Abstract
Congenital nephrotic syndrome (CNS) is a heterogeneous group of disorders presenting with massive proteinuria within the first 3 months of life almost inevitably leading to end-stage kidney disease. The Work Group for the European Reference Network for Kidney Diseases (ERKNet) and the European Society for Pediatric Nephrology (ESPN) has developed consensus statement on genetic aspects of CNS diagnosis and management. The presented expert opinion recommends genetic diagnostics as the key diagnostic test to be ordered already during the initial evaluation of the patient, discusses which phenotyping workup should be performed and presents known genotype–phenotype correlations.
Similar content being viewed by others
Introduction
Congenital nephrotic syndrome (CNS) is a heterogeneous group of disorders presenting in utero or during the first 3 months of life with marked edema and massive proteinuria [1]. In the vast majority of cases, CNS is a primary glomerular disorder due to genetic defects; occasionally it can however be caused by congenital infections or alloimmune maternal disease [2].
Originally, the disorder has been referred to as the Finnish-type nephrotic syndrome due to its high incidence in Finland (1:8.000 live births), with two NPHS1 founder mutations (i.e., Fin-major and Fin-minor) in most cases [3,4,5]. However, over the years several other genes have been implicated in the CNS etiology when mutated, both in isolated cases or in the the less common syndromic forms of the disease [6,7,8].
NPHS1 (encoding nephrin) or NPHS2 (encoding podocin) biallelic pathogenic variants are the most common causes of the disease in patients of both European and other descents. Among syndromic forms, WT1-associated glomerulopathy is the most frequent, followed by Pierson syndrome caused by LAMB2 biallelic pathogenic variants [6,7,8,9,10,11,12,13,14]. The exact data on the prevalence of CNS in various populations are limited. With the advent of massive-parallel sequencing, rapid-WES usage in daily clinical practice in particular, it is anticipated that the information on the associated mutational spectrum and the incidence of various types of hereditary CNS will be soon achieved. Public lists of disease-causing variants are available online (www.ncbi.nlm.nih.gov/clinvar/; www.lovd.nl; www.hgmd.cf.ac.uk). Also, online case consultations, including discussion of variants in question, are available via the European Rare Kidney Disease Reference Network (http://www.cpms.erknet.org).
CNS management is very challenging due to high morbidity and mortality rates [15]. Patients with CNS are prone to severe complications related to hypoproteinemia such as recurrent infection, thrombosis, and impaired growth and most progress to end-stage kidney disease (ESKD) within a few years [1, 5, 16].
In 2018, the European Reference Network for Kidney Diseases (ERKNet) and the European Society for Pediatric Nephrology (ESPN) have jointly established a Work Group to develop recommendations for clinical diagnostics, management and treatment of CNS. This resulted in clinical recommendations for CNS (Boyer et al. in prep.) and the present consensus statement on genetic aspects of CNS diagnosis and management.
Because evidence is frequently missing or inadequate in CNS management, this article is an expert opinion paper rather than a clinical practice guideline. Detailed methodology, including formulation of PICO (Patient or Population covered, Intervention, Comparator, Outcome) questions, approaches to literature search and three-step revision process is presented in the parallel clinical paper (Boyer et al. in prep.).
Content list—questions addressed in the presented recommendations:
-
1.
Which phenotyping workup should be performed?
-
2.
What is the preferred time-point for genetic diagnostics?
-
3.
What is the appropriate genetic testing approach?
-
4.
Is there a role for karyotyping?
-
5.
What kind of samples are needed for genetic testing?
-
6.
Is there a role for prenatal diagnosis/genetic counseling?
-
7.
What are the phenotype/genotype correlations?
-
8.
How to manage syndromic forms?
-
9.
What is the inheritance pattern of a hereditary CNS?
-
10.
Parents as kidney donators.
-
1.
Which phenotyping workup should be performed?
In addition to standard clinical care phenotypic workup in infants with CNS (Boyer et al. in prep.) we recommend additional evaluation with the aim of identifying extrarenal signs and symptoms suggestive of an underlying genetic disease.
Clinical examination should be performed to identify the possible extrarenal manifestations of a hereditary form of CNS. Table 1 summarizes possible phenotypic manifestations of syndromic forms of CNS.
