Abstract
The high density of macromolecules affecting proteins due to volume exclusion has been discussed in theory but numerous in vivo experiments cannot be sufficiently understood taking only pure entropic stabilization into account. Here, we show that the thermodynamic stability of a beta barrel protein increases equally at all atomic levels comparing crowded environments with dilute conditions by applying multidimensional high-resolution NMR spectroscopy in a systematic manner. Different crowding agents evoke a pure stabilization cooperatively and do not disturb the surface or integrity of the protein fold. The here developed methodology provides a solid base that can be easily expanded to incorporate e.g. binding partners to recognize functional consequences of crowded conditions. Our results are relevant to research projects targeting soluble proteins in vivo as it can be anticipated that their thermodynamic stability increase comparably and has consequently to be taken into account to coherently understand intracellular processes.
Similar content being viewed by others
Introduction
All essential processes of life occur within the cellular environment and are therein crowded by high concentrations of macromolecules, such as proteins and nucleic acids. For the cellular lumen of E. coli, a volume fraction of 20–40% of macromolecules has been reported thus contributing to an overall concentration of up to 400 g/L of macromolecules which are present in a living cell1,2. Consequently, this dense environment has to be considered in the study of protein stability, dynamics, and function since the wealth of various macromolecules affects local concentrations of proteins and reactants3,4, potentially hinders molecular diffusion5,6 and has multiple, highly complex effects on molecular interactions in the cytosol7,8,9. In this respect, it has indeed been shown that experimental results that are exclusively based on in vitro investigations of proteins, which have been isolated from their cellular environment are possibly misleading in conclusions drawn about their inherent stability, and conformational dynamics that are relevant for performing intracellular interactions. Thus it has been reported that macromolecular crowding (MC) has pronounced effects in numerous cases as it was found e.g. for FlgM, an intrinsically disordered protein, that it gains a distinct structure upon crowding10 or for the enzyme Pin1, that it is able to form unexpected complexes in a crowded environment11. Further, protein kinetics12 and dynamics13,14, enzymatic reaction rate constants15, ribonucleic acid folding stability16, ribozyme activity17, and the cytoskeletal properties itself18 were shown to be altered under MC. Moreover, the association of proteins is influenced by MC differing strongly from in vitro results in respect to aggregation19 and amyloid fibril formation20,21. Regarding protein folding and stability it was found that MC leads to a narrower folding funnel concerning the native state22, to a change in solvent accessibility23 or to a compaction of intrinsically disordered proteins24 or an unfolded protein ensemble25.
Despite those advances, a consistent universal model of the general influence of MC on the thermodynamic stability and conformational space proteins naturally occupy is missing. To fill this gap the field of MC advances top-down approaches. Here, protein signals are recorded in whole cells or in cellular lysates before degradation of reporter proteins takes place due to the cellular proteostasis machinery that is inherently present in vivo. Those measurements yield indeed fundamental results26,27 but are limited in spatial resolution and are not easily reproducible since the composition of cells, and their cellular extract is highly diverse and strongly depends on individual culturing conditions. Therefore, we have chosen here a bottom-up approach to investigate effects of MC on protein stability using precisely defined conditions. This setup presents no restrictions concerning experimental time due to the prevention of potential degradation of the sample under investigation. One can achieve the highest spectral resolution possible in order to monitor effects of MC at a residue and even an atomistic level on the protein under study. Here, we present an analysis of the thermodynamic stability of the cold shock protein B from Bacillus subtilis, BsCspB, at atomic resolution by probing almost all 15N and 13C bound protons. BsCspB as a prominent model protein representing two-state folding (Fig. 1) has been chosen here as it has been structurally well characterized by using X-ray crystallography and high-resolution NMR spectroscopy before28,29, and its thermodynamic stability has been thoroughly investigated under dilute conditions applying multiple experimental techniques30,31,32,33. Here, the experimentally obtained data have been monitored by conducting chemically induced unfolding of BsCspB in absence and presence of MC applying high-resolution heteronuclear multidimensional NMR spectroscopy. We have acquired both two-dimensional heteronuclear 1H-15N as well as 1H-13C HSQC spectra of BsCspB, respectively, varying in the concentration of urea ranging between curea = 0 M and curea = 6.3 M. The globular sugar polymer dextran having 20 kDa (Dex20) and the smaller, less polar polyethylene glycol of 1 kDa molecular weight (PEG1) have been used as crowding agents. This strategic choice is based on a recent finding obtained by applying a combination of fluorescence, circular dichroism and NMR spectroscopy showing that PEG and Dex20 molecules have a pronounced effect on the overall thermodynamic stability of BsCspB23. It could be shown that the presence of equally concentrated solutions containing PEG1, PEG8, PEG35, or Dex20 molecules impacts the overall thermodynamic stability of BsCspB quantitatively in the same manner regardless of differences in molecular weight and polarity. Selecting PEG1 and Dex20 as done here enables to perform highly-resolved NMR spectroscopic experiments on a protein, which is significantly larger than the used crowding molecules regarding PEG1 on the one hand or smaller regarding Dex20 on the other hand.
The application of dilute conditions is shown in a whereas the presence of macromolecular crowding agents is highlighted in b by using ellipsoids colored in gray. The PDB ID 1NMG has been used to illustrate the protein backbone of BsCspB representing the native state (right). One structure indicating the unfolded protein ensemble of BsCspB has been modeled (left).
Beyond that the present study aims to expand the current knowledge regarding the impact the crowded environment possesses on protein thermodynamic stability by mapping NMR derived data at single residue resolution to resonances originating from the main chain, and even from side chain methylene and methyl groups. This procedure enables to obtain a profound understanding of crowding effects on a protein at an atomic level.
Results
Unfolding of BsCspB under dilute conditions and in crowded environments monitoring 1H signals collectively
First, unfolding of BsCspB induced by an increasing concentration of urea has been monitored by applying one-dimensional 1H NMR spectroscopy (Supplementary Figs. 1a and 2). Under dilute conditions the midpoint of this transition, CM, has been determined to CM = 2.7 M urea (Fig. 2a, b) using aliphatic protons comprising BsCspB. This is in accordance to the result obtained by fluorescence spectroscopy, where a transition midpoint of CM = 2.5 M urea was found23. As expected, addition of c = 120 g/L PEG1 to the buffer leads to a significant increase in thermodynamic stability of BsCspB represented by a shift in CM of about 0.6 M urea resulting in CM = 3.3 M urea (Fig. 2a and Supplementary Table 1). This increase in thermodynamic stability of BsCspB is repeatedly observed for the addition of c = 120 g/L Dex20 (Fig. 2b and Supplementary Table 1). Notably, the gain in thermodynamic stability which has been observed here by using one-dimensional proton NMR spectroscopy is qualitatively in line with probing intrinsic fluorescence and circular dichroism following the unfolding of BsCspB23. Focusing on the change in free energy by unfolding, \({\Delta}G_{\text{N} \leftrightarrow \text{U}}^0\), we determined \({\Delta}G_{\text{N} \leftrightarrow \text{U}}^0\) = 8.4 kJ/mol under dilute conditions, compared to 9.7 kJ/mol in the presence of 120 g/L Dex20 and 9.8 kJ/mol in the presence of 120 g/L PEG1 (Supplementary Table 1). This difference in the change of free energy of unfolding, \({\Delta}{\Delta}G_{\text{N} \leftrightarrow \text{U}}^0\), of 1.3 kJ/mol for Dex20 and 1.4 kJ/mol for PEG1, resembles the range of change in \({\Delta}G_{\text{N} \leftrightarrow \text{U}}^0\) that would be anticipated from considerations of the excluded volume theory, a crowding model that has been intensively discussed in literature since Minton and Wilf coined the term of macromolecular crowding34. Following a pure excluded volume model as discussed by Christiansen et al.35, there is an expected gain in stability, ΔΔGN↔U, of about 3.5 kJ/mol by addition of 120 g/L Dex20, which is in the range of our results considering 3.6 kJ/mol for 100 g/L Dex20 measured by fluorescence spectroscopy23, and 1.3 kJ/mol for 120 g/L Dex20 measured by one-dimensional proton NMR spectroscopy. Comparing different spectroscopic techniques though, the absolute values of \({\Delta}{\Delta}G_{\text{N} \leftrightarrow \text{U}}^0\) slightly differ, which is inherent to the methodology as the applied techniques differ in the structural features they monitor. Here, we focus on the evaluation of atomically resolved NMR spectroscopic data monitoring the transition region of protein unfolding and do not include data extending to a concentration of 0 M urea, where the absolute value of \({\Delta}G_{\text{N} \leftrightarrow \text{U}}^0\) can be precisely estimated. Consequently, we choose the change of the midpoint of protein unfolding, ΔCM(urea), caused by the addition of crowding agents for the detailed discussion of our results.
