin. Cancer Res. 13, 1663–1674 (2007).
Tosoni, A., Ermani, M. & Brandes, A. A. The pathogenesis and treatment of brain metastases: a comprehensive review. Crit. Rev. Oncol. Hematol. 52, 199–215 (2004).
Hanisch, U. K. & Kettenmann, H. Microglia: active sensor and versatile effector cells in the normal and pathologic brain. Nat. Neurosci. https://doi.org/10.1038/nn1997 (2007).
Wolf, S. A., Boddeke, H. W. G. M. & Kettenmann, H. Microglia in physiology and disease. Annu. Rev. Physiol. 79, 619–643 (2017).
Hammond, T. R., Robinton, D. & Stevens, B. Microglia and the brain: complementary partners in development and disease. Annu. Rev. Cell Dev. Biol. 34, 523–544 (2018).
Quail, D. F. & Joyce, J. A. The microenvironmental landscape of brain tumors. Cancer Cell 31, 326–341 (2017).
Goldmann, T. et al. Origin, fate and dynamics of macrophages at central nervous system interfaces. Nat. Immunol. https://doi.org/10.1038/ni.3423 (2016).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, Aging, and disease. Immunity 48, 380–395.e6 (2018).
Jordão, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science https://doi.org/10.1126/science.aat7554 (2019).
Duchnowska, R. et al. Immune response in breast cancer brain metastases and their microenvironment: the role of the PD-1/PD-L axis. Breast Cancer Res. 18, 43 (2016).
Coniglio, S. J. et al. Microglial stimulation of glioblastoma invasion involves epidermal growth factor receptor (EGFR) and colony stimulating factor 1 receptor (CSF-1R) signaling. Mol. Med. https://doi.org/10.2119/molmed.2011.00217 (2012).
Pyonteck, S. M. et al. CSF-1R inhibition alters macrophage polarization and blocks glioma progression. Nat. Med. 19, 1264–1272 (2013).
Quail, D. F. et al. The tumor microenvironment underlies acquired resistance to CSF-1R inhibition in gliomas. Science https://doi.org/10.1126/science.aad3018 (2016).
Yan, D. et al. Inhibition of colony stimulating factor-1 receptor abrogates microenvironment-mediated therapeutic resistance in gliomas. Oncogene https://doi.org/10.1038/onc.2017.261 (2017).
Qiao, S., Qian, Y., Xu, G., Luo, Q. & Zhang, Z. Long-term characterization of activated microglia/macrophages facilitating the development of experimental brain metastasis through intravital microscopic imaging. J. Neuroinflammation https://doi.org/10.1186/s12974-018-1389-9 (2019).
Guldner, I. H. et al. CNS-native myeloid cells drive immune suppression in the brain metastatic niche through Cxcl10. Cell https://doi.org/10.1016/j.cell.2020.09.064 (2020).
Prinz, M. & Priller, J. Tickets to the brain: role of CCR2 and CX3CR1 in myeloid cell entry in the CNS. J. Neuroimmunol. 224, 80–84 (2010).
Rojo, R. et al. Deletion of a Csf1r enhancer selectively impacts CSF1R expression and development of tissue macrophage populations. Nat. Commun. 10, 3215 (2019).
Bos, P. D. et al. Genes that mediate breast cancer metastasis to the brain. Nature https://doi.org/10.1038/nature08021 (2009).
Lorger, M. & Felding-Habermann, B. Capturing changes in the brain microenvironment during initial steps of breast cancer brain metastasis. Am. J. Pathol. 176, 2958–2971 (2010).
Kienast, Y. et al. Real-time imaging reveals the single steps of brain metastasis formation. Nat. Med. 16, 116–122 (2010).
Valiente, M. et al. Serpins promote cancer cell survival and vascular co-option in brain metastasis. Cell https://doi.org/10.1016/j.cell.2014.01.040 (2014).
Kettenmann, H., Hanisch, U.-K., Noda, M. & Verkhratsky, A. Physiology of microglia. Physiol. Rev. https://doi.org/10.1152/physrev.00011.2010 (2011).
Bowman, R. L. et al. Macrophage ontogeny underlies differences in tumor-specific education in brain malignancies. Cell Rep. 17, 2445–2459 (2016).