Particular attention should be paid to neurological examination, including brain imaging in selected cases. Abnormal cerebral gyration or cerebellar atrophy in Galloway–Mowat syndrome (GAMOS) [17,18,19]; cerebral and cerebellar atrophy and stroke-like lesions in CoQ10 nephropathies [20,21,22]; ventriculomegaly in patients with CRB2-glomerulopathy [23] and subcortical changes in patients with SGPL1 biallelic pathogenic variants [24] have been described. Ophthalmological examination and hearing evaluation should also be performed in all cases.
-
2.
What is the preferred time-point for genetic diagnostics?
We recommend genetic testing as a first choice diagnostic test in every CNS patient. It should be performed as part of the initial patient evaluation and should be considered prior to a renal biopsy.
Once the clinical suspicion of CNS is raised, genetic testing should be initiated. Genetic testing in CNS is a fast, noninvasive and reliable one-time diagnostic measure. Prompt genetic testing has profound effects on clinical decision making as it reduces the time for diagnosis in infants during hospital stay and may enhance the cost-effectiveness of clinical management [25, 26]. Establishing the genetic diagnosis is essential for proper patient management, facilitates the anticipation, and/or swift identification of extrarenal manifestations, informs recurrence risk counseling, and may lead to the identification of genetic defects that may represent phenocopies of nephrotic syndrome [27]. As a general rule, in genetic forms of CNS the use of immunosuppressive drugs should be avoided; instead, appropriate fluid management and proteinuria-lowering RAAS (renin–angiotensin–aldosterone system) blockade at postneonatal age should be introduced promptly to stabilize the patient’s condition and slow renal failure progression (see Boyer et al. in prep. for further details). Particular issues around therapeutic decision making in syndromic forms are discussed further in Question 8. These include but are not limited to (1) being aware of the risk of urogenital tumors and meticulous monitoring of them in WT1-associated disease; (2) promptly initiating CoQ10 supplementation in CoQ10-related nephropathies; (3) planning appropriate therapy in GAMOS individuals or subjects with CRB2 or SGPL1-associated disorders; (4) managing endocrine manifestations in individuals with SGPL1 or WT1-related syndromes.
-
3.
What is the appropriate genetic testing approach?
We recommend one- or two-step genetic testing depending on the presenting phenotype and financial and/or technical restrictions related to the diagnostics.
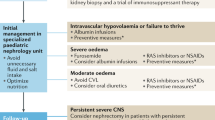
Genetic screening strategies might vary depending on country specific peculiarities including availability and access to genetic testing and reimbursement policy of national healthcare systems (Fig. 1).
In CNS, basic genetic screening of NPHS1, NPHS2, WT1, and LAMB2 genes will uncover underlying genetic abnormalities in >80% of cases. Several other less commonly mutated genes account for an additional ~5% of diagnoses [6, 7, 9, 10, 28, 29]. Due to the wide range of phenotypic variability and genetic heterogeneity of the disease [6, 7, 9, 14, 28, 29] a comprehensive genetic screening comprising all SRNS-related genes is recommended as the first tier method using next generation sequencing technology with either an expanded gene panel or whole exome sequencing (WES). Targeted gene panels have higher depth of coverage of the genes related to a specific phenotype, which allow a higher diagnostic rate. Also, gene panels will not yield incidental findings in genes unrelated with CNS. However, the number of genes that can be examined using a gene panel approach is limited and the covered list of genes should be regularly updated as new genes are continuously identified. Conversely, WES allows detecting variants not only in an established list of known, but also in novel disease-causing genes [30]. However WES has also some important limitations. Complex rearrangements, small copy-number variants and changes within regulatory fragments (promotor, introns, enhancers, silencers etc) might be missed by standard protocols. In particular, WES may have suboptimal coverage of some clinically relevant regions: fragments with high GC content, the mitochondrial genome or duplicated regions (pseudogenes) [31]. These limitations may, to some extent, be overcome by use of hybridization-based custom target enrichment for NGS gene panels.
For patients who present a clear phenotype associated with a syndromic form of CNS, testing the particular related gene can be performed as the first step followed by comprehensive genetic testing if no pathogenic variant is detected. In populations where founder mutations or specific genetic abnormalities have already been well described such as CNS of the Finnish type (NPHS1 Fin-major and Fin-minor recessive pathogenic variants), the genetic screening strategy may be modified considering population specific features.
