Abstract
Memory consolidation after learning involves spontaneous, brain-wide network reorganization during rest and sleep, but how this is achieved is still poorly understood. Current theory suggests that the hippocampus is pivotal for this reshaping of connectivity. Using fMRI in male mice, we identify that a different set of spontaneous networks and their hubs are instrumental in consolidating memory during post-learning rest. We found that two types of spatial memory training invoke distinct functional connections, but that a network of the sensory cortex and subcortical areas is common for both tasks. Furthermore, learning increased brain-wide network integration, with the prefrontal, striatal and thalamic areas being influential for this network-level reconfiguration. Chemogenetic suppression of each hub identified after learning resulted in retrograde amnesia, confirming the behavioral significance. These results demonstrate the causal and functional roles of resting-state network hubs in memory consolidation and suggest that a distributed network beyond the hippocampus subserves this process.
Similar content being viewed by others
Introduction
The formation of enduring memory in the brain is a distributed and dynamic process, the mechanism of which is not fully understood. Current theory of systems memory consolidation suggests that the hippocampus mediates the encoding of information from segregated sensory, motor, or motivation brain networks that are engaged during learning, gradually reshaping their connectivity to form long-term memory1,2. This is facilitated by the reactivation of learning-associated neuronal populations (replay) and the coordinated interaction of the hippocampal-neocortical network during post-encoding periods of quiet wakefulness (resting state) and sleep3,4,5,6. This process is highly dynamic, with the hippocampus initially mediating cortical plasticity, but after which the neocortical network becomes more active1,7,8. Apart from the hippocampus, where, when and how other regions are involved in facilitating this system-wide reconfiguration are still unclear. Whole-brain functional imaging during task performance has revealed broad engagement of not only neocortical but also subcortical areas when encoding or recalling memory in humans9 and rodents10,11. However, the regions involved in consolidating memory in the ill-defined, “offline” period are difficult to pinpoint without aligning them to activities associated with replay12.
A major advance over the last decade has been the identification of the brain-wide network involved in spontaneous activity during task-free conditions13. Functional connectivity (FC), measured as the correlation between regional activities at resting state, forms large-scale networks of functionally associated areas that indicate an intrinsic organization of the brain14,15. The disruption of these resting-state networks (RSNs) in association with cognitive impairment in aging and disease provides evidence for their involvement in the etiology and progression of these conditions16,17,18. Advances in network neuroscience have further revealed that the topology of RSNs changes with performance or symptom, suggesting the behavioral relevance of their organization and dynamics19,20. Importantly, learning can induce ongoing remodeling of the RSNs over time in humans21,22 and rodents23,24. Increased association between hippocampus-neocortical FC and performance over repeated training25,26, reconfiguration after sleep27,28, and reactivation of the learning-related activity pattern29,30 have suggested that post-encoding RSNs reflect systems memory consolidation.
However, a fundamental question that remains is whether the spontaneous network changes are causative of the behavior- or disease-associated states with which they correlate31. Due to the observational nature of the experimental designs, unconstrained imaging environments, and correlation-based FC measures, it remains possible that the observed RSN changes are epiphenomena which are driven by alternative neural, physiological or pathological factors32,33,34,35,36. Furthermore, if they are causative, the activity and areas that drive such large-scale network remodeling remain unresolved. Analytical methods can allow the inference of causality from the RSNs (for review, see ref. 37). Nonetheless, they only estimate inter-dependency between regional activities within a network, instead of the causality to behavior. Critically, whether a network or its hub is causally required for cognition (e.g., episodic memory), such that its dysfunction leads to a disability (e.g., amnesia) whereas its facilitation improves performance, has not been directly tested experimentally with prospective interventions.
In this study, we reveal the brain networks that are instrumental in consolidating memory during post-encoding rest by identifying and functionally manipulating RSN hubs. We examined two hypotheses for defining causal hubs of behavior, one based on a common network and the other on network integration. Certain elements of the hippocampal-neocortical network, particularly between the hippocampal formation (HPF, including the hippocampus, subiculum and entorhinal cortex), retrosplenial cortex (RSC) and medial prefrontal cortex (mPFC), have been identified in different spatial or contextual learning paradigms8,38,39,40,41. The storage of various forms of the spatial memory trace (engram) in these areas8 indicates that a common, task-invariant network may be involved in the systems consolidation1,42, although the full extent of this common network is still unclear. In addition, consolidation incorporates new information from functionally segregated areas, such that this integration could be manifested at the network level. Indeed, topological features of network integration, such as global efficiency43, or segregation, such as modularity44, have been shown to correlate with performance during or after learning. Altered network integration is also found after cognitively demanding tasks45,46 or overnight consolidation27. These findings indicate that network integration is an essential feature in learning and memory. Thus, influential hubs for network integration (“integrator” hubs) may have a causal role in memory formation.
To test whether common network hubs (behaviorally influential brain regions consistently invoked by different learning paradigms) or integrator hubs (behaviorally influential brain regions contribute to network integration) are causally involved in memory consolidation, we trained mice in two versions (early and late retrieval) of active place avoidance (APA), a spatial memory task, and subsequently acquired resting-state functional magnetic resonance imaging (rsfMRI) data to characterize behavior-induced RSN changes during post-learning period (Fig. 1a). Previous studies indicated that different retrieval intervals may form memory via different mechanisms, with elevated network activity and transcription factors within short (1–5 h) retrieval intervals facilitating the integration of information, whereas reactivation of stored memory is involved after a longer interval47,48. Whether the same circuitry is engaged in these processes is unclear. Furthermore, to test whether the RSN during consolidation transitions from hippocampal- to neocortical-dependent, we conducted rsfMRI at 1 day and 1 week after learning to track the dynamics of network reorganization. We also developed methods for detecting common or integrator hubs from post-encoding RSNs, with our results revealing that the sensory cortices were commonly engaged following both tasks, and the prefrontal cortex and the striatal and thalamic nuclei were important for network integration. We then validated the behavioral impact of the identified hubs by silencing each hub individually during the consolidation period using inhibitory Designer Receptors Exclusively Activated by Designer Drugs (DREADDs)49.
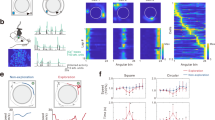
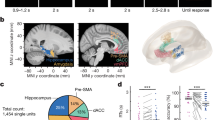
a The schematic diagram of the APA-rsfMRI experiment. b The left diagram shows the setup of the APA task. Four distinct pictures were hung on the surrounding walls as visual cues. The orange arrow indicates the direction of rotation. The 60o sector in red shows the location of the invisible “shock zone”. The two plots on the right show the progressive decrease in the number of shocks over the trials (two-way ANOVA, F5, 80 = 14.22, p < 0.0001), which is comparable between the 1-Day (N = 10) and 5-Day APA (N = 8) groups (F1, 16 = 0.55, p = 0.47 for groups). Similar trends can be seen in the time to first entrance of the shock zone (two-way ANOVA, F5, 80 = 6.63, p < 0.0001 for training trials, F1, 16 = 1.27, p = 0.28 for groups). Post hoc comparisons were performed between the last training trial (T5) and other training trials (T1–T4) or probe test (PT) with Dunnett’s multiple comparison test. The number of shocks for 1-Day APA: T1, p = 7.1 × 10−6; T2, p = 7.1 × 10−6; T3, p = 0.00016; 5-Day APA: T1, p = 1.9 × 10−5; T2, p = 0.00051. The time to first entrance for 1-Day APA: T1, p = 0.016; T2, p = 0.017; T3, p = 0.018; 5-Day APA: T1, p = 0.034. Data are represented as mean ± SEM. *p < 0.05; ***p < 0.001; ****p < 0.0001. c The seed-based correlation analysis used to create the FC matrix of each animal. d Changed functional connections in the 1-Day and 5-Day APA, compared to their corresponding controls, on post-training day 1 and post-training day 8 (two-sample t-test, two-tailed, p < 0.05, FDR corrected; see Supplementary Table S4 for N of each group). The red connections represent APA > control while the blue connections represent APA <control. The line thickness indicates the t value. e Two functional connections from the 1-Day APA post-training day 8 correlated with the memory retention probe test (N = 8; Pearson correlation, two-tailed). See Supplementary Table S1 for the abbreviations of brain regions. Number of animals is from biologically independent mice. Source data are provided as a Source Data file. Significant connections were overlaid on the 3D-rendered brain atlas using BrainNet Viewer for (d). (https://www.nitrc.org/projects/bnv/), Copyright © 2007 Free Software Foundation, Inc.
Results
Similar behavior leads to distinct post-encoding RSNs
It has been shown that the HPF, mPFC and RSC are typically involved in the consolidation of spatial or contextual memory. To investigate whether similar spatial learning invokes a common network in post-encoding rest, we conducted two APA tasks with the same training trials but different inter-trial intervals: one hour (1-Day APA, as the learning was completed in one day) versus one day (5-Day APA; Fig. 1a). APA allows spatial navigation and memory to be assessed in mice with less stress than that associated with the water maze by training them to avoid a shock zone based on spatial cues over repeated trials (Fig. 1b)50,51. After learning, two sessions of rsfMRI were performed on post-training days 1 and 8 to examine the plasticity of the RSNs. One day after the second rsfMRI scan, a probe test was performed to assess memory retention. The number of shocks (Nshock) that the animals received and the time to first entrance into the shock zone (Tenter) were used to measure their behavioral performance. Nshock gradually decreased and Tenter increased during learning; this, together with the similar values obtained during the probe tests (Fig. 1b; Supplementary Results), demonstrated that the mice could remember both APA tasks equally well after 9 days, and the formation of long-lasting memory.