O’Flanagan, C. H. et al. Dissociation of solid tumor tissues with cold active protease for single-cell RNA-seq minimizes conserved collagenase-associated stress responses. Genome Biol. 20, 1–13 (2019).
Motenko, H., Neuhauser, S. B., O’Keefe, M. & Richardson, J. E. MouseMine: a new data warehouse for MGI. Mamm. Genome 26, 325–330 (2015).
Dey, K. K., Hsiao, C. J. & Stephens, M. Visualizing the structure of RNA-seq expression data using grade of membership models. PLoS Genet. 13, 1–23 (2017).
Mathys, H. et al. Temporal tracking of microglia activation in neurodegeneration at. single-cell resolution. Cell Rep. 21, 366–380 (2017).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e6 (2019).
Ochocka, N. et al. Single-cell RNA sequencing reveals functional heterogeneity of glioma-associated brain macrophages. Nat. Commun. 12, 1–14 (2021).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14, 36–49 (2014).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Blasius, A. L. et al. Bone marrow stromal cell antigen 2 is a specific marker of type I IFN-producing cells in the naive mouse, but a promiscuous cell surface antigen following IFN stimulation. J. Immunol. https://doi.org/10.4049/jimmunol.177.5.3260 (2006).
Neil, S. J. D., Zang, T. & Bieniasz, P. D. Tetherin inhibits retrovirus release and is antagonized by HIV-1 Vpu. Nature https://doi.org/10.1038/nature06553 (2008).
Ting, J. P. Y. & Trowsdale, J. Genetic control of MHC class II expression. Cell https://doi.org/10.1016/s0092-8674(02)00696-7 (2002).
Schröder, B. The multifaceted roles of the invariant chain CD74—more than just a chaperone. Biochim. Biophys. Acta 1863, 1269–1281 (2016).
Stanley, E. R. et al. Biology and action of colony-stimulating factor-1. Mol. Reprod. Dev. 46, 4–10 (1997).
Tokunaga, R. et al. CXCL9, CXCL10, CXCL11/CXCR3 axis for immune activation—a target for novel cancer therapy. Cancer Treat. Rev. 63, 40–47 (2018).
Spangenberg, E. et al. Sustained microglial depletion with CSF1R inhibitor impairs parenchymal plaque development in an Alzheimer’s disease model. Nat. Commun. https://doi.org/10.1038/s41467-019-11674-z (2019).
Munro, D. A. D. et al. CNS macrophages differentially rely on an intronic Csf1r enhancer for their development. Development 147, dev194449 (2020).
McQuade, A. et al. Development and validation of a simplified method to generate human microglia from pluripotent stem cells. Mol. Neurodegener. https://doi.org/10.1186/s13024-018-0297-x (2018).
Hasselmann, J. et al. Development of a chimeric model to study and manipulate human microglia in vivo. Neuron https://doi.org/10.1016/j.neuron.2019.07.002 (2019).
Rongvaux, A. et al. Development and function of human innate immune cells in a humanized mouse model. Nat. Biotechnol. https://doi.org/10.1038/nbt.2858 (2014).
Varešlija, D. et al. Transcriptome characterization of matched primary breast and brain metastatic tumors to detect novel actionable targets. J. Natl Cancer Inst. 111, 388–398 (2019).
Marsh, S. E. et al. Dissection of artifactual and confounding glial signatures by single-cell sequencing of mouse and human brain. Nat. Neurosci. 25, 306–316 (2022).
Ocañas, S. R. et al. Minimizing the ex vivo confounds of cell-isolation techniques on transcriptomic and translatomic profiles of purified microglia. eNeuro 9, ENEURO.0348- (2022).
Klemm, F. et al. Compensatory CSF2-driven macrophage activation promotes adaptive resistance to CSF1R inhibition in breast-to-brain metastasis. Nat. Cancer https://doi.org/10.1038/s43018-021-00254-0 (2021).
Huang, Y. et al. Repopulated microglia are solely derived from the proliferation of residual microglia after acute depletion. Nat. Neurosci. 21, 530–540 (2018).
Lund, H. et al. Competitive repopulation of an empty microglial niche yields functionally distinct subsets of microglia-like cells. Nat. Commun. 9, 4845 (2018).
Bruttger, J. et al. Genetic cell ablation reveals clusters of local self-renewing microglia in the mammalian central nervous system. Immunity 43, 92–106 (2015).