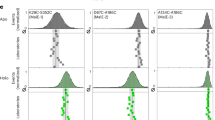
a, b Analysis of aliphatic protons representing the native state, fn, observed by one-dimensional 1H NMR spectroscopy in absence of MC (colored in gray), in presence of c = 120 g/L PEG1 (colored in blue, a), and c = 120 g/L Dex20 (colored in red, b). c, d Difference in the transition midpoint, ΔCM, of amide protons observed in two-dimensional 1H-15N HSQC NMR spectra comparing dilute conditions with the presence of c = 120 g/L PEG1 (colored in blue, c) or c = 120 g/L Dex20 (colored in red, d). Beta sheet regions according to PDB ID 1NMG are indicated by using a background colored in gray. The average of ΔCM is shown by the horizontal line (continuous mode) whereas the mean plus or minus one standard deviation of ΔCM is represented by two additional horizontal lines (dotted mode). The hydropathy score76 concerning BsCspB has been shown on top by using the online tool ProtScale available at the Bioinformatics Resource Portal ExPASy77.
Unfolding of BsCspB under dilute conditions and in crowded environments monitoring individual HN signals
In a next step, we have acquired two-dimensional heteronuclear 1H-15N HSQC NMR spectra of BsCspB dependent on the concentration of urea in order to significantly increase the spectral resolution (Supplementary Fig. 3). In accordance with the analysis done for the acquisition of one-dimensional 1H NMR data (Fig. 2a, b), folding-to-unfolding transitions of amide protons comprising BsCspB at a residue-by-residue level show a shift of CM to a higher concentration of urea, when comparing crowded with non-crowded conditions (Supplementary Fig. 4a–c). The analysis of individual folding-to-unfolding transitions of BsCspB was performed by using linear extrapolation of experimental data36,37 (see “Methods” section), since the approach for data analysis introduced by Santoro and Bolen38 is not indicated here as the baseline for native BsCspB is poorly defined in folding-to-unfolding transitions (Supplementary Fig. 4a–c). It has been shown before that poorly defined baselines potentially falsify the thermodynamic analysis39. Thus, analyzing the transition region ranging between curea = 2 M and curea = 4 M for individual amino acids comprising BsCspB omits efficiently the impact of the native baseline on the overall thermodynamic stability \({\Delta}G_{\text{N} \leftrightarrow \text{U}}^0\) and the folding cooperativity m (Supplementary Fig. 4d–f). In the present study, 53 amide proton folding-to-unfolding transitions characterizing BsCspB could be entirely monitored in the absence of crowding agents yielding a sufficient height of cross-peaks corresponding to nonoverlapping, nonproline residues that have been unambiguously assigned (Supplementary Fig. 1b). In the presence of c = 120 g/L PEG1 or Dex20, 48 out of 53 individual folding-to-unfolding transitions could be monitored. The analysis of these folding-to-unfolding transitions by linear extrapolation has been performed for individual residues taking the three data sets for dilute conditions, c = 120 g/L PEG1 and c = 120 g/L Dex20 into account sharing the folding cooperativity m as global residue specific parameter during the fitting procedure (Supplementary Fig. 4d–f). The sum of all individual folding-to-unfolding transitions observed for NH cross-peaks gives rise to a mean normalized transition for each of the three different conditions, which have been probed here (Supplementary Fig. 5 and Supplementary Table 1). Data analysis by linear extrapolation results in CM = 1.9 M for dilute conditions, \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\), and an increase by about ΔCM = 0.5 M or ΔCM = 0.6 M urea upon addition of c = 120 g/L Dex20 or PEG1, respectively (Supplementary Table 1). Again, this increase in thermodynamic stability of BsCspB by about ΔCM = 0.5 M urea in presence of MC is in accordance with folding-to-unfolding transitions based on the analysis of aliphatic protons applying one-dimensional 1H NMR spectroscopy (Fig. 2a, b). Note that the absolute value of CM differs by about 0.7 M urea when comparing \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) of one-dimensional 1H NMR analysis with mean \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) obtained for native state amide protons derived from the analysis of two-dimensional 1H-15N HSQC spectra. This deviation can be attributed to the fact that in one-dimensional NMR spectra the change in the unfolded ensemble of BsCspB is accounted for by integrating signals representing both native state as well as unfolded protein ensemble, respectively. In contrast, the analysis of folding-to-unfolding transitions of amide proton signals in two-dimensional HSQC NMR spectra focusses on the loss of the population of the native state only omitting information about the unfolded protein ensemble. In this respect, a shift of CM to higher values can be seen when analyzing the increase in the population of the unfolded protein ensemble induced by an increasing concentration of urea monitored by two-dimensional 1H-15N HSQC spectra (Supplementary Fig. 6). Quantitatively, the mean in CM is shifted to \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) = 4.2 M monitoring 16 transitions that could be unambiguously assigned and analyzed throughout the course of protein unfolding (Supplementary Fig. 1c). The mean transition midpoint for the folding-to-unfolding transition of BsCspB which is calculated by using folded as well as unfolded amide proton signals separately can be therefore specified with about 3.0 M urea, which is comparable to \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) = 2.7 M obtained for one-dimensional 1H NMR analysis. Analyzing the unfolded protein ensemble directly, there is also a stabilization effect due to the presence of MC agents evident. This thermodynamic stabilization can be determined to about ΔCM = 0.2 M and ΔCM = 0.7 M for c = 120 g/L PEG1 or Dex20, respectively (Supplementary Fig. 6e, f and Supplementary Table 1). The difference in stabilization induced by the two different MC agents as seen here is not reflected in the analysis of the native state amide proton folding-to-unfolding transitions. Moreover, the standard deviations of the mean transition midpoints derived from the unfolded protein ensemble reach an overlapping range. Therefore, we must conclude that this deviation seen in the unfolded ensemble is not a difference in the extent of stabilization of the native state exerted by the two MC agents, but rather reflects a limit of accuracy in the determination of the stabilization by evaluation of the amide proton signals of the unfolded ensemble. Similar observations have been illuminated by protein folding studies of e.g., the protein barstar in comparing NMR spectroscopic data with fluorescence detected transitions40,41. Note that an additionally performed concentration dependent analysis of the folding-to-unfolding transition of BsCspB conducted under dilute conditions does not show a significant difference in CM (Supplementary Fig. 7). In addition, another control has shown that using either height or volume of cross-peaks for the analysis of individual folding-to-unfolding transitions leads to similar results in ΔCM (Supplementary Fig. 8). Therefore, all cross-peaks acquired in two-dimensional heteronuclear NMR spectra in this study have been analyzed using peak height enabling a reliable readout of data even if cross-peaks are in close spectral proximity.