We distinguished post-encoding RSNs by comparing the FC between 230 highly parcellated brain regions in a brain template (Fig. 1c; Supplementary Table S1) of the APA groups versus their own controls. In the control group, animals were exposed to the APA training procedures without any foot shock being delivered as we found that random shock elicited a strong stress response. Despite comparable behaviors during learning and retrieval of both APA tests, we found distinct post-encoding RSNs. On post-training day 1 (Fig. 1d; two-sample t-tests, p < 0.05, false discovery rate [FDR]-corrected), the 1-Day APA increased the sparse FC in the left hemisphere between the entorhinal cortex and pontine nucleus, which is a pivotal relay and transformer for motor signal between the cerebellum and cerebral cortex52; and between the dorsal anterior cingulate cortex ([A24a], part of the mPFC) and olfactory tubercle, which is involved in sensory-guided reward/motivation behaviors53; but decreased the FC between the somatosensory and prefrontal cortices and between the HPF and pons in the right hemisphere. In contrast, the 5-Day APA increased the FC mostly in the right hemisphere, including the HPF, prefrontal cortex and sensory areas. This highly lateralized FC (12 out of 17 connections) is consistent with studies reporting that the right hemisphere is predominant in memory processing54,55,56. One week after APA training, the network was reorganized. In the 1-Day APA group, even more inter-hemispheric FC was found, with increased FC being observed among the somatosensory cortex, lateral accumbens shell (LAcbSh, an area involved in feeding, reward and motivated behavior57) and ventral entorhinal cortex, whereas the FC between the lateral orbital cortex (LO, a prefrontal region involved in decision making and the acquisition of hippocampus-dependent memories58,59,60), somatosensory cortex, thalamus, and pons decreased. In the 5-Day APA group, only sparse FC between the HPF, prefrontal cortex and thalamus in the right hemisphere was found. Comparable post-encoding plasticity could also be identified using independent component analysis (ICA), revealing distributed network in the sensory cortex, mPFC, hippocampus, basal ganglia and thalamus (Supplementary Results and Fig. S1). These results indicated that post-encoding RSNs are task- and time-dependent and involve distant neocortical and subcortical areas, similar to the findings of previous studies using the Morris water maze23,24. The overall connectivity with the HPF and mPFC is consistent with their critical roles in memory consolidation, although the specific subregions involved differed between tasks. A common network between the two APA tasks could be obscured with such detailed parcellation. Alternatively, this could be due to the much stricter false positive rate when calculating the overlap between two FDR-corrected connectivity matrices.
It is generally expected that behaviorally relevant connections predict performance. Many studies have found an association between memory performance and FC during encoding or retrieval61,62,63, yet little is known about the relationship with post-encoding FC. To test whether post-encoding FC is associated with memory retention, we calculated Pearson’s correlation between FC strength and Nshock or Tenter in the probe trial (Fig. 1e). Only two connections on post-training day 8 in the 1-Day APA group correlated with behavior: between the left posterolateral cortical amygdala (PLCo) and the LAcbSh (r = 0.87, p = 0.0045) and between the right primary somatosensory cortex jaw region and the inferior colliculus (r = 0.82, p = 0.012). Although several connections were enhanced in the 5-Day APA, no correlation with behavior was found. This indicates that the most significant FC may not be influential for behavior.
Locating common network hubs that correlate with behavior
We predicted that behavior-correlated RSNs commonly induced by both kinds of APA tasks are influential for memory consolidation. As combining two FDR-corrected thresholds reduces the true positive rate, we first lowered the threshold for the two-sample t-tests to an uncorrected p < 0.05 to discover common RSNs induced by both tasks, (Fig. 2a). Despite being similar in their task designs, only a small fraction of the FC was found in both APA tasks, with 3.56% (post-training day 1) and 2.85% (post-training day 8) of the connections overlapping. To identify the causal hub, we selected the FC that correlated with behavioral performance in the probe test. Using a permutation test, imposing two uncorrected network thresholds and behavioral correlation with Nshock (threshold at p < 0.05) together resulted in an equivalent family-wise error of p < 0.05 for common connections on both post-training day 1 and 8 (Fig. 2b, c). When using Tenter as a behavioral index, the family-wise false positive rate of common connections was p = 0.019 on post-training day 1 but was not significant on post-training day 8 (Supplementary Fig. S2). Here, we chose to use Nshock as the primary behavioral index.
a Procedures for identifying target hubs based on common networks that correlate with behaviors in the probe test (PT). The difference FC matrix between the APA and control (Ctl) groups was calculated by two-sample t-test. The FC on (b) post-training day 1 (N = 17) and (c) post-training day 8 (N = 16) that correlated with Nshock (Pearson correlation, p < 0.05, two-tailed, uncorrected) within the common networks (p < 0.05, uncorrected) of the 1-Day and 5-Day APA. The red lines show the functional connections that positively correlate with Nshock while the blue lines show those that negatively correlate with Nshock. The line width indicates the absolute r value. The cumulative distribution of the 5000 permutation tests is shown on the right. The red bars represent the cumulative probability before reaching the real number of edges (9 connections on post-training day 1 and 5 connections on post-training day 8). Behavioral correlation (Pearson correlation, two-tailed) of the functional connections with the target hubs on (d) post-training day 1 (N = 17) and (e) post-training day 8 (N = 16). See Supplementary Table S1 for the abbreviations of brain regions. Number of animals is from biologically independent mice. Source data are provided as a Source Data file. Significant connections were overlaid on the 3D-rendered brain atlas using BrainNet Viewer for (b) and (c) (https://www.nitrc.org/projects/bnv/), Copyright © 2007 Free Software Foundation, Inc.
We found behavior-correlated common networks composed of the hippocampus, mPFC and thalamus in the left hemisphere, and the connection between the primary somatosensory and primary visual (V1) cortex in the right hemisphere on post-training day 1 (Fig. 2b, Supplementary Table S2). Excluding the HPF and mPFC, which are known to be engaged in memory consolidation, and subcortical areas, the FC of the left primary somatosensory cortex barrel field (S1BFL) had the highest behavioral correlation (CA3-OrL−S1BFL, r = −0.68, Cohen’s d = 0.93), followed by right V1 (V1R−S1R, r = 0.57, Cohen’s d = −0.89; Fig. 2d and Supplementary Fig. S3). On post-training day 8 (Fig. 2c, Supplementary Table S2), the FC with the mPFC was gone. Instead, we observed FC with the reticular nucleus (Rt), which drives the neural oscillations important for memory consolidation during sleep5, and the RSC. The engagement of the mPFC on day 1 with silencing one week later is consistent with the temporal dynamic of the engram in this area8. Excluding the entorhinal and retrosplenial cortices, the right secondary somatosensory cortex (S2R) had the highest behavioral correlation (CA3-OrL - S2R, r = −0.80, Cohen’s d = −1.26; Fig. 2e and Supplementary Fig. S3). Overall, we found that expanded behavior-correlated common networks beyond the HPF, mPFC, and RSC were engaged at different times after learning. Here, we chose the S1BFL, V1R, and S2R which had large effect size as the targets for validation.
Learning alters network integration
Based on the importance of network integration in learning and memory27,43,45,46,63, we predicted that post-encoding RSNs would be more integrated after spatial learning. To investigate this, we applied graph theory analysis, which simplifies the brain network as nodes (brain regions) and edges (FC strengths). To evaluate the network integration and segregation, we used several graph measures: the global efficiency, modularity, transitivity, size of the giant component and the small-world topology. Global efficiency measures the shortest path length which reflects integration. Modularity, which calculates the size and number of network component and intra-component connections, is a measure of segregation. Transitivity measures how tightly that nodes are tightly connected within a cluster thus reflects segregation. Giant component is the largest cluster of interconnected nodes which represents network integration. Small-world topology is a key feature of the brain network presenting local segregation and long-range integration64. We evaluated the small-world features using the normalized characteristic path length, lambda; normalized clustering coefficient, gamma; and small-worldness, sigma.
With an increased t-score threshold from 2.0 to 3.8 (uncorrected), the connectivity matrices after two-sample t-tests became more fragmented, resulting in a reduced giant component, global efficiency and transitivity but increased modularity (Fig. 3 and Supplementary Fig. S4). To test the overall difference, we calculated the area under the curve (AUC)65 and compared it to the distributions of 5000 random networks. Based on the null distribution, both APA training protocols, except the 5-Day APA on post-training day 8, significantly increased the size of the giant component (Fig. 3a, and Supplementary Fig. S4a). The global efficiency was only increased in 1-Day APA on post-training day 8 (Fig. 3b, and Supplementary Fig. S4b). Interestingly, the modularity was significantly increased except the 5-Day APA on post-training day 8 (Fig. 3c, and Supplementary Fig. S4c). 1-Day, but not 5-Day, APA learning led to a significant decrease in transitivity, indicating that the post-encoding network is rather distributed instead of tightly connected (Fig. 3d, and Supplementary Fig. S4d). Similar trends could also be observed using an unweighted network except for 5-Day APA on post-training day 8 (Supplementary Fig. S5 and Supplementary Fig. S6). Small-world features were calculated on individual RSNs because the two-sample t-test matrices were too sparse. We found a trend towards an increase in the small-worldness, sigma, after learning due to a trend of higher local segregation (increased gamma) and long-range integration (reduced lambda) compared to the control (Supplementary Fig. S7). Interestingly, the small-worldness on post-training day 1 in 5-Day APA was significantly decreased (Supplementary Fig. S7c) due to a significantly longer path length and lower clustering, suggesting sparse segregation.
Trends of (a) giant component size, (b) global efficiency, (c) modularity and (d) transitivity of post-encoding RSNs for 1-Day (left) and 5-Day (right) APA on post-training day 1 (red) compared to random networks (black), thresholded at 2 ≤ t ≤ 3.8. The black lines show the mean ± SD of 5000 random networks generated based on the real network. The embedded bar graphs show the probability distribution of the area under curve (AUC) of the random networks. The red arrows indicate the AUC for the real network and the corresponding p value estimated based on the permutation test. PD probability distribution. Source data are provided as a Source Data file.
Such paradoxically increased network integration and segregation is also found in a recent study that reported repeated training, which automates a cognitively demanding task, can increase the integration and segregation of post-encoding RSNs46. To understand the cause of these features, we examined the key elements behind the modularity measure: the number of components and intra-component connections. We found a steady increase in the proportion of connections within components but a plateau in the number of components with increased threshold (Supplementary Fig. S8). This indicates that much stronger intra-component connections than those between components caused an increase in modularity. Together these results indicate that APA learning increases network integration and segregation by forming loose-linked, larger and more network components while also strengthening the connectivity within components.