Han, J., Harris, R. A. & Zhang, X. M. An updated assessment of microglia depletion: current concepts and future directions. Mol. Brain 10, 1–8 (2017).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. https://doi.org/10.1038/nn.3599 (2014).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science https://doi.org/10.1126/science.aal3222 (2017).
Campbell, J. P., Merkel, A. R., Masood-Campbell, S. K., Elefteriou, F. & Sterling, J. A. Models of bone metastasis. J. Vis. Exp. https://doi.org/10.3791/4260 (2012).
Graphpad Prism. https://www.graphpad.com (2023).
stardist. GitHub https://github.com/stardist/stardist (2018).
McGinnis, C. S. et al. MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat. Methods https://doi.org/10.1038/s41592-019-0433-8 (2019).
MULTI-seq. GitHub. https://github.com/chris-mcginnis-ucsf/MULTIseq (2021).
Living Image. https://www.perkinelmer.com/category/in-vivo-imaging-software (2017).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinf. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
jnc_2018. GitHub https://github.com/npriedig/jnci_2018/blob/master/brainMetPairs.salmon.cts.txt (2018).
Acknowledgements
Research reported in this publication was supported by the National Institute of Health/ National Cancer Institute award number R01CA237376-01A1, to D.A.L., the American Cancer Society award number IRG-98-279-10 to D.A.L., the Center for Complex Biological Systems, University of California, Irvine NIH/NCI U54-CA-217378 to D.A.L. and M.B.-J., the NIH/NINDS award number R01NS120060 to S.A.V., the National Institutes of Health T32AI 0602573, the Stanley Behrens Fellows in Medicine award and the UCI Graduate Dean’s Dissertation Fellowship to K.T.E., the National Institute of Health award number NIHT32EB009418 and the UCI Public Impact Fellowship to K.B., the National Institutes of Health award T32NS121727-01 to A.L., the National Institute of Neurological Disorders, Stroke/National Institutes of Health under award number T32NS082174 to M.A.C., the Feodor-Lynen fellowship from the Alexander-von-Humboldt Stiftung to J.I.-R., the National Institutes of Health award T32GM008620 to T.P.M., the Canadian Institutes of Health Research Postdoctoral Fellowship and the NIH/NCI K99/R00 Transition to Independence Award (1K99CA267160-01) to D.M., and the National Institutes of Health T32GM136624 and F31CA281331 to T.L. The authors wish to acknowledge the support of the Chao Family Comprehensive Cancer Center Experimental Tissue Shared Resource, supported by the National Cancer Institute of the National Institutes of Health under award number P30CA062203. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. Experiments using the GFP-expressing iPSC line AICS-0036 were made possible through the Allen Cell Collection, available from Coriell Institute for Medical Research. The authors also sincerely thank O. Shivashankar for sharing their advice and expertise, M. Pein for reviewing the manuscript and for her generation of GFP- and luciferase-labelled EO771 cells, D. Tifrea for obtaining human specimens, J. Atwood for assistance and training in flow cytometry and cell sorting, I. Smith for his generous gift of MDA-231-BR2 cells, and I. Chang and the UCI Research Cyberinfrastructure Center for maintenance and development of computational resources. This research is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conceptualization, K.T.E., K.B., A.L., M.A.C., J.I.-R., K.N.G., S.A.V., M.B.-J. and D.A.L.; methodology, K.T.E., K.B., A.L. and M.A.C.; investigation, K.T.E., K.B., A.L., M.A.C., J.I.-R., T.P.M., Q.H.N., D.M., T.L., G.A.H., D.O. and A.K.O.; resources, D.A.L., M.B.-J., C.P. and R.A.E.; writing—original draft, K.T.E., K.B., A.L. and D.A.L.; writing—review and editing, D.A.L., C.P. and M.B.-J.; project administration, D.A.L.; funding acquisition, D.A.L. and M.B.-J.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analysis of TAMs and astrocytes in BCBM.