Informed consent should be obtained before initiating genetic studies. Informed consent forms should clearly describe the methods that will be applied, and the interpretation and handling of results and incidental findings that would have clinical and psychosocial impacts, for instance male-to-female sex-reversal related to WT1 dominant pathogenic variants.
-
4.
Is there a role for karyotyping?
We recommend karyotype testing to be performed in each CNS patient with ambiguous genitalia and in each phenotypic female with a causative WT1 mutation.
Establishing chromosomal gender of a patient with CNS with ambiguous genitalia and/or a WT1 dominant pathogenic variant is necessary for further proper management, endocrine and oncological follow-up in particular [32, 33].
For karyotype testing, a heparin blood sample should be obtained according to local genetic laboratory recommendations.
-
5.
What kind of samples are needed for genetic testing?
We recommend performing genetic testing on whole blood (EDTA) or umbilical cord tissue/blood sample.
A blood sample for genetic testing should be obtained as soon as possible as it informs further clinical management of a CNS patient. However, in those patients who have individual blood-sampling limitations or less than 3 months from a blood transfusion, buccal swabs or newborn dried blood spots may also be used [34, 35].
-
6.
Is there a role for prenatal diagnosis/genetic counseling?
We recommend prompt genetic counseling in families with a past history or prenatal signs of CNS.
In families with a past history of CNS, recurrence risk counseling by a clinical geneticist/clinical counsellour should be promptly provided. The decision regarding preimplantation genetic diagnosis and prenatal genetic testing should be discussed in light of the local financial, social, and legal settings [36].
-
7.
What are the phenotype/genotype correlations?
We suggest to take the following phenotype–genotype information into account for genetic counseling:
NPHS1
The majority of patients in whom CNS manifests in the first week of life will harbor biallelic pathogenic variants in the nephrin gene [10, 14]. The frequency of the disorder is significantly higher in Finland with two NPHS1 founder mutations (i.e., Fin-major and Fin-minor) in most cases [3,4,5]. Renal pathology does not exclusively appear as “Finnish type” NS in CNS caused by recessive NPHS1 pathogenic variants [37]. There is no difference between truncating NPHS1 variants (i.e., nonsense, frameshift, and splice site) and missense NPHS1 variants in terms of age at diagnosis, proportion of age at ESKD or death, and proportion of patients who achieved at least transient albumin withdrawal [38]. A subset of patients with CNS and a milder clinical course were described associated with the NM_004646.3: c.3478C>T(p.(Arg1160Ter)) variant [10, 39].
PLCE1
Patients with PLCE1 (NM_016341.3) splice site recessive pathogenic variants have an earlier age of onset than patients with C-terminal truncating variants (after amino acid residue 1000) or missense variants [6]. PLCE1 pathogenic variants are mostly associated with diffuse mesangial sclerosis histopathology [6]. Anecdotally, incomplete penetrance has been reported [40, 41].
NPHS2
NPHS2 (NM_014625.3) biallelic pathogenic variants are the main genetic cause of CNS beginning >1 month after birth [10]. There is no correlation regarding age of onset for truncating versus missense variants [7]. The European founder mutation c.413G>A (p.Arg138Gln) is frequently detected in those CNS patients who do not have NPHS1-related disease [10]. Individuals homozygous for the c.413G>A (p.Arg138Gln) variant present NS at a median age of 2 months and generally progress to ESKD after the age of 5 years (median 79 months; range 24–159 months) [10]. Individuals with the c.779T>A (p.(Val260Glu)) variant have an earlier age at onset of nephrotic syndrome and progress more rapidly to ESKD when compared to subjects with the c.413G>A (p.Arg138Gln) variant [10].
The disease-associated allele c.686G>A (p.Arg229Gln) has variant dependent pathogenicity [42, 43]. It is considered pathogenic only when located in trans to another recessive pathogenic variant in exon 7 or 8 of NPHS2. Such compound heterozygosity generally causes late onset nephrotic syndrome (after the median age of 17 months) [6, 10, 42, 43]. However, a few subjects developed nephrotic syndrome in the first month of life; yet, progressed slowly to ESKD [10].