Optimal method for distinguishing integrator hubs
As network integration is a feature of post-encoding RSNs, pinpointing the integrator hubs would allow us to test whether this is causally required in memory consolidation. We predicted that when an integrator hub is removed (inhibited), the network integration would be greatly impeded, leading to the breakdown of the giant component. The best method for hub identification would be one which can reduce the size of the giant component with the removal of the fewest number of nodes. Centrality, which describes the importance of network communication and integration, is typically regarded as reflecting network integration. However, simulation showed that centrality is a poor measure of causal inference66.
To determine the method for hub identification, we compared four centrality measures (degree centrality, closeness centrality, betweenness centrality, eigenvector centrality), a link authority HITS (Hyperlink Induced Topic Search) score, and collective influence (CI), which searches for nodes that can quickly break down large networks in an optimal percolation model67,68. We calculated the reduction in the giant component in the post-encoding RSNs by removing high ranking/centrality nodes one by one. Figure 4b shows an example in which the normalized giant component size quickly dropped by removing nodes detected by the CI, followed by those identified by the degree centrality and betweenness centrality, whereas closeness centrality, HITS and eigenvector centrality had a slower effect. Similar trends were observed in networks from both APA groups on post-training days 1 and 8 (Supplementary Fig. S9). Comparing the AUC of the giant component changes, the CI analysis resulted in the quickest collapse of the giant component (Fig. 4c, Supplementary Fig. S10). This indicates that CI is a better method for identifying integrator hubs.
a Procedures for identifying target hubs based on network integration. The difference FC matrix between the APA and control (Ctl) groups was analyzed by CI to select top ranking nodes. Nodes with FC that correlated with behaviors in the probe test (PT) were selected. b The relative giant component size decreases by removing high-ranking CI, centrality or HITS hubs one by one from the difference FC matrix between the 5-Day APA and its control on post-training day 1 (p < 0.01, two-sample t-test, two-tailed, uncorrected). Six hub selection methods, including CI (blue), degree centrality (red), closeness centrality (green), betweenness centrality (purple), eigenvector centrality (orange), and HITS (black) were compared. The quicker the giant component shrinks, the more effective the original network breaks into smaller networks (reduced network integration). c Comparison of the area under the curve (AUC) of the giant component reduction curves (N = 4 experimental groups). The data points were derived from the difference FC matrices comparing APA and control groups for the 1-Day APA and 5-Day APA datasets on both post-training day 1 and day 8. Each data point was normalized by the corresponding AUC of CI. One-way ANOVA, F5, 15 = 67.45, p < 0.0001. Bonferroni’s multiple comparison tests: degree, p = 0.028; closeness, p = 5.2 × 10−5; betweenness, p = 0.018; eigenvector, p = 7.3 × 10−10; HITS, p = 2.4 × 10−7. Data are represented as mean ± SEM. *p < 0.05; ****p < 0.0001. Source data are provided as a Source Data file.
Identification of integrator hubs that correlate with behavior
We next applied CI analysis to post-encoding RSNs to identify nodes with FC that correlates with memory retention (Fig. 4a). As shown in Fig. 1d, using the FDR-corrected threshold makes the network very sparse without a giant component, thereby excluding the use of CI analysis for hub identification67,69. Here, we used multiple uncorrected thresholds, p < 0.05, 0.01 and 0.005, to determine the averaged ranking of a node in breaking down the RSNs after 1-Day APA training. On post-training day 1, the 10 top-ranking nodes were mostly subcortical, including regions in the basal ganglia, midbrain and brainstem, with cortical areas in the HPF (subiculum and entorhinal cortex) and S2 being ranked lower (Table 1). On post-training day 8, more cortical areas (parietal association, sensory and prefrontal cortices) rose to the top ranking compared to subcortical areas. Compared to the hubs identified by the HITS score, 8 out of the 20 hubs were the same as those found by CI analysis albeit with a different ranking (supplementary Table S3). To identify candidate nodes that are influential on behavior, we selected CI nodes with nodal FC correlated with memory retention in the probe test (Table 2). We found that the right caudate putamen had a connection with the highest correlation with Nshock (CA1-LmolR − CPuR, r = −0.79, p = 0.0063, Cohen’s d = 1.25), and the left LAchSh had a connection with the highest correlation (LAcbShL − RtR, r = 0.95, p = 3.7 × 10−5, Cohen’s d = 1.21) with Tenter on post-training day 1. On post-training day 8, nodal FC with high behavioral correlation included the left ventromedial thalamic nucleus (VML − A30L, r = 0.91, p = 0.0015, Cohen’s d = −1.37), left primary somatosensory cortex forelimb region (S1FLL − MeAR, r = 0.89, p = 0.0033, Cohen’s d = 1.36), right LO (VLL − LOR, r = 0.87, p = 0.0045, Cohen’s d = −1.25), right periaqueductal gray (EAL − PAGR, r = −0.84, p = 0.0088, Cohen’s d = −1.43) and right primary somatosensory cortex trunk region (PoDGR − S1TrR, r = −0.84, p = 0.0095, Cohen’s d = −1.35; Supplementary Fig. S3). Some of these nodes had high CI ranking (top 3) whereas some had low ranking (bottom 3). Based on the averaged CI rank, we chose the LAchShL, LOR, VML, and S1TrR as the high, middle and low ranking hubs for validation.
Validation of causal hubs by DREADDs inhibition
To verify the causal role of selected hubs in memory consolidation, we injected AAV2/1-pSyn-hM4D(Gi)-T2A-mScarlet to transfect inhibitory DREADDs in all neurons in each area individually (Fig. 5a). One month after the surgery, animals went through the 1-Day APA training. Immediately after finishing the five training trials, animals were administered clozapine N-oxide (CNO) by intraperitoneal injection, followed by drinking water containing CNO to maintain inhibition of the targeted hubs for 7 days until one day before the probe test, to allow clearance of the CNO70,71. In addition to a naïve group to control for the effect of CNO, we chose one cortical area, the right frontal association cortex (FrAR), and one subcortical area, the right ventral posteriomedial thalamus (VPMR), which did not present in our analyses, as negative controls. Figure 5b, e, g illustrates good viral expression in the targeted areas in both the experimental and control groups, although we did notice that there was some viral expression in nearby brain areas, such as S2R and LAcbShL.
a Schematic diagram of target validation using DREADDs inhibition. Representative DREADDs expression (red) in the target areas selected from (b) the negative control, (e) common networks, and (g) network integration. In each subgraph, the left image shows the injection location of AAV-pSyn-hM4D(Gi)-T2A-mScarlet, and the right image shows the fluorescence imaging together with the DAPI staining (blue). c The number of shocks during the learning trials (T1–T5) show a progressive decrease and insignificant change during the probe test (PT) after inhibition of the negative controls, VPMR (F5, 20 = 10.80, p < 0.0001; N = 5) and FrAR (F5, 20 = 10.65, p < 0.0001; N = 5). d The behavior was comparable to that of the CNO-control (CTL) group (F5, 40 = 7.41, p < 0.0001; N = 9). f Similar trends during learning can be seen with DREADDs in the common hubs V1R (F5, 45 = 6.95, p < 0.0001; N = 10), S2R (F5, 35 = 6.02, p = 0.0004; N = 8) and S1BFL (F5, 50 = 10.67, p < 0.0001; N = 11), or in (h) integrator hubs LAchShL (F5, 30 = 5.31, p = 0.0031; N = 7), LOR (F5, 35 = 6.13, p = 0.0004; N = 8), VML (F5, 45 = 7.16, p < 0.0001; N = 10) and S1TrR (F5, 30 = 6.40, p = 0.0004; N = 7). After hub inhibition, impaired memory recall was seen, except S1TrR. i Compared with the CTL (two-sample t-test, one tailed, uncorrected), an increased number of shocks (PT−T5) after inhibition of S1BFL (Cohen’s d = 0.76, p = 0.046), V1R (Cohen’s d = 0.78, p = 0.045), S2R (Cohen’s d = 0.94, p = 0.025), LAcbShL (Cohen’s d = 1.04, p = 0.017) or LOR (Cohen’s d = 1.10, p = 0.0090) was found. Data are represented as mean ± SEM. Unless noted, statistical test is one-way ANOVA with post hoc Dunnett’s multiple comparison test with respect to T5. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001. See Supplementary Table S1 for the abbreviations of brain regions. Number of animals is from biological independent mice. Source data are provided as a Source Data file. The brain outlines were created with BioRender.com.
Animals successfully learned the 1-Day APA task (Fig. 5). The consistent improvement over the five training trials in the two negative control groups showed that the surgery itself did not affect spatial learning, based on comparison of the first and last trials (t = 6.06, p = 0.0038 for VPMR; t = 8.55, p = 0.0010 for FrAR; Fig. 5c). After receiving CNO for one week, the mice in the negative control (FrAR and VPMR) or in the CNO control group (Fig. 5d) showed intact memory recall in the probe test when compared to their own last training trial (T5) (t = 2.28, p = 0.085 for VPMR; t = 1.63, p = 0.18 for FrAR; t = 1.76, p = 0.12 for CNO control). In contrast, the Nshock was significantly increased after inhibiting each of the common hubs, S1BFL (t = 4.23, p = 0.0017), V1R (t = 3.76, p = 0.0045) and S2R (t = 3.57, p = 0.0091), and the high or middle ranking integrator hubs, LAcbShL (t = 3.85, p = 0.0084), LOR (t = 3.63, p = 0.0084) and VML (t = 2.49, p = 0.034), during the consolidation period (Fig. 5f, h). No difference was found after inhibiting a low-ranking node, the S1TrR (t = 1.14, p = 0.30). Compared to the CNO control, a significantly larger ΔNshock between T5 and the probe test was found when inhibiting S1BFL (Cohen’s d = 0.76, p = 0.046), V1R (Cohen’s d = 0.78, p = 0.045), S2R (Cohen’s d = 0.94, p = 0.025), LAcbShL (Cohen’s d = 1.04, p = 0.017) or LOR (Cohen’s d = 1.10, p = 0.0090), but not VML (Cohen’s d = 0.40, p = 0.20) or S1TrR (Cohen’s d = −0.045, p = 0.47) (Fig. 5i). The Tenter also exhibited similar trends in these regions except for V1R (Supplementary Fig. S11). These results demonstrate that inhibition of common hubs, or middle to high-ranking integrator hubs can impair memory consolidation.