a Whole mount brightfield and fluorescent microscopy images of metastatic brains (Met1-3) used to generate the scRNA-seq dataset described in Fig. 1d. Metastatic lesions are GFP+ (green). Results are representative of a single experiment. Scale bar = 50 mm. b IF staining shows IBA1+ cells (red) in normal human brain and three resected patient BCBM tumors. Insets show cell morphology, exhibiting evenly spaced, ramified microglia in normal human brain contrasting heavily infiltrated ameboid microglia in BCBM patients. Results are representative of a single experiment. Scale bar = 50 μm. c Representative FACS plots show gating for single, live (Sytox negative) myeloid cells (CD45+CD11b+), astrocytes (CD45−ASCA2+) and 231BR cells (CD45−GFP+) isolated for scRNA-seq. d Identification of mouse and human cells by the frequency of reads that align to the mm10 mouse genome. Cutoffs used to identify mouse cells (>0.875 aligned, n = 51,418 cells), human cells (<0.05 aligned, n = 7336 cells) and doublets (0.05–0.875 aligned, n = 913 cells) are shown. e Violin plots show cell distributions for key quality control metrics pre- and post- filtering and removal of poor quality cells. Cells were removed that displayed <500 or >2000 genes (nFeature_RNA), or >10% of genes mapped to the mitochondrial genome (percent mito genes). f Bar chart shows the frequency of cells contributed by each mouse that localize to each cell type in Fig. 1f. g tSNE plot shows astrocytes colored by control or metastatic condition. h Volcano plot shows genes differentially expressed (n = 6,542) between astrocytes from control and metastatic brains determined by Wilcoxon rank sum test, (p < 0.01). See Supplementary Table 2 for full list. Select genes with an absolute value average natural logFC >0.35 are colored and labeled. The y-axis represents the -log10 of Bonferroni corrected P values, and the x-axis represents average natural logFC between conditions.
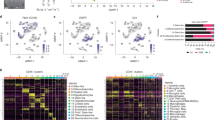
Extended Data Fig. 2 Identification of myeloid cell types and subclustering analysis of proinflammatory microglia.
a tSNE plot shows myeloid cells (n = 15,288) colored and labeled by cell type. mDC = mature dendritic cell. Mono.Macro = monocytes and macrophages. b Dot plot shows top marker genes for each cell type ranked by average natural logFC. Dot size represents the percentage of cells that express the gene, and dot greyscale represents the average expression level. See Supplementary Table 4. c Bar chart shows the frequency of cells contributed by each mouse that localize to each cell type in b. d Feature plots show myeloid cells colored by canonical cell type marker genes or features. Stressed cells were identified by increased expression of mitochondrial genome (percent.mito) genes, and decreased number of genes detected (nFeature_RNA). e Bayesian information criterion (BIC) for microglia topic models from Fig. 2d with the listed number of topics (K), each fit to an error tol = 10. f Bar plot shows the relative enrichment of each topic in control and metastatic animals from Fig. 2d. The relative enrichment was determined by subtracting the average topic assignment for the control mice from the average topic assignment across all cells in each mouse. Highlighted topics show four core topics where all three metastatic mice have a higher relative enrichment than all three control mice (that is min(Metastatic) > max(Control)).
Extended Data Fig. 3 Pro-inflammatory marker expression in microglia from BCBM models.
a Gating strategy for identification of microglia. Dot plots (top) show gating for single, live (zombie negative) CD45loCD11b+Ly6C− microglia. Histogram plots (bottom) show subsequent gating for CD74, BST2, and MHC-II in microglia. b Flow cytometry analysis of CD74, BST2 and MHC-II in microglia harvested 14 days post intracardiac injection of 4T1-GFP (100,000) cells into BALB/c animals. Bar graph shows the percent of microglia that express each marker in control (n = 7) and metastatic (n = 7) brains. P values were generated by an unpaired two-sided student’s t-test, and error bars indicate mean +/− standard deviation. c Quantification of microglia in tumor and distal regions of mice bearing EO771-GFP tumors. Representative images (left panels) show microglia localization relative to other cell types using a machine learning classifier (see Methods). Pie graphs (right panels) show the proportion of microglia and other cell types in each region. Frequencies are as follows: other non-microglia cells (TMEM119−CD74−MHC-II−ISG15−), distal = 0.94, tumor = 0.22; microglia (TMEM119+), distal = 0.05, tumor = 0.22; tumor cells (ISG15+TMEM119−), distal < 0.01 tumor = 0.14; other immune cells (TMEM119−CD74+MHC-II+), distal < 0.01, tumor = 0.41. Scale bar= 100um.