LAMB2
In LAMB2 (NM_002292.3), N-terminal truncating recessive pathogenic variants tend to manifest before 2 months of life, whereas C-terminal truncating variants later. Missense variants and small in frame deletions have a higher mean age at onset of renal disease and lack of neurologic abnormalities [6]. Diffuse mesangial sclerosis has been identified in 61% of individuals with LAMB2-associated NS [44].
WT1
WT1 dominant pathogenic variants are associated with a wide range of clinical phenotypes that are clearly associated with the type and location of the causative WT1 variant. More than 90% of the deleterious variants reside in the hot spot region (exons 8 and 9 and their intronic junctions) [6, 32, 45, 46].
Classically, individuals with WT1 dominant pathogenic variants have been subclassified as having Denys–Drash or Frasier syndrome, however these two syndromes may overlap phenotypically to a certain extent. A number of patients present with a milder phenotype that cannot be easily classified as one of these syndromes [32, 45, 46].
Missense substitutions affecting DNA-binding residues are associated with diffuse mesangial sclerosis, early-onset steroid-resistant nephrotic syndrome and rapid progression to ESKD. Truncating variants confer the highest risk of Wilms tumor (~80%) and congenital anomalies of kidney and urinary tact (~25%) but are typically associated with late-onset steroid-resistant nephrotic syndrome [32, 45, 46]. Intronic (KTS) variants usually present as isolated SRNS with the histological picture of FSGS and slow progression to ESKD. Patients with isolated SRNS are genotypic and phenotypic females.
Male-to-female sex reversal (46,XY complete gonadal dysgenesis) occurs exclusively in individuals with intronic KTS dominant pathogenic variants and exonic variants. Urogenital abnormalities have also been described in patients with all types of WT1 deleterious variants [32, 45, 46].
CoQ10-associated CNS
Primary CoQ10 deficiencies that stem from autosomal recessive pathogenic variants in genes involving endogenous CoQ10 biosynthesis can cause nephropathies that are collectively referred to as CoQ10-associated nephropathies. Of them, COQ2, COQ6, and PDSS2 biallelic variants have been shown to be related to CNS. As CoQ10 is essential for mitochondrial electron transport, many organs can be affected, therefore multisystemic involvement is a cardinal feature of these disorders. There are no established genotype–phenotype correlations, potentially due to the small number of patients in the literature. In addition to progressive nephropathy, COQ2 pathogenic recessive variants typically manifest with signs of progressive encephalopathy (including ataxia, generalized amyotrophy, retinitis pigmentosa, bilateral sensorineural deafness, hypotonia, and psychomotor delay), hypertrophic cardiomyopathy, as well as diabetes [20, 21]. COQ6 recessive pathogenic variants are associated with severe infantile onset progressive SRNS resulting in end-stage renal failure and sensorineural hearing loss, central nervous system involvement, congenital heart disease, and motor retardation [13, 47]. PDSS2 recessive pathogenic variants are associated with Leigh syndrome, a progressive and severe neurodegenerative disorder, which may become evident within the first months of life and may result in early death. Affected individuals usually show global developmental delay or developmental regression, hypotonia, ataxia, dystonia, and ophthalmologic abnormalities such as nystagmus or optic atrophy. Basal ganglia and/or brainstem or brain imaging show T2-weighted hyperintensities [22].
It most commonly presents as a progressive and severe neurodegenerative disorder with onset within the first months or years of life, and may result in early death. Affected individuals usually show global developmental delay or developmental regression, hypotonia, ataxia, dystonia, and ophthalmologic abnormalities, such as nystagmus or optic atrophy. The neurologic features are associated with the classic findings of T2-weighted hyperintensities in the basal ganglia and/or brainstem on brain imaging.
Galloway–Mowat syndrome
GAMOS is a phenotypically heterogeneous disorder characterized by neurodevelopmental defects combined with podocytopathy. Individuals with GAMOS may not present overt dysmorphic features, however pre- or postnatal microcephaly should be considered its hallmark. Central nervous system abnormalities result in severely delayed psychomotor development and propensity to seizures. Additional clinical features include skeletal anomalies and various degrees of growth retardation [18].
There is high inter- and intrafamilial variability concerning renal involvement with regard to age at onset and type of kidney disease; some individuals may not even have renal disease [17, 48].