Discussion
Defining brain regions and their functional involvement in the spontaneous, brain-wide reorganization that occurs after learning is essential for understanding the circuitry and mechanism of memory consolidation. Although spontaneous network activity presented in the RSNs has been identified for decades and proposed to play a role in learning and memory, this function has not been directly demonstrated. Here we demonstrate that, in addition to the HPF and mPFC, sensory areas are commonly involved following APA learning and that prefrontal, striatal and thalamic areas are pivotal for network integration. We confirm that inhibition of these RSN hubs after successful learning impairs memory formation. Our results demonstrate a causal link between post-encoding RSNs and memory consolidation, and reveal that a distributed network mediates this process, as well as providing effective methods for inferring causal hubs of behavior. This expands our understanding of the brain-wide network involved in memory formation. Considering the comparable organization and properties of human and rodent RSNs72,73, our validated approaches have the potential to identify targets for intervention to modulate cognition and behavior.
Network hubs are typically defined based on their importance in network topology using measures such as centrality, rich club, and HITS20. However, a high centrality node may not necessary be the most influential node74. In particular, it is unclear whether and how a brain network hub causally impacts behavior. In this study, we combined behavioral and topological features to identify two kinds of post-encoding RSN hubs that are influential on behavior: common network and integrator. We found that behaviorally defined common network hubs (shown in both APA tasks and having connections correlated with memory retention) and topologically (breakdown of giant component) and behaviorally defined integrator hubs can causally affect the behavior. From topological point of view, an integrator hub would be similar to a connector node that links two network modules20. However, a connector node may not necessarily be influential on behavior. We also found that CI analysis can effectively detect integrator hubs among the network measures tested. Identifying influential node remains a challenge in network science. Other approaches, such as k-shell decomposition74, integrated value influence75 and VIP76, would be useful for selecting candidate nodes for testing their behavioral effects.
The HPF and mPFC have been the most common targets in memory research. Apart from these areas, we verified the engagement of an extended network that commonly supports systems consolidation. We found that several subcortical areas in the thalamus and basal ganglia were invoked by both APA tasks, consistent with the highly distributed subcortical engrams reported in a recent study of contextual fear conditioning10. Despite only a few neocortical connections being found, their hubs are required in memory consolidation. We discovered involvements of sensory areas (V1, S1BF, and S2) in systems consolidation. The early visual cortex has been reported to play a role in consolidating visual working memory77; however, its involvement in long-term memory consolidation has not been demonstrated. Although the hippocampus does not directly drive primary sensory areas, replay of maze-running activity patterns during slow-wave sleep has been observed in V178, supporting its involvement. S1 has been found to engage in motor, but not spatial, memory consolidation79. Recently spatially selective activity, similar to that of the place cells in the hippocampus, was found in S1, providing a mechanism for location-body coordination80. Our result provides evidence showing the involvement of S2 in memory consolidation, likely due to its role in integrating somatosensory information involved in the foot shock. Our analysis indicated that S2 is functionally connected to the CA3 region of the hippocampus, warranting further investigation of its interaction with the HPF in spatial memory formation.
Our results demonstrate the essential roles of integrator hubs in memory formation and support the notion that network integration is a key factor in memory processes. This is consistent with a rodent study which demonstrated that inhibition of brain regions estimated from covariate c-fos activity networks led to a reduction in the giant component correlating with the behavioral impairment11. We also showed that CI is more efficient than centrality in identifying integrator hubs, with another measure of hub importance, the HITS score only detecting the LO but missing other integrator hubs (supplementary Table S3). In particular, we found a graded behavioral effect with high-ranking integrator hubs (LAcbSh and LO) having large effect sizes, whereas a mid-ranking hub (VM) had a moderate effect size and a low-ranking hub (S1Tr) had a minimal effect. This indicates that our analysis can predict behavioral effects. Among the integrator hubs tested, the identification of LAcbSh is consistent with its involvement in learning and memory (for review see ref. 81) and the integration of spatial information82. It is also a hub that is active in both APA tasks (Fig. 2b, c). LO is a critical prefrontal region for both decision making and the acquisition of hippocampus-dependent memories58,59,60, but its role in memory consolidation is less understood. VM, part of the motor thalamus, is the site of convergence of sensory (including nociceptive) and motor information and projects to the neocortex, particularly the mPFC83,84. It is involved in decision making but its role in learning and memory remains unclear.
Post-encoding replay of the spatiotemporal activity during learning in the hippocampal-neocortex network has been shown to be an important mechanism for memory maintenance and consolidation. In the neocortex, replay has been observed in the sensory (such as visual and auditory) or motor cortex engaged during learning in animals78,85 and humans86,87. Based on correlating with fMRI activation during learning, hippocampal replay has also been found during post-encoding rest in humans88,89,90. However, whether FC changes, such as the post-encoding RSNs observed here, reflects hippocampal-neocortical replay is unclear. High-frequency oscillations, called ripples91, which facilitate replay, has been reported to couple the hippocampus and association cortex after learning92, suggesting the presence of post-encoding FC. Combining fMRI and electrophysiology, a study in anesthetized monkey showed that hippocampal ripples coincide with the activation of the default mode network93. A similar result was recently found in mice by optical imaging94 and in humans by magnetoencephalography95. These findings suggest that post-encoding RSNs may reflect or coordinate replay.
Sleep plays several essential roles in supporting memory consolidation96. Replaying of the information that is encoded during wakefulness, and enhancing the crosstalk between the neocortex, hippocampus and thalamus are most active during slow-wave sleep5. Sleep also restores synaptic homeostasis, such as synaptic strength renormalization and dendritic spine down-selection, which prepares the brain for the next day’s experiences97. Relevant activity has also been observed using fMRI during or after sleep. Sleep can strengthen the hippocampal-prefrontal functional connectivity and stabilize the network induced by learning98,99. The post-encoding RSN between cortical and subcortical areas, particularly the striatum, was enhanced during and after sleep27,28, consistent with our findings that a broader network is involved. Replay could also be induced by presenting previously associated cues during sleep, leading to enhanced hippocampal-cortical FC and, particularly, increased network integration100. Together these findings indicate an important role of sleep in facilitating brain network reorganization to consolidate memory. As the post-encoding RSNs of both kinds of APA learning were measured after sleep, they may reflect the effects of sleep. The multiple days of sleep involved in the 5-Day APA may contribute to the different post-encoding RSNs compared to that of the 1-Day APA.
Systems consolidation can last for weeks, months or even years101,102, but networks transform and interact with each other over time remains unclear. Non-invasive rsfMRI allows longitudinal imaging in both animal models and humans to complement invasive imaging, such as c-fos imaging, which captures a snapshot during memory encoding or recall10,11. The changes in RSNs and their hubs that we observed across two time points support this ongoing plasticity. Among the common hubs, the primary sensory areas were identified on post-training day 1 whereas S2 was identified on post-training day 8. This suggests a transition from primary areas during early memory consolidation to association areas later in this process. On the other hand, the integrator hubs transit from the HPF and subcortical areas on post-training day 1 to neocortical areas on post-training day 8. This is consistent with the gradual reduction in the involvement of the HPF in systems consolidation2,42.
The hubs identified during the post-encoding period could be involved in forming, storing or recalling memory. We only inhibited the hubs after learning until one day before the probe test. This allowed us to determine their involvement in memory consolidation without affecting memory recall. Whether these hubs are also regions for memory storage will require further investigation. For instance, a recent study using activity tagging techniques reported that the ventrolateral orbital cortex, but not the sensory cortex, could store the contextual fear engram10. Future studies could combine similar techniques to determine the specific role of RSN hubs in memory storage.
We conducted the rsfMRI in this study using a sedative protocol that is reliable in detecting RSNs103,104,105 and post-encoding plasticity23,24, with a previous study showing that it does not affect memory consolidation106. Nonetheless, the detection of certain networks, such as the amygdala, which is involved in aversive learning paradigms, may be affected107. Ultrafast fMRI allows improved sensitivity for the ventral part of the brain108 and enables the detection of FC with amygdala nuclei, such as the PLCo (Fig. 1). Further development of awake imaging should facilitate more comprehensive mapping and testing of functional networks not only post-encoding but also during learning or recall. Due to the varying volume of the atlas-based seed region-of-interest (ROI), the sensitivity of a smaller region would be inferior due to less signal averaging. The highly sampled rsfMRI data (6000 time points) in this study partly compensated for this sensitivity issue. The use of a regionally optimized seed ROI109 or cryoprobe would further improve the sensitivity in future investigation.
Methods
Animals
121 male C57BL/6 mice (10–16 weeks old) were used in the experiments. Animals were housed in transparent cages and maintained on a 12 h light-dark cycle (lights on at 7 a.m. and off at 7 p.m.), 20–22 °C and 40–60% humidity. Food and water were provided ad libitum. Experiments were performed during the light phase. All experimental procedures were approved by the Animal Ethics Committee of the University of Queensland and conducted in compliance with the Queensland Animal Care and Protection Act 2001 and the Australian Code of Practice for the Care and Use of Animals for Scientific Purposes.
Experimental design
Two sets of animal experiments were conducted, one for hub identification (Fig. 1) and the other for hub verification (Fig. 5). Four groups of mice were used for hub identification: 1-Day APA (n = 10), 1-Day control (n = 9), 5-Day APA (n = 7), and 5-Day control (n = 5). Ten groups were used for hub verification: three groups for the common network, four groups for network integration, two negative controls (see surgical section for details) and one CNO control (n = 9) in naïve mice without surgery.