Extended Data Fig. 4 Quantification of tumor size in FIRE-WT and FIRE-KO animals.
a IVIS images show EO771 luciferase luminescence signal change over time in FIRE-WT and FIRE-KO animals. Representative animals that displayed continuous signal increase (tumor growth, solid line) vs. signal decrease (tumor rejection, dashed line) are shown. Pseudocoloring of luminescence shows quantification of radiance (p/sec/cm2/sr). b Line graphs show quantification of luminescence signal change over time in all FIRE-WT and FIRE-KO animals. Solid lines indicate animals that demonstrated tumor growth and dashed lines indicate those that showed tumor rejection. Growth was defined by signal increase over time, and rejection was defined as either baseline signal (<106) or >5-fold decrease in signal relative to maximum. c Serial dilution analysis of EO771 cell engraftment in FIRE-WT and FIRE-KO animals. 10–200 × 104 EO771 cells were transplanted intracranially into each mouse strain. Ex vivo whole brain luminescence images show signal from tumor cells in each tissue at day 14. Fractions denote the number of grafts that produced macroscopic tumors in each condition. d Dot plots quantify luminescent signal (total flux) from each tissue shown in Extended Data Fig. 4c at day 14. P values were generated by an unpaired two-sided student’s t-test.
Extended Data Fig. 5 Analysis of NK, T, and monocyte responses to BCBM in FIRE-WT and FIRE-KO animals.
a Quantification of tumor burden in FIRE-WT and FIRE-KO animals (n = 8/group). Mice were injected with EO771 GFP-Luc cells as described in Fig. 5a and tumors were harvested and analyzed by IVIS on day 7. Images (left panels) show pseudocoloring of radiance (p/sec/cm2/sr), and bar graph shows quantification of total flux (p/s). P value shown is the result of a student’s unpaired two sided t-test. Error bars represent mean +/− standard deviation. b Box plots show frequency of T cell subsets from Fig. 5c (n = 7 FIRE-WT, n = 8 FIRE-KO). Frequencies shown are out of all T cells. Bounds of box and whiskers are indicative of the first through fourth interquartile range. P value shown is the result of a student’s un-paired two sided t-test. c Analysis of T cell activation in tumor bearing FIRE-WT and FIRE-KO brain tissues by flow cytometry. CD44 and CD62L expressions were measured in CD4 and CD8 T cells to delineate T effector (Teff), T central memory (TCM) and naive T cell subsets. Representative FACS plots (top panels) show gating for each subset after gating for single, live (Sytox negative) cells. Bar graphs (bottom panels) show quantification of T cell counts for each group. Error bars represent mean +/− standard deviation. Pairwise comparisons of counts between groups were not significant. d Quantification of monocytes in tumor bearing FIRE-WT and FIRE-KO brain tissues by flow cytometry. CD11b+Ly6C+ monocytes were identified following gating for CD3−NK1.1−, single, live (Sytox negative) cells. Top panels show representative FACS plots, and bottom panels show quantification of cell counts. Error bars represent mean +/− standard deviation. Pairwise comparisons of counts between groups were not significant. e Linear regression model of CD8+ T cell and Treg quantification from Extended Data Fig. 5b. R-squared and P values determined by simple linear regression function.
Extended Data Fig. 6 Analysis of tumor burden and the immune response in T cell deficient mice.
a Gating scheme for analysis of T cells in brain tissue harvested from vehicle treated, FTY720 treated and RAG1-KO mice. FACS plots show gating of TCRb+ T cells from single, live (sytox negative) CD45hiCD11b− cells from Extended Data Fig. 6b. b Gating scheme for analysis of microglia and monocytes in brain tissue from vehicle, FTY720 and RAG1-KO animals. FACS plots (top panels) show gating for CD45hi-int and CD11b+ cell populations, followed by gating for CD45hiLy6C+ monocytes and CD45intLy6C− microglia (bottom panels). c Bar graphs show the percentage of microglia and monocytes out of total live, single cells in brains harvested from vehicle (veh, n = 6), FTY720 (FTY, n = 6) and RAG1-KO (RAG1, n = 4) animals. Pairwise comparisons of percentages between groups were not significant. Error bars represent standard deviation. d Quantification of EO771 tumor burden at endpoint on day 12 by IVIS. Pseudocolor shows radiance (p/sec/cm2/sr) in each whole brain.