Monitoring folding-to-unfolding transitions based on NH cross-peaks further facilitates structurally resolved mapping of the individual thermodynamic stability at a residue-by-residue level under both dilute and MC conditions. First, mapping of individual folding-to-unfolding transition midpoints under dilute conditions to the primary sequence (Supplementary Fig. 9a) indicates that BsCspB does not possess regions, which are characterized by a significant local increase or decrease in thermodynamic stability compared to the average confirming a two-state folding character of this protein. There are only few residues possessing CMdilute values which do not lie within the range covered by the mean and one standard deviation. These are specifically Asn10, Ile18, Asp25, Phe27, Leu41, and Ala61 that possess a \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) > 2.2 M whereas Gly4, Lys5, Ala32, Ile33, Lys39, Gln45, Ala46, and Glu50 own \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) < 1.6 M (Supplementary Table 2). Mapping those residues on the structure of BsCspB (Supplementary Fig. 10a) underlines that there is no structural relation present that would connect residues of raised or lowered stability. Next, addition of MC agents increases the residue specific thermodynamic stability equally using either PEG1 or Dex20 (Fig. 2c, d). Again, only single, structurally separated residues show a moderate difference to the range of CM which is spanned by the mean and one standard deviation. Specifically, amino acids Gly4, Asn10, Ala32, Ile33, Leu41, and Ala61 mirror deviations larger than the mean plus one standard deviation in CM (Supplementary Fig. 10b, c). However, these residues have been already identified analyzing CMdilute. The general thermodynamic stabilization by about ΔCM = 0.5 M in the presence of either c = 120 g/L PEG1 or c = 120 g/L Dex20 is distributed over the entire primary sequence and is indifferent of surface or core regions, inherent properties of amino acids or the presence of secondary structural elements. This observation is in contrast to studies which have suggested that the effects of MC agents rely on the exposed surface area of the protein42,43, on a specific protein composition such as hydropathy or on the electrostatics of certain exposed residues44,45 due to direct interactions between MC agents and the protein under study. The results presented here underscore—at least for BsCspB—the general character of thermodynamic stabilization when PEG1 or Dex20 molecules are present at a concentration of about c = 120 g/L. This increase in thermodynamic stability is independent of direct binding or specific chemical interactions between the MC agent and defined sites in the protein, which can be seen from the secondary structure and hydropathy notation presented in Fig. 2c, d. To sum up, the detailed analysis of amide proton signals does not provide evidences for the presence of local hot spots responsible for thermodynamic stabilization of BsCspB and the extent of stabilization as induced by Dex20 and PEG1 is highly similar.
Unfolding of BsCspB under dilute conditions and in crowded environments monitoring individual CH, CH2, and CH3 signals
The analysis of folding-to-unfolding transitions of resonance signals comprising BsCspB has been further expanded by the acquisition of two-dimensional heteronuclear 1H-13C HSQC spectra in dependence on the concentration of urea (Supplementary Figs. 11 and 12). All folding-to-unfolding transitions of cross-peaks representing CH, CH2, and CH3 groups have been analyzed by linear extrapolation using both dilute and two MC conditions sharing the folding cooperativity m as global residue specific parameter during the fitting procedure (Supplementary Fig. 13a–f and Supplementary Tables 3–5). First, carbon bound protons comprising BsCspB show an overall thermodynamic stability of about \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) = 2.1 M urea which is in accordance with \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) analyzed for cross-peaks representing NH groups (Supplementary Table 1 and Supplementary Fig. 9a, c). Secondly, for almost all amino acids, the residue specific value of \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) determined for CH correlations lies within a range covered by one standard deviation of the mean. Thus, there are no structural regions present in BsCspB which possess a thermodynamic stability that is significantly increased or decreased (Fig. 3a, b). Consequently, the two state folding character of BsCspB30 monitored in equilibrium by using high-resolution NMR spectroscopy here is underlined by showing a well-defined midpoint of the folding-to-unfolding transitions for both individual NH and individual CH correlations. The few residues of BsCspB whose methine groups show a deviation in \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) from the mean value plus one standard deviation are Lys7, Ile18*, His29, Ile33*, Leu41*, Ala46, Ile51, Pro58, Val63, and Thr64. This list comprises three residues which have been identified before analyzing \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) using NH cross-peaks and have been labeled by using “*” here. Thus, the independent and profound analysis of both NH and CH cross-peaks comprising BsCspB dependent on the concentration of urea observed under dilute conditions does not illuminate a cluster of residues, which deviate in \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) regarding the overall thermodynamic stability. Therefore, we conclude that deviations observed in \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) are rather based on the very local flexibility and environment of nuclei comprising the amino acid under study.
a–f Difference in the transition midpoint, ΔCM, of CH protons (a, b), CH2 protons (c, d), and CH3 protons (e, f) observed in two-dimensional 1H-13C HSQC NMR spectra comparing dilute conditions with the presence of c = 120 g/L PEG1 (colored in blue) or c = 120 g/L Dex20 (colored in red). Beta sheet regions according to PDB ID 1NMG are indicated by using a background colored in gray. The mean of ΔCM is shown by the horizontal line (continuous mode) whereas the mean plus or minus one standard deviation of ΔCM is represented by two additional horizontal lines (dotted mode). The presence of more than a single CH, CH2 or CH3 group per residue is highlighted by applying dark and bright mode of red or blue color, respectively. For the assignment of cross-peaks in two-dimensional 1H-13C HSQC NMR spectra see Supplementary Fig. 8.