We used DREADDs for targeted inhibition (see surgical section for detail). 4 weeks after the surgery, animals were trained in the 1-Day APA task. Immediately after they finished the last training trial (T5), they were given a water-soluble CNO (CNO dihydrochloride; cat #6329, Tocris Bioscience) via intraperitoneal (i.p.) injection (1 mg/kg dissolved in saline), followed by CNO (1 mg/kg/day) dissolved in their drinking water to continuously suppress the network hub until one day before the probe test. This one-day interval allowed the CNO to be cleared from the body, thereby minimizing its interference in the probe test70. Memory retention in the probe test was used to examine whether memory consolidation was affected by hub inhibition.
Behavior
In the APA task, an animal stands in a rotating circular arena (diameter: 0.9 m; rotation speed: 1 rpm) with four pictures as spatial cues on each side of the wall (APA equipment: Bio-Signal Group). Once the animal enters an invisible sector (shock zone) that is stable in relation to a spatial cue, a mild electric shock (0.5 mA, 60 Hz, 500 ms) is administered. The animal needs to learn to use the visual cues to identify the exact location of the aversive zone and to avoid it. Two training protocols were used:
-
i.
In the 1-Day APA, animals received five 10 min training sessions with an inter-session interval of ~1 h completed in one day.
-
ii.
In the 5-Day APA, animals received one 10 min training session each day for 5 consecutive days.
Training started with a habituation session (15 min) one day before the training, during which the animal did not receive any shock. Nine days after the last training day, a 10 min probe test was performed to measure memory retention in the same environment. A foot shock was delivered when the animal entered the aversive zone in the probe test. During each training or probe session, behavior was recorded by a video camera. Two control groups were included: 1-Day sham control and 5-Day sham control. In these groups, animals went through the same APA procedure as the experimental group but did not receive any foot shocks. To ensure consistency, all the behavioral experiments were started at the same time of the day. For data analysis, the number of shocks and the time to first entrance of the shock zone were analyzed by Bio-Signal Track software. Repeated measures one-way ANOVA was performed using Prism (GraphPad Software LLC).
MRI
MRI was conducted on a 9.4T system (BioSpec 94/30, Bruker BioSpin MRI GmbH). Two rsfMRI scan sessions were performed on each animal for hub identification. Animals were initially anesthetized using 3% isoflurane in a 2:1 air and oxygen mixture. After being secured in an MRI-compatible holder using custom-made tooth and ear bars, a bolus of medetomidine was delivered via an i.p. catheter (0.05–0.1 mg/kg) and the isoflurane level was progressively reduced to 0.25–0.5% over 10 min, after which sedation was maintained by a constant i.p. infusion of medetomidine (0.1 mg/kg/h) using a syringe pump. Key physiological parameters, including arterial oxygenation saturation (SpO2), rectal temperature, heart rate and respiratory rate, were measured by an MRI-compatible monitoring system (SAII Inc). Body temperature was maintained at 36.5 °C with a heated waterbath.
After high-order shimming, structural T2-weighted MRI (resolution = 0.1 × 0.1 × 0.3 mm3) and the visual task were first conducted to ensure optimal physiology and neurovascular coupling. A flashing blue light at 5 Hz was delivered by an optical fiber in a block design with 21 s on and 39 s off. The rsfMRI scan was then acquired using multiband gradient-echo echo-planar imaging108 with TR/TE = 300/15 ms, 4 slice bands, matrix size = 128 × 64 (7/8 partial Fourier), thickness = 0.5 mm, gap = 0.1 mm, 16 axial slices covering the whole cerebrum with in-plane resolution of 0.3 × 0.3 mm2. 2000 volumes were acquired in 10 min and repeated three times with an inter-run interval of 2 min. To ensure consistency, the rsfMRI scan started ~45 min after the bolus injection of medetomidine.
Surgical procedure for DREADDs
Surgeries were performed at least one month before behavioral training. The animal was anesthetized with 1.5–2% isoflurane during surgery. Enrofloxacin (6 mg/kg) and carprofen (5 mg/kg) were injected subcutaneously to prevent infection and relieve pain and inflammation, respectively. Body temperature was maintained at 37 °C with a heating pad. During surgery, 0.25–0.3 μL of virus (AAV2/1-pSyn-hM4D(Gi)-T2A-mScarlet) was injected into the following target areas based on the rsfMRI connectivity map and the Paxions and Franklin Mouse Brain atlas, fifth edition. The coordinates (relative to Bregma) were:
Common hubs:
-
(i)
V1R (n = 10). ML: −2.30 mm; AP: −4.15 mm; DV: −0.50 mm.
-
(ii)
S2R (n = 8). ML: −3.70 mm; AP: −0.71 mm; DV: −1.40 mm.
-
(iii)
S1BFL (n = 11). ML: +2.88 mm; AP: −1.07 mm; DV: −0.85 mm.
Integrator hubs:
-
(iv)
LAcbShL (n = 7). ML: +1.70 mm; AP: +1.10 mm; DV: −3.85 mm.
-
(v)
LOR (n = 8). ML: −1.65 mm; AP: +2.57 mm; DV: −1.9 mm.
-
(vi)
VML (n = 10). ML: +0.75 mm; AP: −1.18 mm; DV: −4.05 mm.
-
(vii)
S1TrR (n = 7). ML: −1.63 mm; AP: −1.60 mm; DV: −0.57 mm.
Negative control:
-
(viii)
VPMR (n = 5). ML: −1.5 mm; AP: −1.80 mm; DV: −3.3 mm.
-
(ix)
FrAR (n = 5). ML: −1.70 mm; AP: +3.00 mm; DV: −0.50 mm.
Virus was injected using a Nanoject III (Drummond Scientific) with a slow injection rate (0.03 μL/min) over 10 min. The glass pipette was retained in place for another 6 min and then slowly retracted. After injection, the wound was closed using Vetbond (3 M) and sutured. Enrofloxacin and carprofen were administered for another two days. Animals were kept in their home cage (group housing of 2–4 animals per cage) for 4 weeks to recover and to allow expression of the virus before APA testing.
Behavior and CNO treatment
The same 1-Day APA task was used for spatial memory training. Immediately after the 1-Day APA training, water-soluble CNO was administered to the animals (1 mg/kg, i.p.) to inhibit the neuroactivity of the target brain regions. Water containing CNO (1 mg/kg/day) was then provided for 7 days to keep the target brain areas inhibited during memory consolidation. This was replaced with normal water 24 h before the probe test to minimize the effects of CNO on behavioral performance. On post-training day 8, a 10 min probe trial was performed to test memory retention.
Histology
Mice were administed an overdose of sodium pentobarbitone and transcardially perfused with 40 ml of phosphate-buffered saline (PBS), followed by 45 ml of 4% paraformaldehyde in PBS for fixation. The brain was extracted and fixed at 4 °C for 12–24 h. It was then washed once with PBS and transferred to a 30% sucrose solution for 36 h prior to sectioning. 40 μm thick sections were cut using a sliding microtome and collected in a 1:6 series. Cell nuclei were stained by 4’,6-diamidino-2-phenylindole (DAPI, catalog #6329; Sigma Aldrich). Sections were first washed once in PBS for 10 min, and then incubated in 1:5000 DAPI-PBS solution for 15 min at room temperature. After two washes, the sections were mounted on SuperFrost slides using fluorescence mounting medium (Dako, Agilent). Images were captured using a slide scanner (Metafer VSlide Scanner, MetaSystems) and microscope (Axio Imager Z2, Zeiss) with a 20 × 0.8 NA/0.55 mm objective lens.
rsfMRI data processing
The rsfMRI data were processed using MATLAB (MathWorks Inc), FSL (v5.0.11, https://fsl.fmrib.ox.ac.uk/fsl), AFNI (ver 17.2.05, National Institutes of Health, USA) and ANTs (v2.3.1, http://stnava.github.io/ANTs). The k-space data of the multiband EPI were first phase-corrected and reconstructed in MATLAB. After motion correction by FSL mcflirt, the geometric distortion was corrected by FSL TOPUP. The brain mask was extracted automatically using PCNN3D110, followed by manual editing. Nuisance signals, including quadratic drift, six motion parameters and their derivatives, ten principal components from tissues outside the brain which included muscle and scalp, and mean signal of the cerebrospinal fluid from a manually drawn ventricular mask, were then regressed out111. The data were band-pass filtered at 0.01–0.3 Hz to account for any potential frequency shift under sedation. This frequency range could also remove the aliased respiratory and cardiac signal variations in the high sampling rate data. The rsfMRI was coregistered to an EPI template by linear and nonlinear transformations using ANTs. The data were then smoothed by a 0.6 mm Gaussian kernel.
Seed-based correlation analysis was used to measure FC across the brain. Based on the Australian Mouse Brain Mapping Consortium (AMBMC) atlas (https://imaging.org.au/AMBMC/AMBMC), the brain was divided into 190 bilateral ROIs in the cortex, hippocampus, thalamus, and basal ganglia in accordance with the parcellation in the Paxions and Franklin mouse brain atlas112. The DSURQE atlas (https://wiki.mouseimaging.ca/) was used to label regions not yet defined in the AMBMC atlas (40 ROIs), such as the amygdala, hypothalamus, midbrain and brainstem (pons). The combined 230 ROIs were used in the following seed-based correlation analysis. The mean time-series of each brain region was extracted as a seed signal. Pearson’s correlation coefficients between seed time-courses were calculated using AFNI 3dNetCorr. Fisher’s z-transformation was used to convert correlation coefficients to z values. Connectivity matrices from the three repeated scans were calculated for each animal. Quality control (QC) was conducted based on the presence of visual task activation. If the visual activation was not detectable, the physiological condition and neurovascular coupling were regarded as sub-optimal, and the scan was discarded. Based on this criterion, 24% of scans were discarded (Supplementary Table S4). As an animal’s physiological condition can vary between scanning sessions, it may show a visual response on post-training day 1 but not on post-training day 8, or vice versa, as a result of which the dataset was not one-to-one matched at the two time points. The matrices of each animal at each time point that passed the QC were averaged.