Recessive pathogenic variants in a number of genes, including WDR73, LAGE3, OSGEP, TP53RK, and TPRKB encoding subunits of the KEOPS complex [17, 18, 48]; WDR4 encoding an enzyme required for a specific posttranscriptional modification of tRNA [49]; and NUP107 and NUP133 encoding nuclear pore complex proteins [50] have been implicated in the pathogenesis of the disorder. This significant genetic heterogeneity and extreme rarity of the disorder with less than 50 patients described so far hamper precise genotype–phenotype analyses.
SGPL1
Biallelic pathogenic variants in SGPL1 result in a podocytopathy and primary adrenal insufficiency. Additional features include ichthyosis, acanthosis, immunodeficiency manifesting as lymphopenia, and recurrent bacterial infections. About half had variable neurologic abnormalities including ataxia, cognitive decline, loss of motor skills, impaired speech, and sensorineural hearing loss. There was a significant variability of the extra-adrenal and extrarenal features in the ~30 individuals reported so far [24, 51, 52].
CRB2
Biallelic pathogenic variants in CRB2 result in a glomerulopathy with additional systemic features in a minority of cases [53]. The more severe disease manifests already prenatally with renal corticomedullary cysts and structural abnormalities of the central nervous system, ventriculomegaly in particular. In some, additional defects of the radius or postaxial polydactyly is also noted. Most affected pregnancies have been terminated [23]. No genotype–phenotype data exist due to the extreme rarity of the disorder.
Histopathology
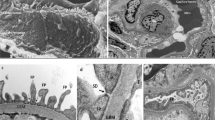
Diffuse mesangial sclerosis is associated with dominant pathogenic variants in WT1 (23.1%) and biallelic pathogenic variants in PLCE1 (17.8%), LAMB2 (13.6%), and NPHS1 (4.9%) [6]. CoQ10-associated nephropathies may be associated with focal and segmental glomerulosclerosis, focal mesangial sclerosis, and collapsing glomerulopathy [20, 54]. Increased and dysmorphic mitochondria in podocytes in electron microscopy are highly suggestive for CoQ10-associated nephropathy. Yet, absence of mitochondrial abnormalities does not exclude CoQ10-associated nephropathy diagnosis.
-
8.
How to manage syndromic forms?
In addition to standard clinical management of CNS described in detail elsewhere (Boyer et al. in prep.) we recommend that all syndromic CNS patients should be managed by a multidisciplinary team as described below.
NPHS2
We recommend cardiac evaluation in patients with NPHS2 biallelic pathogenic variants as cardiac anomalies have been shown in 89% of patients with the c.412C>T(p.(Arg138*)) variant [55]. Despite a few case reports describing partial or complete remission after immunosuppressive treatment, by principle patients with biallelic NPHS2 pathogenic variants respond neither to standard steroid treatment nor to intensified immunosuppressive treatment [7, 56, 57]. Therefore, we recommend not to use immunosuppresive regimens but to use RAAS blockade in such patients.
WT1
We recommend individuals with WT1-glomerulopathy to be evaluated for urogenital malformations. Oncological surveillance for Wilms tumor and gonadoblastoma should be applied. Subjects with exonic variants should be monitored for Wilms tumor with abdominal US performed every 3 months until the age of 7 years [58]. After reaching ESKD, bilateral nephrectomy should be considered to prevent the development of Wilms tumor, in particular in individuals carrying truncating variants [32, 59]. In subjects with a 46,XY karyotype and a female phenotype (i.e., complete gonadal dysgenesis), we recommend bilateral gonadectomy due to increased gonadoblastoma risk [32].
WT1 patients should be managed by a multidisciplinary team comprising a clinical geneticist, pediatric oncologist for Wilms tumor and gonadoblastoma surveillance, pediatric endocrinologist, pediatric surgeon, and psychologist in cases of disorders of sex development.
LAMB2
We recommend detailed ophthalmological examination in children with LAMB2 biallelic pathogenic variants, even though individuals with missense pathogenic variants may display variable phenotypes ranging from a milder variant of Pierson syndrome to an isolated CNS. Surviving children may have neurodevelopmental deficits and blindness [44].
Individuals with LAMB2-associated glomerulopathy need to be managed by multidisciplinary team composed of pediatric optalmologist, clinical geneticist, pediatric neurologist and rehabilitation team.