Next, 1H-13C HSQC spectra of BsCspB in presence of c = 120 g/L PEG1 or Dex20 have been acquired dependent on the concentration of urea. The addition of PEG1 or Dex20 is capable to increase the midpoint of folding-to-unfolding transitions of BsCspB probed by individual CH signals on average to the same degree as seen for the analysis of NH signals (Supplementary Fig. 9b, d). Specifically, the presence of c = 120 g/L PEG1 leads to an increase in CM of the folding-to-unfolding transition of BsCspB of about 0.3 M urea whereas the same concentration of Dex20 increases the transition midpoint by about 0.6 M urea (Supplementary Table 1). Analyzing the folding-to-unfolding transitions of CH2 groups comprising BsCspB leads quantitatively to the same result as seen for CH groups, illuminating an increase in CM of 0.2 M urea in presence of c = 120 g/L PEG1, whereas the presence of the same amount of Dex20 increases CM by about 0.6 M urea (Fig. 3c, d and Supplementary Table 4). The reliability of the experimental setup used for the determination of the thermodynamic stability of a protein at an atomic level outlined here is corroborated by the fact, that the increase in CM caused by a crowded environment seen for one proton present in the CH2 group equals the extent of stabilization seen for the second proton in the same CH2 group. Even though the absolute values might be divergent, the gain in stability is precisely conserved (Supplementary Table 4). Last but not least, individual folding-to-unfolding transitions of CH3 groups comprising BsCspB have been followed under dilute and MC conditions, respectively. In this regard, the midpoint CM reporting on the unfolding of BsCspB increases by about 0.5 M urea when c = 120 g/L PEG1 is present whereas c = 120 g/L Dex20 causes ΔCM = 0.7 M urea (Fig. 3e, f and Supplementary Table 5). Note again that an increase in CM observed for one methyl group comprising isoleucine, leucine or valine has been mirrored by the second methyl group present in the same residue (Supplementary Table 5).
Analyzing potential interaction of urea with crowding agents and with BsCspB being present in crowded environments
Further, we have analyzed the change in chemical shifts of NH cross-peaks present in 1H-15N HSQC NMR spectra to evaluate potential effects of urea on native BsCspB which is supplemented with MC agents. Applying dilute conditions, the addition of urea to BsCspB induces chemical shift perturbations (CSPs) of residues comprising predominantly the highly exposed loop ranging between Gly35 and Thr40 (Fig. 4a). The presence of crowding agents (c = 120 g/L PEG1 or Dex20) does neither qualitatively nor quantitatively alter the pattern of CSPs seen for BsCspB under dilute conditions when comparing the absence with the presence of c = 3 M urea (Fig. 4b, Supplementary Fig. 14). Therefore, we conclude that the presence of crowding agents does not structurally perturb the folding-to-unfolding transitions of individual NH cross-peaks comprising BsCspB, which have been induced by urea. In other words, the chemically induced unfolding in presence of PEG1 or Dex20 processes similar to the unfolding of BsCspB under dilute conditions. Consequently, it can be reasonably anticipated that also urea-induced folding-to-unfolding transitions of CH, CH2, and CH3 cross-peaks are not structurally perturbed when comparing dilute with crowding conditions. Furthermore, one-dimensional proton NMR spectroscopy has been applied to monitor the potential impact of urea on crowding agents used in this study. Neither perturbations in chemical shifts nor modifications in line shape of resonance signals comprising PEG1 and Dex20 could be observed comparing curea = 0 M with curea = 6 M (Supplementary Fig. 15). This result indicates an inertness of PEG1 and Dex20 regarding urea even if present at a high concentration.
a Changes in chemical shifts of 1H-15N cross-peaks comprising the native state of BsCspB in presence of c = 1.1 M urea (light gray), c = 2.25 M urea (dark gray), and c = 3.26 M urea (black) compared to the absence of urea applying dilute conditions. b Changes in chemical shifts of 1H-15N cross-peaks comprising the native state of BsCspB comparing c = 2.97 M with c = 0 M urea applying dilute conditions (colored in black), c = 2.94 M with c = 0 M urea using c = 120 g/L Dex20 (colored in red) and c = 3.08 M with c = 0 M urea using c = 120 g/L PEG1 (colored in blue). Beta sheet regions according to PDB ID 1NMG are indicated using a background colored in gray.
Following hydrodynamic dimensions of BsCspB upon unfolding
Finally, the experimental data set reporting on the folding-to-unfolding reaction of BsCspB in absence and presence of crowding agents has been complemented by applying diffusion NMR methodology. The determination of the diffusion coefficient of BsCspB dependent on the concentration of urea in absence and presence of c = 120 g/L Dex20 makes it possible to evaluate the folding-to-unfolding transition in terms of hydrodynamic dimensions, rH, taking native and unfolded conformations into account. First, applying dilute conditions cause a transition midpoint for the unfolding of BsCspB of \(C_{\mathrm{M}}^{{\mathrm{dilute}}}\) = 3.2 M urea taking averaged values of rH into account (Supplementary Fig. 16). Analyzing the diffusion of BsCspB in presence of c = 120 g/L Dex20 dependent on the concentration of urea increases CM by about 0.5 M urea to CM = 3.7 M (Supplementary Fig. 16). This increase in thermodynamic stability of BsCspB obtained by analyzing the diffusion properties once c = 120 g/L Dex20 is present reproduces the results independently obtained by applying one-dimensional (Fig. 2b and Supplementary Table 1) and two-dimensional NMR spectroscopy (Fig. 2d and Supplementary Table 1). Notably, the hydrodynamic dimensions representing native and unfolded BsCspB determined here possessing rH,nativ = 15.7 Å and rH,unf = 22.3 Å match very well with theoretically expected values of \(r_{{\mathrm{H,nativ}}}^{{\mathrm{th}}} = 16.1\,{\mathrm{{\AA}}}\) and \(r_{{\mathrm{H}},{\mathrm{unf}}}^{{\mathrm{th}}} = 24.3\,{\mathrm{{\AA}}}\) for a protein of this size46.
Discussion
Applying NMR spectroscopy enabled the elucidation of the folding-to-unfolding transition of BsCspB at highest resolution by observing cross-peaks belonging to backbone amide (Fig. 5), backbone carbon as well as side chain aliphatic protons dependent on the concentration of urea (Supplementary Fig. 17). Thereby, we have shown on all levels a cooperative effect in thermodynamic stabilization of BsCspB due to adding of PEG1 or Dex20. We will discuss how this thermodynamic stabilization can be understood and focus for this reason on excluded volume theory, potential enthalpic contributions arising from crowding agents, potential changes in water structure or the hydration shell as well as potential effects due to the combination of different cosolutes.
The increase in the transition midpoint, ΔCM, obtained for amide protons is mapped onto the three-dimensional structure of BsCspB (PDB ID: 1NMG) applying Dex20 (left) or PEG1 (right). The following color code has been used: 0.2 M ≤ ΔCM ≤ 0.3 M in light orange, 0.4 M ≤ ΔCM ≤ 0.6 M in orange and 0.7 M ≤ ΔCM ≤ 0.9 M in bright orange whereas residues colored in gray report on lack of experimental data. Numerical values for ΔCM obtained under these experimental conditions can be found in Fig. 2c, d and Supplementary Table 2.
Starting with Dex20, which can be modeled as a hard rod cosolute47, the thermodynamic stabilization of BsCspB can be qualitatively understood by the excluded volume effect47,48. Dex20 acting as an inert macromolecule reduces the space available for BsCspB in solution due to mutual exclusion and consequently BsCspB favors the compact, native state compared to the extended unfolded ensemble3. Note that this purely entropic origin of the MC effect can be potentially accompanied by additional enthalpic contributions due to chemical interactions. Thus, it has been discussed that enthalpic contributions may add on to or counteract the entropic stabilization of the native state27,47,49,50. Focusing on PEG, the effects which have been reported for this cosolvent are highly diverse. Thus it has been shown that PEG can act as a denaturant51 leading to thermal destabilization52 whereas cosolute exclusion leads to purely entropic stabilization of protein complexes53,54. Moreover, early as well as recently performed experimental studies have reported both stabilizing and destabilizing effects of PEG, respectively52,55. Shortly speaking, PEG has shown so far to exert a stabilizing effect due to steric repulsion that could in some cases be diminished or overridden by specific or unspecific hydrophobic interactions with protein side chains supposedly depending on the properties of the protein under investigation3,42,56. There have been attempts to classify proteins by their hydrophobicity or primary sequence in order to predict negative or positive enthalpic contributions based on chemical interactions with the protein43,57. Quantitatively, the steric repulsion or excluded volume effect of PEG can be modeled in the same way as for Dex20 since PEG conformations in solution have been described as low density random coils20 that can be sufficiently modeled as hard particles58.