Between-group differences were calculated by two-sample t test and thresholded at p < 0.05 (FDR corrected) using the Network Based Statistics toolbox (https://sites.google.com/site/bctnet/comparison/nbs). To detect common network hubs, the network and behavioral correlations were each thresholded at p < 0.05, uncorrected (see details in the subsection “Behavior-correlated common networks between 1-Day and 5-Day APA” below). To identify integrator hubs, three uncorrected thresholds, p < 0.05, p < 0.01 and p < 0.005, were used to generate unweighted (t-score) network matrices for CI analysis. Significant connections were overlaid on the 3D-rendered brain atlas using BrainNet Viewer (https://www.nitrc.org/projects/bnv/).
Group independent component analysis with dual regression
Group ICA was performed on preprocessed rsfMRI datasets for the 1-Day APA and 1-Day control using FSL MELODIC (https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/MELODIC). After separation into 30 components, dual regression (http://fsl.fmrib.ox.ac.uk/fsl/fslwiki/DualRegression) was performed to determine the between-group difference. A two-sample t-test was conducted using FSL-glm with the cluster-level correction estimated by AFNI 3DClusterSim (p < 0.05, two-tail, overall family-wise error rate p < 0.05). The ICA components were classified into signal and artifact based on other rsfMRI studies in mice113. 21 components on post-training day 1 and 18 components on post-training day 8 were identified as the signal. The group-level spatial ICA maps were thresholded at |Z| ≥ 1.96 (equivalent to p < 0.05, uncorrected) for visualization.
Graph theory analysis
To characterize the RSNs, both weighted and unweighted versions, when applicable, of the following graph theory parameters were calculated by the Brain Connectivity Toolbox (http://www.brain-connectivity-toolbox.net) and the graph functions in Matlab:
Global efficiency:
where \({d}_{{ij}}\) is the shortest path length between nodes \(i\) and \(j\), and N is the total number of nodes. A value of 1 indicates maximum efficiency.
Modularity was calculated using the Newman’s spectral community detection algorithm:
where the network is fully subdivided into a set of nonoverlapping modules \(M\), and \({e}_{{uv}}\) is the proportion of all links that connect nodes in module \(u\) with nodes in module \(v\). The higher the Q value, the larger degree of network segregation.
Transitivity:
where \({k}_{i}\) is the degree of a node \(i\), and \({t}_{i}\) is the number of triangles around a node \(i\). Transitivity is a variant of the clustering coefficient. The higher the T value, the larger the degree of network segregation.
Degree centrality of a node \(i\):
where \({a}_{{ij}}\) is the connection between nodes \(i\) and \(j\). \({a}_{{ij}}\) = 1 when a link (\(i\), \(j\)) exists, and \({a}_{{ij}}\) = 0 otherwise (\({a}_{{ii}}\) = 0 for all \(i\)).
Closeness centrality of node \(i\):
Betweenness centrality of node \(i\):
where \({\rho }_{{hj}}\) is the number of shortest paths between nodes \(h\) and \(j\), and \({\rho }_{{hj}}(i)\) is the number of shortest paths between \(h\) and \(j\) that pass through \(i\).
Eigenvector centrality of node \(i\):
where \(\lambda\) is a constant and x is the eigenvector of the binarized network matrix.
The HITS score is a link analysis algorithm used to assign authority and hub indices to a network114. The authority of a node indicates how many high-quality nodes link to it while the hub index of a node indicates how many links of this node are connected to high-quality nodes. Here we used a Matlab implementation (https://people.sc.fsu.edu/~jburkardt/m_src/hits/hits.html) to rank a post-encoding RSN hub based on its hub index, as the authority showed similar ranking (data not shown).
The giant component is defined as the largest connected component in a network11. It can be represented by the number of nodes in the largest connected component. The ratios of the giant components in the CI analysis were used to represent the change in the giant component when nodes were removed from the network.
To evaluate the small-world property of the brain network, the normalized characteristic path length, lambda, the normalized clustering coefficient, gamma, and the small-world index, sigma, were calculated. Individual FC matrices were first thresholded in a predefined range (0.01 < z < 0.07, step 0.01, with z = 0.0254 corresponding to p = 0.05, z = 0.07 corresponding to p < 0.00001). The maximum threshold selected was the value for which all FC matrices were fully connected (no isolated node). Sigma, gamma and lambda were calculated from the thresholded network using a Matlab code (https://github.com/mdhumphries/SmallWorldNess). To calculate sigma, the Erdos Renyl random graph was used to estimate the path lengths and clustering coefficients of random network by inputting the total node number (230) and mean degree of the thresholded network. After plotting the trends in the above range, AUC was calculated and a two-sample t-test was performed to examine the difference between the APA and control groups.
FC–behavior correlation
We correlated the strength of each significant FC with two behavioral indices (the number of shocks and the time to first entrance of the shock zone) in the probe test using Pearson’s correlation coefficient, with p < 0.05 (two-tailed) regarded as significant. After correlation analysis, we sorted the significant connections based on their absolute value of the correlation coefficient.
Behavior-correlated common networks between 1-Day and 5-Day APA
We defined the behavior-correlated common network as a connection shown in both tasks, the connection strength of which correlated with memory retention. This required a connection to fulfill two network thresholds and one behavioral threshold, which together are equivalent to a much stricter threshold that reduces both true and false positive rates. Therefore, the node-wise threshold was reduced to achieve suitable power while controlling for the false positive rate. We detected overlapping connections between the 1-Day and 5-Day APA tasks from their network matrices obtained by two-sample t-test between the APA and control groups with each network being threhsolded at p < 0.05, uncorrected. From the overlapping connections, we calculated FC–behavior correlations and sorted (ranked) the significant connections (p < 0.05, uncorrected) by their absolute value of behavior correlation. Combining these uncorrected thresholds together resulted in a family-wise error rate of p < 0.05 according to the null distribution estimated by a permutation test.
CI analysis
CI analysis was applied to the between-group difference network using an optimized implementation69 (https://github.com/zhfkt/ComplexCi/releases) that calculates the value of each node with the following formula:
where \({k}_{i}\) is the degree of node \(i\), and \(\delta B(i,\,l)\) is the frontier of the ball of radius \(l\) which is the set of nodes at a particular distance from \(i\). The value is calculated iteratively by removing nodes until all nodes in the network are eliminated67. In this analysis, we used the ball radius \(l=2\) as we found that a larger radius gave nearly the same results. We ranked the nodes according to how fast the size (number of nodes) of the “giant component” collapsed by removing the selected node. To find the reliable ranking list, we calculated the mean CI ranking of each node under three thresholds (p < 0.05, p < 0.01, p < 0.005) and sorted the node by the mean CI rank value. The top 10 CI nodes were then defined as high (1–3), middle (4–7) and low (8–10) ranking hubs.
Null distribution of network property analysis
To determine whether the difference matrices between the APA and control groups were significantly different from random networks, 5000 random networks were created using the function “null_model_und_sign” of the Brain Connectivity Toolbox that preserves the degree and strength distributions of the real network. For each of the 5000 random networks, curves of three network properties, including the giant component, global efficiency and modularity were generated under thresholds ranging from t = 2 to 3.8 (step of 0.2). The AUC of each curve was calculated to form the null distribution for each network property.
Null distribution of common network analysis
To estimate the null distribution of the common network, 5000 permutations of FC matrices were created by randomly assigning each individual FC matrix into the APA groups and controls. Between-group differences of these permutated matrices were tested by two-sample t-tests and thresholded at p < 0.05, uncorrected. The common FC of the permutated 1-Day APA and 5-Day APA data was correlated with the Nshock or Tenter of the probe test and thresholded at p < 0.05. The number of connections surviving these thresholds from the 5000 permutations formed the null distribution of the common network detection (Supplementary Fig. S2a).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The fMRI data and the AMBMC atlas labels generated in this study have been deposited in the Zenodo database at https://zenodo.org/deposit/8161802 (https://doi.org/10.5281/zenodo.8161802). Source data in this paper are provided in the Supplementary Information and a Source Data file. Source data are provided with this paper.
Code availability
Data analyses were conducted using public domain software, including: FSL (https://www.fmrib.ox.ac.uk/fsl), AFNI (https://afni.nimh.nih.gov/), ANTs (http://stnava.github.io/ANTs/), Brain connectivity toolbox (https://sites.google.com/site/bctnet/), Network Based Statistics toolbox (https://sites.google.com/site/bctnet/comparison/nbs), HITS score (https://people.sc.fsu.edu/~jburkardt/m_src/hits/hits.html), Small-worldness (https://github.com/mdhumphries/SmallWorldNess), ComplexCI (https://github.com/zhfkt/ComplexCi/releases), 3D-PCNN (https://sites.google.com/site/chuanglab/software/3d-pcnn), and BrainNet Viewer (https://www.nitrc.org/projects/bnv/).
References
Frankland, P. W. & Bontempi, B. The organization of recent and remote memories. Nat. Rev. Neurosci. 6, 119–130 (2005).
Squire, L. R., Genzel, L., Wisted, J. T. & Morris, R. G. M. Memory consolidation. Cold Spring Harb. Perspect. Biol. 7, a021766 (2015).
Fell, J. & Axmacher, N. The role of phase synchronization in memory processes. Nat. Rev. Neurosci. 12, 105–118 (2011).
Jutras, M. J. & Buffalo, E. A. Synchronous neural activity and memory formation. Curr. Opin. Neurobiol. 20, 150–155 (2010).
Klinzing, J. G., Niethard, N. & Born, J. Mechanisms of systems memory consolidation during sleep. Nat. Neurosci. 22, 1598–1610 (2019).
Joo, H. R. & Frank, L. M. The hippocampal sharp wave–ripple in memory retrieval for immediate use and consolidation. Nat. Rev. Neurosci. 19, 744–757 (2018).