Extended Data Fig. 7 Analysis of BCBM immune repertoire in T cell deficient and replete mice.
a Dot plot shows top marker genes for each cell type in total CD45hi-int sorted cells, ranked by the average log2 fold-change and determined by the Wilcoxon rank sum test. Dot size represents the percentage of cells that express each gene, and dot greyscale represents the average expression level. Macro.DCs = macrophages and dendritic cells, mDCs = mature dendritic cells, pvMacro = perivascular macrophages. Bar graphs illustrate relative contribution of each cluster to total leukocytes, separated by mouse strain and timepoint. b UMAPs show T cells (n = 1949 cells) from C57BL/6 mice at day 4 and 10, colored by cluster (left) and timepoint (right). c Bar graph illustrates the distribution of T cell clusters in each animal (n = 6) separated by timepoint. d Dot plot shows expression of top marker genes for each T cell cluster from Extended Data Fig. 7b. CD4.eff = CD4+ effector T cell, CD8.eff = CD8+ effector T cell, Lt.stg.eff = late stage effector T cell, γδ = gamma delta T cell. e Dot plot shows expression of top marker genes for each microglia cell cluster from Fig. 7b.
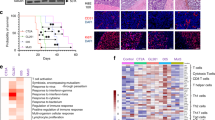
Extended Data Fig. 8 Analysis of humanized mouse model of BCBM and patient BCBM data.
a Whole mount brightfield and fluorescence microscopy images show brains from MITRG mice transplanted with GFP-labeled iHPSC cells and mCherry-labeled 231BR cells from Fig. 8a. b Identification of mouse and human cells by the frequency of reads that align to the mm10 mouse genome. Cutoffs used to identify mouse cells (>0.95 aligned, n = 641 cells), human cells (<0.1 aligned, n = 25,287 cells) and doublets (0.1–0.95 aligned, n = 387 cells) are shown. c Violin plots show cell distributions for key quality control metrics pre- and post- filtering and removal of poor quality cells. Cells were removed that displayed >20% of genes mapped to the mitochondrial genome (percent mito genes). d tSNE plot shows human cells, colored by cluster and labeled by cell type. pvMacro = perivascular macrophages, Cycling = cycling myeloid cells. See Supplementary Table 6 for full gene list. e Dot plot shows top marker genes for each cell type determined by the Wilcoxon rank sum test and ranked by average natural logFC. Dot size represents the percentage of cells that express the gene, and dot greyscale represents the average expression level. See Supplementary Table 6 for full gene list. pvMacro = perivascular macrophages. f Bar chart shows the frequency of cells contributed by each mouse to the cell types shown in e. g tSNE plots colored to show the expression of BST2, CD74 and CCL3. h tSNE plot shows the distribution in human microglia of the three core topics identified in mouse microglia in response to BCBM. Gene scores for each topic from Fig. 2e were generated using the AddModuleScore function in Seurat (See Methods). Each topic is indicated by color, where only cells with a topic score > 0.25 are colored. Contrast gray scale indicates topic weights. Scaling was performed by dividing all topic scores by the maximum topic score across the dataset. i Bar plot shows the relative enrichment of each topic score in human microglia. j Kaplan-Meier plots show overall survival probability stratified by MHC-II, CSF1, and BST2 expression in bulk RNA-seq data from human patient BCBM tumors (Varešlija et al.50).
Supplementary information
Supplementary Information
FACS gating strategies.
Supplementary Table
Supplementary Tables 1–6.
Source data
Source Data Fig. 1
Quantification of IF images.
Source Data Fig. 3
Quantification of flow cytometry data and cytokine array.
Source Data Fig. 4
Quantification of tumour burden in FIRE mice.
Source Data Fig. 5
Quantification of flow cytometry data.
Source Data Fig. 6
Quantification of flow cytometry and IVIS data.
Source Data Extended Data Fig./Table 3
Quantification of flow cytometry data.
Source Data Extended Data Fig./Table 4
Quantification of IVIS data.
Source Data Extended Data Fig./Table 5
Quantification of flow cytometry and IVIS data.
Source Data Extended Data Fig./Table 6
Quantification of flow cytometry data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Evans, K.T., Blake, K., Longworth, A. et al. Microglia promote anti-tumour immunity and suppress breast cancer brain metastasis. Nat Cell Biol 25, 1848–1859 (2023). https://doi.org/10.1038/s41556-023-01273-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-023-01273-y