Secondly, our results may indicate that in the context of MC and cosolute properties protein stabilization caused by a crowded environment might take place indirectly by changes in the water structure namely in the hydrogen bonding network of the solution59,60. Consequently, a perturbation of the pure water hydrogen bonding network might favor or disfavor the hydration of either the native or the unfolded protein species, if the thermodynamic stabilization of the protein under study gets influenced by changes in the water structure. In the present study, it is unlikely that the addition of PEG significantly changes the hydrogen bonding network composed of water and urea as PEG does not comprise a wealth of hydroxy groups being able to interact with urea. In the case of dextran though, the amino groups comprising urea may accept protons not only from water but also to a significant amount from Dex20 molecules which can be seen in the difference in pH value probing a pure water-urea solution (pH = 8.4 at curea = 6 M) compared to a water-urea—Dex20 solution (pH = 7.4 at curea = 6 M and cDex20 = 100 g/L). This decrease in pH value mirrors the reduced number of hydroxide ions which are set free once the amino groups comprising urea get protonated. In other words, urea potentially accepts protons from hydroxy groups in a solution which contains a polyol like Dex20. Thus, Dex20 is possibly involved in changes of the water structure and may influence the hydrogen bonding network. On the contrary, PEG does not perturb the water structure to this extent as the value of a solution containing water, urea and PEG1 (pH = 8.1 at curea = 6 M and cPEG1 = 100 g/L) does not significantly differ to a solution containing water and urea only (pH = 8.4 at curea = 6 M). Hence, we conclude that it is not very likely that PEG1 or Dex20 manipulate the structure of water and the hydration shell which is spanned around BsCspB, as they inherently possess different effects on water structure, but influence the thermodynamic stability of BsCspB to the same extent. We acknowledge that the extent of entropic repulsion—thus the size of the excluded volume caused by the presence of MC—depends on the thickness of the hydration shell. But for BsCspB being dissolved in Dex20 or PEG1, the impact on water structure and the hydration shell, if even present, has a negligible effect on the thermodynamic stability as our results show.
Moving on, it has been shown that destabilizing agents such as urea and stabilizing cosolutes can interact with each other in a way that diminishes individual contributions of the cosolutes61. Does the denaturation strength and the mechanism urea possesses regarding protein unfolding change once MC agents are present? Mechanistically, urea denatures proteins by changes in the hydration shell accompanied by a direct contact with the protein surface62,63. A class of small molecules, so-called “protective osmolytes”, stabilizes proteins against unfolding caused by urea. In this way, TMAO, inositol, sugars or polyols in general61,64,65 represent potent protective osmolytes that show stabilizing effects which mainly originate from interactions with the protein backbone66. Is Dex20 possibly capable of interactions, which are typically seen for small polyols? Even though dextran is widely accepted as an inert MC agent, it has also been shown that it is able to interact with proteins and thereby contributes a non-negligible enthalpic component42. For the model protein BsCspB studied here, the increase in thermodynamic stability seen for the protein backbone is the same for both crowding agents used in our experimental setup. Consequently, the stabilizing effects caused by the possibly affected hydrogen bonding network are either identical—which is unlikely as PEG1 and Dex20 differ significantly in polarity—or they are negligible. The present difference in polarity comparing PEG1 with Dex20 cannot be seen in an interaction with the protein backbone which gets inherently stabilized by hydrogen bonds. This also underlines that there is no direct interaction between amide protons and MC agents. One can assume that such an interaction would be dependent on the polarity of MC agents and would thus have to differ between PEG1 and Dex20. But, the folding-to-unfolding transitions which have been monitored here by applying one-dimensional NMR spectroscopy as well as by two-dimensional 1H-15N HSQCs report no differences in the thermodynamic stabilization of BsCspB, which has been induced by addition of PEG1 or Dex20, two crowding agents that differ in their hydropathicity. The analysis of carbon bound protons using two-dimensional 1H-13C HSQC spectra shows minor differences in the increase of the thermodynamic stability induced by the addition of Dex20 compared to PEG1, as the residue averaged mean ΔCM is 0.4 M urea for PEG1 but 0.6 M urea for Dex20 (Supplementary Table 1). This may indicate a potential influence of the difference in the polarity of Dex20 and PEG1, but could also result from possible destabilizing hydrophobic interactions of PEG1. To the best of our knowledge, any kind of destabilizing interactions having an impact on folding-to-unfolding transitions of CH groups must also be monitored for the atom, which is next to it which is in this case the amide proton. Since this destabilizing behavior is not present in 1H-15N HSQC derived transitions, we conclude that this minor difference in mean stabilization seen for carbon bound protons comparing the addition of Dex20 with the addition of PEG1 is negligible. However, what holds as a reason that an enthalpic contribution (if existent) for BsCspB can only be of negligible or equal extent for PEG1 and Dex20 as no hydrophobic interactions can be detected whereas a couple of different proteins have shown such a contribution67? This feature is potentially based on the low hydrophobicity BsCspB owns and its polyanionic net charge, estimated as −5.8 at pH 7. Note that it has been shown that the presence of charged residues increases the repulsive interaction of proteins with PEG, especially the presence of anionic charges enables to diminish hydrophobic interactions of proteins with PEG even more pronounced than cationic charges do57.
Taken all results together, we postulate that the increase in thermodynamic stability observed for BsCspB here is purely entropically driven. We propose that the effect of excluded volume is dominantly responsible for the increase in thermodynamic stability of BsCspB taking crowding agents covering a range of molecular weight between 1 to 20 kDa and concentrations of about 120 g/L into account. Interestingly, the pronounced difference in molecular weight of crowding agents used here does not change the magnitude of the stabilization seen for BsCspB. This demonstrates that a crowded environment which is composed of molecules possessing a molecular weight of about 1 kDa only procures an excluded volume effect regarding a protein of about 7 kDa in molecular weight (further discussion in the Supplementary Information).
Motivated by the lack of highly resolved experimental data from systematic bottom-up studies that are required to complement the theoretical discussion of MC50, this work contributes essential insights into the excluded volume induced stabilization at highest resolution possible. Probing the impact a crowded environment has on the thermodynamic stability of BsCspB at the level of 1H, 15N, and 13C one-dimensional and two-dimensional NMR spectroscopy enables to illuminate the dominance of entropic stabilization over enthalpic contributions at every level of resolution independent of the local site of detection. This finding narrows the gap between theoretical predictions and experimental proofs regarding the estimation of crowding effects on proteins.