Frankland, P. W., O’Brien, C., Ohno, M., Kirkwood, A. & Silva, A. J. Alpha-CaMKII-dependent plasticity in the cortex is required for permanent memory. Nature 411, 309–313 (2001).
Kitamura, T. et al. Engrams and circuits crucial for systems consolidation of a memory. Science 356, 73–78 (2017).
Nyberg, L., Forkstam, C., Petersson, K. M., Cabeza, R. & Ingvar, M. Brain imaging of human memory systems: Between-systems similarities and within-system differences. Cogn. Brain Res. 13, 281–292 (2002).
Roy, D. S. et al. Brain-wide mapping reveals that engrams for a single memory are distributed across multiple brain regions. Nat. Commun. 13, 1799 (2022).
Vetere, G. et al. Chemogenetic interrogation of a brain-wide fear memory network in mice. Neuron 94, 363–374.e4 (2017).
Staresina, B. P., Alink, A., Kriegeskorte, N. & Henson, R. N. Awake reactivation predicts memory in humans. Proc. Natl. Acad. Sci. USA. 110, 21159–21164 (2013).
Fox, M. D. & Raichle, M. E. Spontaneous fluctuations in brain activity observed with functional magnetic resonance imaging. Nat. Rev. Neurosci. 8, 700–711 (2007).
Smith, S. M. et al. Functional connectomics from resting-state fMRI. Trends Cogn. Sci. 17, 666–682 (2013).
Power, J. D., Schlaggar, B. L. & Petersen, S. E. Studying brain organization via spontaneous fMRI signal. Neuron 84, 681–696 (2014).
Pievani, M., Filippini, N., van den Heuvel, M. P., Cappa, S. F. & Frisoni, G. B. Brain connectivity in neurodegenerative diseases—from phenotype to proteinopathy. Nat. Rev. Neurol. 10, 620–633 (2014).
Yu, M., Sporns, O. & Saykin, A. J. The human connectome in Alzheimer disease—relationship to biomarkers and genetics. Nat. Rev. Neurol. 17, 545–563 (2021).
Greicius, M. D., Krasnow, B., Reiss, A. L. & Menon, V. Functional connectivity in the resting brain: a network analysis of the default mode hypothesis. Proc. Natl. Acad. Sci. USA. 100, 253–258 (2003).
Mišić, B. & Sporns, O. From regions to connections and networks: New bridges between brain and behavior. Curr. Opin. Neurobiol. 40, 1–7 (2016).
Bullmore, E. & Sporns, O. Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009).
Albert, N. B., Robertson, E. M. & Miall, R. C. The resting human brain and motor learning. Curr. Biol. 19, 1023–1027 (2009).
Sami, S., Robertson, E. M. & Chris Miall, R. The time course of task-specific memory consolidation effects in resting state networks. J. Neurosci. 34, 3982–3992 (2014).
Nasrallah, F. A., To, X. V., Chen, D.-Y., Routtenberg, A. & Chuang, K.-H. Functional connectivity MRI tracks memory networks after maze learning in rodents. Neuroimage 127, 196–202 (2016).
Shah, D. et al. Acquisition of spatial search strategies and reversal learning in the Morris water maze depend on disparate brain functional connectivity in mice. Cereb. Cortex 29, 4519–4529 (2019).
Dresler, M. et al. Mnemonic training reshapes brain networks to support superior memory. Neuron 93, 1227–1235.e6 (2017).
Tambini, A., Ketz, N. & Davachi, L. Enhanced brain correlations during rest are related to memory for recent experiences. Neuron 65, 280–290 (2010).
Debas, K. et al. Off-line consolidation of motor sequence learning results in greater integration within a cortico-striatal functional network. Neuroimage 99, 50–58 (2014).
Vahdat, S., Fogel, S., Benali, H. & Doyon, J. Network-wide reorganization of procedural memory during NREM sleep revealed by fMRI. Elife 6, e24987 (2017).
Schlichting, M. L. & Preston, A. R. Memory reactivation during rest supports upcoming learning of related content. Proc. Natl. Acad. Sci. USA. 111, 15845–15850 (2014).
Tompary, A. & Davachi, L. Consolidation promotes the emergence of representational overlap in the hippocampus and medial prefrontal cortex. Neuron 96, 228–241.e5 (2017).
Reid, A. T. et al. Advancing functional connectivity research from association to causation. Nat. Neurosci. 22, 1751–1760 (2019).
Duncan, N. W. & Northoff, G. Overview of potential procedural and participant-related confounds for neuroimaging of the resting state. J. Psychiatry Neurosci. 38, 84–96 (2013).
Gonzalez-Castillo, J., Kam, J. W. Y., Hoy, C. W. & Bandettini, P. A. How to interpret resting-state fMRI: ask your participants. J. Neurosci. 41, 1130–1141 (2021).
Cole, D. M., Smith, S. M. & Beckmann, C. F. Advances and pitfalls in the analysis and interpretation of resting-state FMRI data. Front. Syst. Neurosci. 4, 8 (2010).
Drew, P. J., Mateo, C., Turner, K. L., Yu, X. & Kleinfeld, D. Ultra-slow oscillations in fMRI and resting-state connectivity: neuronal and vascular contributions and technical confounds. Neuron 107, 782–804 (2020).
Matthews, P. M. & Hampshire, A. Clinical concepts emerging from fMRI functional connectomics. Neuron 91, 511–528 (2016).
Bielczyk, N. Z. et al. Disentangling causal webs in the brain using functional magnetic resonance imaging: a review of current approaches. Netw. Neurosci. 3, 237–273 (2019).
Paul, C. M., Magda, G. & Abel, S. Spatial memory: theoretical basis and comparative review on experimental methods in rodents. Behav. Brain Res. 203, 151–164 (2009).
Milczarek, M. M. M. et al. Spatial memory engram in the mouse retrosplenial cortex. Curr. Biol. 28, 1975–1980.e6 (2018).
Ranganath, C. & Ritchey, M. Two cortical systems for memory-guided behaviour. Nat. Rev. Neurosci. 13, 713–726 (2012).
Eichenbaum, H. Prefrontal–hippocampal interactions in episodic memory. Nat. Rev. Neurosci. 18, 547–558 (2017).
Tonegawa, S., Morrissey, M. D. & Kitamura, T. The role of engram cells in the systems consolidation of memory. Nat. Rev. Neurosci. 19, 485–498 (2018).
Cohen, J. R. & D’Esposito, M. The segregation and integration of distinct brain networks and their relationship to cognition. J. Neurosci. 36, 12083–12094 (2016).
Bassett, D. S. et al. Dynamic reconfiguration of human brain networks during learning. Proc. Natl. Acad. Sci. USA. 108, 7641–7646 (2011).
Shine, J. M. et al. The dynamics of functional brain networks: Integrated network states during cognitive task performance. Neuron 92, 544–554 (2016).
Finc, K. et al. Dynamic reconfiguration of functional brain networks during working memory training. Nat. Commun. 11, 2435 (2020).
Chowdhury, A. & Caroni, P. Time units for learning involving maintenance of system-wide cFos expression in neuronal assemblies. Nat. Commun. 9, 1–11 (2018).
Smolen, P., Zhang, Y. & Byrne, J. H. The right time to learn: mechanisms and optimization of spaced learning. Nat. Rev. Neurosci. 17, 77–88 (2016).
Roth, B. L. DREADDs for neuroscientists. Neuron 89, 683–694 (2016).
Lesburguères, E., Sparks, F. T., O’Reilly, K. C. & Fenton, A. A. Active place avoidance is no more stressful than unreinforced exploration of a familiar environment. Hippocampus 26, 1481–1485 (2016).
Cimadevilla, J. M., Fenton, A. A. & Bures, J. New spatial cognition tests for mice: passive place avoidance on stable and active place avoidance on rotating arenas. Brain Res. Bull. 54, 559–563 (2001).
Nagao, S. Pontine nuclei-mediated cerebello-cerebral interactions and its functional role. Cerebellum 3, 11–15 (2004).
Wesson, D. W. & Wilson, D. A. Sniffing out the contributions of the olfactory tubercle to the sense of smell: Hedonics, sensory integration, and more? Neurosci. Biobehav. Rev. 35, 655–668 (2011).
Tucker, D. M., Hartry-Speiser, A., McDougal, L., Luu, P. & Degrandpre, D. Mood and spatial memory: emotion and right hemisphere contribution to spatial cognition. Biol. Psychol. 50, 103–125 (1999).
Metcalfe, J., Funnell, M. & Gazzaniga, M. S. Right-hemisphere memory superiority: studies of a split-brain patient. Psychol. Sci. 6, 157–164 (1995).
Shinohara, Y. et al. Right-hemispheric dominance of spatial memory in split-brain mice. Hippocampus 22, 117–121 (2012).
Castro, D. C., Cole, S. L. & Berridge, K. C. Lateral hypothalamus, nucleus accumbens, and ventral pallidum roles in eating and hunger: interactions between homeostatic and reward circuitry. Frontiers in Systems Neuroscience 9, 90 (2015).
Vafaei, A. A. & Rashidy-Pour, A. Reversible lesion of the rat’s orbitofrontal cortex interferes with hippocampus-dependent spatial memory. Behav. Brain Res. 149, 61–68 (2004).
Frey, S. & Petrides, M. Orbitofrontal cortex and memory formation. Neuron 36, 171–176 (2002).
Lesburguères, E. et al. Early tagging of cortical networks is required for the formation of enduring associative memory. Science 331, 924–928 (2011).
Hampson, M., Driesen, N. R., Skudlarski, P., Gore, J. C. & Constable, R. T. Brain connectivity related to working memory performance. J. Neurosci. 26, 13338–13343 (2006).
Sneve, M. H. et al. Decoupling of large-scale brain networks supports the consolidation of durable episodic memories. Neuroimage 153, 336–345 (2017).
Keerativittayayut, R., Aoki, R., Sarabi, M. T., Jimura, K. & Nakahara, K. Large-scale network integration in the human brain tracks temporal fluctuations in memory encoding performance. Elife 7, e32696 (2018).