It thereby underlines the high relevance of intracellular crowding, which affects essentially all cellular processes in vivo, and contributes to the evolution of a precise understanding of cellular protein action and interaction. Thus, a boost in following highly resolved in-cell studies enabling investigations performed also at atomic resolution can be proposed. Finally, we believe that due to the systematic experimental setup and comprehensive data analysis applied here a generalization of the results to proteins showing folding characteristics which are comparable to BsCspB is feasible.
Methods
Protein expression and purification
Protein expression and purification occurred as described30,68 with minor modifications. Buffer exchange and sample concentration was carried out using centrifugal filters (Amicon, Merck). Protein concentration was determined using an absorbance spectrometer; ε280nm = 5800 M−1 cm−1 30, aliquots were shock frozen and stored at −80 °C. Expression of 15N and 13C labeled BsCspB occurred in M9 minimal medium using 15N ammonium chloride (Cortecnet) and 13C glucose (Cambridge Isotope Laboratories) as nitrogen and carbon sources.
Sample preparation
Samples were prepared from stock solutions of Urea (ultra pure, MP), sodium cacodylate (Alfa Aesar) and the following crowding agents: Dextran T20 (Dex20) of a molecular weight of 20 kDa (Pharmacosmos), polyethylene glycol (PEG) of a molecular weight of 1 kDa (Roth). Dilute conditions refer to 20 mM sodium cacodylate at pH 7 and T = 298 K.
NMR measurements
1D 1H NMR spectra, 2D 1H-15N HSQC-TROSY and 2D 1H-13C HSQC spectra were recorded for a BsCspB solution (95% H2O/5% D2O, 20 mM sodium cacodylate/HCl, pH 7.0, containing either no crowder, 120 g/L Dex20, or 120 g/L PEG1) on a Bruker 600 MHz spectrometer with a cryogenic probe head. Folding-to-unfolding transitions of BsCspB have been monitored at cBsCspB = 1.09 mM (dilute and 120 g/L Dex20 condition), 0.8 mM (120 g/L PEG1 condition), 0.25 mM (diffusion measurement). An additional folding-to-unfolding transition of BsCspB has been monitored under dilute conditions at cBsCspB = 0.03 mM. Water suppression occurred using presaturation and a Watergate pulse sequence69. Pseudo 2D diffusion spectra of a 250 µM BsCspB solution (95% H2O/5% D2O, 20 mM sodium cacodylate/HCl; pH 7.0, with and without 120 g/L Dex20) were acquired on a Bruker 800 MHz spectrometer operated with a cryogenic probe head dependent on the applied gradient. Due to high molecular concentration of buffer molecules in crowded and denatured samples, further signal suppression was executed via presaturation of buffer specific frequencies.
Data analysis
1H NMR transition data for chemical unfolding was evaluated using integrals of 1D spectra in the aliphatic region, readout occurred with the software TopSpin (Bruker). Integration of signals typical for the native conformation occurred from 0.59 to 0.14 ppm and for the increase in the unfolded population, signals from 0.697 to 1.064 ppm were integrated. See ref. 23 for details. For data regression a thermodynamic two state model was employed as described by Szyperski et al.70. For chemical unfolding, qualitative analysis of two-dimensional 1H-15N HSQC spectra and 1H-13C HSQC spectra occurred via readout of peak height comprising all NH and all nonoverlapping CH, CH2, and CH3 cross-peaks. The standard deviation of ΔCM, ΔΔCM, has been calculated for n residues according to \({\Delta}{\Delta}C_{\mathrm{M}} = \sqrt {\frac{{\mathop {\sum }\nolimits_{i = 1}^{\mathrm{n}} (C_{\mathrm{M}}^{\mathrm{i}} - {\Delta}C_{\mathrm{M}})^2}}{{n - 1}}}\) in which \(C_{\mathrm{M}}^{\mathrm{i}}\) represents the midpoint of residue specific folding-to-unfolding transition.
For diffusion analysis, pseudo-2D spectra were processed using TopSpin. For baseline correction and integration Mestrenova 14 (Mestrelab Research) was used. Qualitative analysis of 2D spectra focused on the change in 1H and 15N chemical shifts. Both axes were referenced using TMSP and 15NH4Cl reference probes in the corresponding buffer / crowder concentration. All two-dimensional HSQC spectra were processed with the software NMR Pipe71, readout of individual peak height and chemical shift occurred using the software NMRView72. Perturbations of chemical shifts, CSPs, reporting on NH cross-peaks as acquired in two-dimensional 1H-15N HSQC NMR spectra were calculated according to Grzesiek et al.73. All folding-to-unfolding transitions were analyzed using the software Origin. For data regression of global chemical unfolding (detection of one-dimensional NMR spectra) the model of Santoro and Bolen was used38. For data regression of individual chemical unfolding transitions comprising single HN, CH, CH2, and CH3 transitions, linear extrapolation of the transition region was used37. NMR diffusion experiments were performed using a pulse sequence comprising a stimulated echo assisted by bipolar gradients74, G, employing a diffusion time, Δ, of 120 ms and gradient length, δ, of 5 ms along the z-axis. Gradients were calibrated as described in the ref. 75. Three different gradient strengths have been repeated three times each to estimate the experimental error. Integrals for proton signals, I, were determined for BsCspB in the spectral range between 9.55 and 6.50 ppm and used for calculation of the diffusion coefficient, D:
where γ is the gyromagnetic ratio of protons. Hydrodynamic radii, RH, were calculated using the respective diffusion coefficients of TMSP in order to account for the microviscosity of the respective sample. Microviscosities were calculated according to the Einstein–Stokes equation using TMSP with rH = 3.4 Å contained in the respective sample. The hydrodynamic radius for TMSP was estimated by using dioxane with a known rH = 2.12 Å as reported in46.
Linear extrapolation of experimental data
For data regression, heights of cross-peaks were normalized to the highest value found for the respective resonance and were reported as normalized values. The normalized values were set equal to fraction native of BsCspB present in solution and the derived ΔG values were plotted over denaturant concentration. Linear regression was performed in the transition region, including all data points in between 2 and 4 M urea, setting the cooperativity m as a residue specific parameter shared for the respective signal using the three conditions “dilute”, c = 120 g/L PEG1 and c = 120 g/L Dex20.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Cayley, S., Lewis, B. A., Guttman, H. J. & Record, M. T. Characterization of the cytoplasm of Escherichia coli K-12 as a function of external osmolarity: implications for protein-DNA interactions in vivo. J. Mol. Biol. 222, 281–300 (1991).
Zimmerman, S. B. & Trach, S. O. Estimation of macromolecule concentrations and excluded volume effects for the cytoplasm of Escherichia coli. J. Mol. Biol. 222, 599–620 (1991).
Zhou, H. X., Rivas, G. & Minton, A. P. Macromolecular crowding and confinement: biochemical, biophysical, and potential physiological consequences. Annu. Rev. Biophys. 37, 375–397 (2008).
Schnell, S. & Turner, T. E. Reaction kinetics in intracellular environments with macromolecular crowding: simulations and rate laws. Prog. Biophys. Mol. Biol. 85, 235–260 (2004).
Wang, Q., Zhuravleva, A. & Gierasch, L. M. Exploring weak, transient protein–protein interactions in crowded in vivo environments by in-cell nuclear magnetic resonance spectroscopy. Biochemistry 50, 9225–9236 (2011).
Han, J. & Herzfeld, J. Macromolecular diffusion in crowded solutions. Biophys. J. 65, 1155–1161 (1993).
Ellis, R. J. & Minton, A. P. Cell biology: join the crowd. Nature 425, 27–28 (2003).
Sarkar, M., Li, C. & Pielak, G. J. Soft interactions and crowding. Biophys. Rev. 5, 187–194 (2013).
Smith, A. E., Zhou, L. Z., Gorensek, A. H., Senske, M. & Pielak, G. J. In-cell thermodynamics and a new role for protein surfaces. Proc. Natl Acad. Sci. USA 113, 1725–1730 (2016).
Dedmon, M. M., Patel, C. N., Young, G. B. & Pielak, G. J. FlgM gains structure in living cells. Proc. Natl Acad. Sci. USA 99, 12681–12684 (2002).
Luh, L. M. et al. Molecular crowding drives active Pin1 into nonspecific complexes with endogenous proteins prior to substrate recognition. J. Am. Chem. Soc. 135, 13796–13803 (2013).
Chen, E. et al. Effects of macromolecular crowding on burst phase kinetics of cytochrome c folding. Biochemistry 51, 9836–9845 (2012).
Echeverria, C. & Kapral, R. Molecular crowding and protein enzymatic dynamics. Phys. Chem. Chem. Phys. 14, 6755–6763 (2012).
Bismuto, E. et al. Effect of molecular confinement on internal enzyme dynamics: Frequency domain fluorometry and molecular dynamics simulation studies. Biopolymers 67, 85–95 (2002).
Akabayov, B., Akabayov, S. R., Lee, S.-J., Wagner, G. & Richardson, C. C. Impact of macromolecular crowding on DNA replication. Nat. Commun. 4, 1615 (2013).
Dupuis, N. F., Holmstrom, E. D. & Nesbitt, D. J. Molecular-crowding effects on single-molecule RNA folding/unfolding thermodynamics and kinetics. Proc. Natl Acad. Sci. USA 111, 8464–8469 (2014).
Desai, R., Kilburn, D., Lee, H.-T. & Woodson, S. A. Increased ribozyme activity in crowded solutions. J. Biol. Chem. 289, 2972–2977 (2014).
Kulp, D. T. & Herzfeld, J. Crowding-induced organization of cytoskeletal elements. III. Spontaneous bundling and sorting of self-assembled filaments with different flexibilities. Biophys. Chem. 57, 93–102 (1995).
Breydo, L. et al. The crowd you’re in with: effects of different types of crowding agents on protein aggregation. Biochim. Biophys. Acta 1844, 346–357 (2014).
Shtilerman, M. D., Ding, T. T. & Lansbury, P. T. Molecular crowding accelerates fibrillization of α-synuclein: could an increase in the cytoplasmic protein concentration induce Parkinson’s disease? Biochemistry 41, 3855–3860 (2002).
Luo, X.-D., Kong, F.-L., Dang, H.-B., Chen, J. & Liang, Y. Macromolecular crowding favors the fibrillization of β2-microglobulin by accelerating the nucleation step and inhibiting fibril disassembly. Biochim. Biophys. Acta 1864, 1609–1619 (2016).
Stagg, L., Christiansen, A. & Wittung-Stafshede, P. Macromolecular crowding tunes folding landscape of parallel alpha/beta protein, apoflavodoxin. J. Am. Chem. Soc. 133, 646–648 (2011).
Köhn, B. & Kovermann, M. Macromolecular crowding tunes protein stability by manipulating solvent accessibility. ChemBioChem 20, 759–763 (2019).
Soranno, A. et al. Single-molecule spectroscopy reveals polymer effects of disordered proteins in crowded environments. Proc. Natl Acad. Sci. USA 111, 4874–4879 (2014).
Mikaelsson, T., Ådén, J., Johansson, L. B. Å. & Wittung-Stafshede, P. Direct observation of protein unfolded state compaction in the presence of macromolecular crowding. Biophys. J. 104, 694–704 (2013).
Danielsson, J. et al. Thermodynamics of protein destabilization in live cells. Proc. Natl Acad. Sci. USA 112, 12402–12407 (2015).
Rivas, G. & Minton, A. P. Macromolecular crowding in vitro, in vivo, and in between. Trends Biochem. Sci. 41, 970–981 (2016).
Schindelin, H., Marahiel, M. A. & Heinemann, U. Universal nucleic acid-binding domain revealed by crystal structure of the B. subtilis major cold-shock protein. Nature 364, 164–168 (1993).
Schnuchel, A. et al. Structure in solution of the major cold-shock protein from Bacillus subtilis. Nature 364, 169–171 (1993).
Schindler, T., Herrler, M., Marahiel, M. A. & Schmid, F. X. Extremely rapid protein folding in the absence of intermediates. Nat. Struct. Biol. 2, 663–673 (1995).
Schindler, T. & Schmid, F. X. Thermodynamic properties of an extremely rapid protein folding reaction. Biochemistry 35, 16833–16842 (1996).
Jacob, M. et al. Microsecond folding of the cold shock protein measured by a pressure-jump technique. Biochemistry 38, 2882–2891 (1999).
Jacob, M., Schindler, T., Balbach, J. & Schmid, F. X. Diffusion control in an elementary protein folding reaction. Proc. Natl Acad. Sci. USA 94, 5622–5627 (1997).
Minton, A. P. & Wilf, J. Effect of macromolecular crowding upon the structure and function of an enzyme: glyceraldehyde-3-phosphate dehydrogenase. Biochemistry 20, 4821–4826 (1981).
Christiansen, A., Wang, Q., Cheung, M. S. & Wittung-Stafshede, P. Effects of macromolecular crowding agents on protein folding in vitro and in silico. Biophys. Rev. 5, 137–145 (2013).
Pace, C. N. Methods in Enzymology Vol. 131, 266–280 (Academic Press, Cambridge, 1986).
Greene, R. F. & Pace, C. N. Urea and guanidine hydrochloride denaturation of ribonuclease, lysozyme, α-chymotrypsin, and β-lactoglobulin. J. Biol. Chem. 249, 5388–5393 (1974).
Santoro, M. M. & Bolen, D. W. Unfolding free energy changes determined by the linear extrapolation method. 1. Unfolding of phenylmethanesulfonyl alpha-chymotrypsin using different denaturants. Biochemistry 27, 8063–8068 (1988).
Haupt, C., Weininger, U., Kovermann, M. & Balbach, J. Local and coupled thermodynamic stability of the two-domain and bifunctional enzyme SlyD from Escherichia coli. Biochemistry 50, 7321–7329 (2011).
Hofmann, H. et al. Fast amide proton exchange reveals close relation between native-state dynamics and unfolding kinetics. J. Am. Chem. Soc. 131, 140–146 (2009).
Hofmann, H., Golbik, R. P., Ott, M., Hübner, C. G. & Ulbrich-Hofmann, R. Coulomb forces control the density of the collapsed unfolded State of Barstar. J. Mol. Biol. 376, 597–605 (2008).
Jiao, M., Li, H.-T., Chen, J., Minton, A. P. & Liang, Y. Attractive protein-polymer interactions markedly alter the effect of macromolecular crowding on protein association equilibria. Biophys. J. 99, 914–923 (2010).