Bassett, D. S. & Bullmore, E. Small-world brain networks. Neuroscientist 12, 512–523 (2006).
Zhang, D., Wang, J., Liu, X., Chen, J. & Liu, B. Aberrant brain network efficiency in Parkinson’s disease patients with tremor: a multi-modality study. Front. Aging Neurosci. 7, 31 (2015).
Dablander, F. & Hinne, M. Node centrality measures are a poor substitute for causal inference. Sci. Rep. 9, 1–13 (2019).
Morone, F. & Makse, H. A. Influence maximization in complex networks through optimal percolation. Nature 524, 65–68 (2015).
Del Ferraro, G. et al. Finding influential nodes for integration in brain networks using optimal percolation theory. Nat. Commun. 9, 2274 (2018).
Zhu, F. Improved collective influence of finding most influential nodes based on disjoint-set reinsertion. Sci. Rep. 8, 14503 (2018).
Jendryka, M. et al. Pharmacokinetic and pharmacodynamic actions of clozapine-N-oxide, clozapine, and compound 21 in DREADD-based chemogenetics in mice. Sci. Rep. 9, 1–14 (2019).
Raper, J. et al. Metabolism and distribution of clozapine-N-oxide: Implications for nonhuman primate chemogenetics. ACS Chem. Neurosci. 8, 1570–1576 (2017).
Xu, N. et al. Functional connectivity of the brain across rodents and humans. Front. Neurosci. 16, 1–27 (2022).
Chuang, K.-H. & Nasrallah, F. A. Functional networks and network perturbations in rodents. Neuroimage 163, 419–436 (2017).
Kitsak, M. et al. Identification of influential spreaders in complex networks. Nat. Phys. 6, 888–893 (2010).
Salavaty, A., Ramialison, M. & Currie, P. D. Integrated value of influence: an Integrative method for the identification of the most influential nodes within networks. Patterns 1, 100052 (2020).
Bando, S. Y. et al. Complex network analysis of CA3 transcriptome reveals pathogenic and compensatory pathways in refractory temporal lobe epilepsy. PLoS One 8, e79913 (2013).
van de Ven, V., Jacobs, C. & Sack, A. T. Topographic contribution of early visual cortex to short-term memory consolidation: a transcranial magnetic stimulation study. J. Neurosci. 32, 4–11 (2012).
Ji, D. & Wilson, M. A. Coordinated memory replay in the visual cortex and hippocampus during sleep. Nat. Neurosci. 10, 100–107 (2007).
Kumar, N., Manning, T. F. & Ostry, D. J. Somatosensory cortex participates in the consolidation of human motor memory. PLOS Biol. 17, e3000469 (2019).
Long, X. & Zhang, S.-J. J. A novel somatosensory spatial navigation system outside the hippocampal formation. Cell Res. 31, 649–663 (2021).
Rinaldi, A., Oliverio, A. & Mele, A. Spatial memory, plasticity and nucleus accumbens. Rev. Neurosci. 23, 527–541 (2012).
Ito, R., Robbins, T. W., Pennartz, C. M. & Everitt, B. J. Functional interaction between the hippocampus and nucleus accumbens shell is necessary for the acquisition of appetitive spatial context conditioning. J. Neurosci. 28, 6950–6959 (2008).
Sieveritz, B., García-Muñoz, M. & Arbuthnott, G. W. Thalamic afferents to prefrontal cortices from ventral motor nuclei in decision-making. Eur. J. Neurosci. 49, 646–657 (2019).
Monconduit, L., Bourgeais, L., Bernard, J.-F., Le Bars, D. & Villanueva, L. Ventromedial thalamic neurons convey nociceptive signals from the whole body surface to the dorsolateral neocortex. J. Neurosci. 19, 9063–9072 (1999).
Ramanathan, D. S., Gulati, T. & Ganguly, K. Sleep-dependent reactivation of ensembles in motor cortex promotes skill consolidation. PLoS Biol. 13, e1002263 (2015).
Eichenlaub, J. B. et al. Replay of learned neural firing sequences during rest in human motor cortex. Cell Rep. 31, 107581 (2020).
Liu, Y., Dolan, R. J., Kurth-Nelson, Z. & Behrens, T. E. J. Human replay spontaneously reorganizes experience. Cell 178, 640–652.e14 (2019).
Tambini, A. & Davachi, L. Persistence of hippocampal multivoxel patterns into postencoding rest is related to memory. Proc. Natl. Acad. Sci. USA. 110, 19591–19596 (2013).
Schuck, N. W. & Niv, Y. Sequential replay of nonspatial task states in the human hippocampus. Science 364, eaaw5181 (2019).
Schapiro, A. C., McDevitt, E. A., Rogers, T. T., Mednick, S. C. & Norman, K. A. Human hippocampal replay during rest prioritizes weakly learned information and predicts memory performance. Nat. Commun. 9, 3920 (2018).
Buzsáki, G. Hippocampal sharp wave-ripple: a cognitive biomarker for episodic memory and planning. Hippocampus 25, 1073–1188 (2015).
Khodagholy, D., Gelinas, J. N. & Buzsáki, G. Learning-enhanced coupling between ripple oscillations in association cortices and hippocampus. Science 358, 369–372 (2017).
Kaplan, R. et al. Hippocampal sharp-wave ripples influence selective activation of the default mode network. Curr. Biol. 26, 686–691 (2016).
Pedrosa, R. et al. Hippocampal gamma and sharp wave/ripples mediate bidirectional interactions with cortical networks during sleep. Proc. Natl. Acad. Sci. USA. 119, e2204959119 (2022).
Higgins, C. et al. Replay bursts in humans coincide with activation of the default mode and parietal alpha networks. Neuron 109, 882–893.e7 (2021).
Rasch, B. & Born, J. About sleep’s role in memory. Physiol. Rev. 93, 681–766 (2013).
Tononi, G. & Cirelli, C. Sleep and the price of plasticity: from synaptic and cellular homeostasis to memory consolidation and integration. Neuron 81, 12–34 (2014).
Himmer, L., Schönauer, M., Heib, D. P. J., Schabus, M. & Gais, S. Rehearsal initiates systems memory consolidation, sleep makes it last. Sci. Adv. 5, eaav1695 (2019).
van den Berg, N. H. et al. Sleep strengthens resting-state functional communication between brain areas involved in the consolidation of problem-solving skills. Learn. Mem. 30, 25–35 (2023).
Berkers, R. M. W. J. et al. Cued reactivation during slow-wave sleep induces brain connectivity changes related to memory stabilization. Sci. Rep. 8, 16958 (2018).
Dudai, Y., Karni, A. & Born, J. The consolidation and transformation of memory. Neuron 88, 20–32 (2015).
Kandel, E. R., Dudai, Y. & Mayford, M. R. The molecular and systems biology of memory. Cell 157, 163–186 (2014).
Grandjean, J. et al. Common functional networks in the mouse brain revealed by multi-centre resting-state fMRI analysis. Neuroimage 205, 116278 (2020).
Nasrallah, F. A., Tay, H. & Chuang, K.-H. Detection of functional connectivity in the resting mouse brain. Neuroimage 86, 417–424 (2014).
Grandjean, J., Schroeter, A., Batata, I. & Rudin, M. Optimization of anesthesia protocol for resting-state fMRI in mice based on differential effects of anesthetics on functional connectivity patterns. Neuroimage 102P2, 838–847 (2014).
Nasrallah, F. A., To, X. V., Chen, D.-Y., Routtenberg, A. & Chuang, K.-H. Resting state functional connectivity data supports detection of cognition in the rodent brain. Data Brief 7, 1156–1164 (2016).
Liang, Z., King, J. & Zhang, N. Neuroplasticity to a single-episode traumatic stress revealed by resting-state fMRI in awake rats. Neuroimage 103, 485–491 (2014).
Lee, H.-L., Li, Z., Coulson, E. J. & Chuang, K.-H. Ultrafast fMRI of the rodent brain using simultaneous multi-slice EPI. Neuroimage 195, 48–58 (2019).
Kreitz, S., Alonso, B., de, C., Uder, M. & Hess, A. A new analysis of resting state connectivity and graph theory reveals distinctive short-term modulations due to whisker stimulation in rats. Front. Neurosci. 12, 1–19 (2018).
Chou, N., Wu, J., Bai Bingren, J., Qiu, A. & Chuang, K.-H. Robust automatic rodent brain extraction using 3-D pulse-coupled neural networks (PCNN). IEEE Trans. Image Process. 20, 2554–2564 (2011).
Chuang, K.-H. et al. Evaluation of nuisance removal for functional MRI of rodent brain. Neuroimage 188, 694–709 (2019).
Ullmann, J. F. P. et al. Segmentation of the C57BL/6J mouse cerebellum in magnetic resonance images. Neuroimage 62, 1408–1414 (2012).
Zerbi, V., Grandjean, J., Rudin, M. & Wenderoth, N. Mapping the mouse brain with rs-fMRI: an optimized pipeline for functional network identification. Neuroimage 123, 11–21 (2015).
Ringstad, G. et al. Brain-wide glymphatic enhancement and clearance in humans assessed with MRI. JCI Insight 3, e121537 (2018).
Acknowledgements
This study was funded by Australian Research Council Discovery Project grant #180103319 to K.C., P.O. and P.S. Z.L. was supported by a Research Training Program scholarship of the University of Queensland. We thank Ms Rowan Tweedale for proof reading and Prof Ethan Scott for helpful discussion.
Author information
Authors and Affiliations
Contributions
K.C., P.O., and P.S. contributed to designing experiments. Z.L., H.L., and D.A. conducted experiments. Z.L. and K.C. conducted data analysis. Z.L., P.O., P.S., and K.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Andreas Hess and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Z., Athwal, D., Lee, HL. et al. Locating causal hubs of memory consolidation in spontaneous brain network in male mice. Nat Commun 14, 5399 (2023). https://doi.org/10.1038/s41467-023-41024-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-41024-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.