Shkel, I. A., Knowles, D. B. & Record, M. T. Separating chemical and excluded volume interactions of polyethylene glycols with native proteins: comparison with PEG effects on DNA helix formation. Biopolymers 103, 517–527 (2015).
Lee, J. C. & Lee, L. L. Preferential solvent interactions between proteins and polyethylene glycols. J. Biol. Chem. 256, 625–631 (1981).
Crowley, P. B., Keith, B. & Jimmy, M. NMR spectroscopy reveals cytochrome c-poly(ethylene glycol) interactions. ChemBioChem 9, 685–688 (2008).
Wilkins, D. K. et al. Hydrodynamic radii of native and denatured proteins measured by pulse field gradient NMR techniques. Biochemistry 38, 16424–16431 (1999).
Minton, A. P. Models for excluded volume interaction between an unfolded protein and rigid macromolecular cosolutes: macromolecular crowding and protein stability revisited. Biophys. J. 88, 971–985 (2005).
Minton, A. P. Effect of a concentrated inert macromolecular cosolute on the stability of a globular protein with respect to denaturation by heat and by chaotropes: a statistical-thermodynamic model. Biophys. J. 78, 101–109 (2000).
Minton, A. P. Excluded volume as a determinant of macromolecular structure and reactivity. Biopolymers 20, 2093–2120 (1981).
Rivas, G. & Minton, A. P. Toward an understanding of biochemical equilibria within living cells. Biophys. Rev. 10, 241–253 (2018).
Arakawa, T., Bhat, R. & Timasheff, S. N. Why preferential hydration does not always stabilize the native structure of globular proteins. Biochemistry 29, 1924–1931 (1990).
Lee, L. L. Y. & Lee, J. C. Thermal stability of proteins in the presence of poly(ethylene glycols). Biochemistry 26, 7813–7819 (1987).
Reddy, M. K., Weitzel, S. E., Daube, S. S., Jarvis, T. C. & von Hippel, P. H. Methods in Enzymology Vol. 262 466–476 (Academic Press, Cambridge, 1995).
Jarvis, T. C., Ring, D. M., Daube, S. S. & von Hippel, P. H. “Macromolecular crowding”: thermodynamic consequences for protein–protein interactions within the T4 DNA replication complex. J. Biol. Chem. 265, 15160–15167 (1990).
Tellam, R. L., Sculley, M. J., Nichol, L. W. & Wills, P. R. The influence of poly(ethylene glycol) 6000 on the properties of skeletal-muscle actin. Biochem. J. 213, 651–659 (1983).
Mukherjee, S. K., Gautam, S., Biswas, S., Kundu, J. & Chowdhury, P. K. Do macromolecular crowding agents exert only an excluded volume effect? A protein solvation study. J. Phys. Chem. B 119, 14145–14156 (2015).
Knowles, D. B. et al. Chemical interactions of polyethylene glycols (PEGs) and glycerol with protein functional groups: applications to effects of PEG and glycerol on protein processes. Biochemistry 54, 3528–3542 (2015).
Zimmerman, S. B. & Minton, A. P. Macromolecular crowding: biochemical, biophysical, and physiological consequences. Annu. Rev. Biophys. Biomol. Struct. 22, 27–65 (1993).
Eggers, D. K. & Valentine, J. S. Crowding and hydration effects on protein conformation: a study with sol-gel encapsulated proteins. J. Mol. Biol. 314, 911–922 (2001).
Panuszko, A., Bruździak, P., Kaczkowska, E. & Stangret, J. General mechanism of Osmolytes’ influence on protein stability irrespective of the type of osmolyte cosolvent. J. Phys. Chem. B 120, 11159–11169 (2016).
Holthauzen, L. M. F. & Bolen, D. W. Mixed osmolytes: the degree to which one osmolyte affects the protein stabilizing ability of another. Protein Sci. 16, 293–298 (2007).
Bennion, B. J. & Daggett, V. The molecular basis for the chemical denaturation of proteins by urea. Proc. Natl Acad. Sci. USA 100, 5142 (2003).
Lim, W. K., Rösgen, J. & Englander, S. W. Urea, but not guanidinium, destabilizes proteins by forming hydrogen bonds to the peptide group. Proc. Natl Acad. Sci. USA 106, 2595 (2009).
Yancey, P. H., Clark, M. E., Hand, S. C., Bowlus, R. D. & Somero, G. N. Living with water stress: evolution of osmolyte systems. Science 217, 1214 (1982).
Manning, M. C., Chou, D. K., Murphy, B. M., Payne, R. W. & Katayama, D. S. Stability of protein pharmaceuticals: an update. Pharm. Res. 27, 544–575 (2010).
Liu, Y. & Bolen, D. W. The peptide backbone plays a dominant role in protein stabilization by naturally occurring osmolytes. Biochemistry 34, 12884–12891 (1995).
Wang, Y., Sarkar, M., Smith, A. E., Krois, A. S. & Pielak, G. J. Macromolecular crowding and protein stability. J. Am. Chem. Soc. 134, 16614–16618 (2012).
Sachs, R., Max, K. E., Heinemann, U. & Balbach, J. RNA single strands bind to a conserved surface of the major cold shock protein in crystals and solution. RNA 18, 65–76 (2012).
Piotto, M., Saudek, V. & Sklenář, V. Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J. Biomol. NMR 2, 661–665 (1992).
Szyperski, T., Mills, J. L., Perl, D. & Balbach, J. Combined NMR-observation of cold denaturation in supercooled water and heat denaturation enables accurate measurement of ΔC p of protein unfolding. Eur. Biophys. J. 35, 363–366 (2006).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Kirby, N. I., DeRose, E. F., London, R. E. & Mueller, G. A. NvAssign: protein NMR spectral assignment with NMRView. Bioinformatics 20, 1201–1203 (2004).
Grzesiek, S., Stahl, S. J., Wingfield, P. T. & Bax, A. The CD4 determinant for downregulation by HIV-1 nef directly binds to nef. mapping of the nef binding surface by NMR. Biochemistry 35, 10256–10261 (1996).
Jones, J. A., Wilkins, D. K., Smith, L. J. & Dobson, C. M. Characterisation of protein unfolding by NMR diffusion measurements. J. Biomol. NMR 10, 199–203 (1997).
Berger, S. & Braun, S. 200 and more NMR Experiments. A Practical Course. (Wiley, Hoboken, 2004).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982).
Gasteiger, E. et al. Protein Identification and Analysis Tools on the ExPASy Server. 571–607 (Humana Press, Totowa, 2005).
Acknowledgements
We thank Holger Hilbert, Julian Schuck, and Dr. Oleksandra Kukharenko for support at the initial state of this project and Professor Magnus Wolf-Watz for fruitful discussion. Financial support from the Young Scholar Fund of the University of Konstanz is gratefully acknowledged.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
B.K. and M.K. designed research; B.K. performed research; B.K. and M.K. analyzed data; B.K. and M.K. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Köhn, B., Kovermann, M. All atom insights into the impact of crowded environments on protein stability by NMR spectroscopy. Nat Commun 11, 5760 (2020). https://doi.org/10.1038/s41467-020-19616-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-19616-w
This article is cited by
-
Monitoring protein unfolding transitions by NMR-spectroscopy
Journal of Biomolecular NMR (2022)
-
Impact of crowded environments on binding between protein and single-stranded DNA
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.