PLCE1
We recommend PLCE1-related nephropathy to be included in the differential diagnosis in subjects with congenital/infantile nephrotic syndrome associated with diffuse mesangial sclerosis in particular. In general, most individuals are resistant to any immunosuppressive therapy but some selected cases may respond to steroid or cyclosporine treatment [40, 60].
CoQ10-related mitochondropathies (COQ2, COQ6, and PDSS2 genes)
We recommend performing complete and repeated screening for extrarenal manifestations in patients with biallelic pathogenic variants in COQ2, COQ6, and PDSS2 or presenting phenotype suggestive of CoQ10-related glomerulopathy (hearing deficit, encephalopathy, seizures, ataxia, hypotonia, motor/intelectual disability, elevated lactate levels, and diabetes).
For diagnosis we recommend massive-parallel sequencing of the corresponding genes in the first place. In case of a non-informative genetic result, muscle or skin biopsies may be needed for measuring mitochondrial enzyme activity [61]. Renal biopsy with electron microscopy allows quantitative and qualitative analysis of mitochondria [62].
Individuals with CoQ10-related mitochondropathy need to be managed with a multidisciplinary approach including a pediatric ophthalmologist, audiologist, clinical geneticist, pediatric neurologist, rehabilitation team, and in case of diabetes, pediatric endocrinologist.
We suggest treating the individuals with biallelic pathogenic variants in COQ2, COQ6, and PDSS2 or presenting phenotype suggestive of CoQ10-related glomerulopathy with oral CoQ10 as early as possible.
Few case reports suggest that patients with defective variants in CoQ10 biosynthesis genes treated early with oral CoQ10 have improved outcome [20, 54, 61, 63, 64]. Individuals who respond to treatment exhibit an improvement in proteinuria and sometimes in neuromuscular complaints, however, refractory encephalopathy and seizures have been reported in subjects who had a beneficial effect of CoQ10 treatment on their kidney disease [21]. As the total number of reported patients is low, the exact dose regimen to improve or reverse glomerular damage is unknown. The initial CoQ10 dose applied in several reported cases is 15–30 mg/kg/day divided in three administrations, which might be increased to 50 mg/kg/day [47, 54, 63]. Leukocyte CoQ10 levels can be normal in these patients, and seem not to be helpful for monitoring therapy [54].
Galloway–Mowat syndrome
We recommend that individuals with GAMOS are managed by a multidisciplinary team including a pediatric nephrologist, pediatric neurologist, clinical geneticist, and physiotherapist. For older children psychotherapeutic, psychological and speech therapy services should be offered. Palliative care may also be considered depending on the severity of the disease.
SGPL1-associated CNS
We recommend patients with SGPL1-glomerulopathy to be carefully monitored for adrenal insufficiency. Individuals should be managed by a multidisciplinary team including a pediatric nephrologist, pediatric endocrinologist, pediatric neurologist, clinical geneticist, and physiotherapist. For older children psychotherapeutic, psychological, and speech therapy services should be offered. Palliative care may also be considered depending on the severity of the disease.
-
9.
What is the inheritance pattern of a hereditary CNS?
We recommend that each individual with confirmed hereditary CNS will have a genetic consult performed to address the issues of recurrence risk in the family.
The majority of forms of hereditary CNS are inherited in an autosomal recessive manner. That implies a 25% risk of recurrence in subsequent pregancies.
The exceptions are:
-
WT1—inherited in autosomal dominant manner; the recurrence risk depends whether the genetic defect is familial or occurred de novo (50% vs. <1% due to gonadal mosaicism respectively).
-
LAGE3—inherited in an X-linked recessive manner; the reccurence risk is 0% for female and 50% for male siblings.
-
10.
Parents as kidney donors.
We recommend that parents undergo genetic counseling prior to kidney donation for their child who has CNS with a confirmed genetic diagnosis.
The majority of forms of hereditary CNS are inherited in an autosomal recessive (AR) manner (see #9 for exceptions). This implies that both parents are obligate carriers of one of the defective variants. An extremely rare omission to this rule would be a de novo mutation or misattributed paternity. Carriers of a heterozygous variant in an AR gene can be kidney donors.
However, it cannot be excluded that the parents actually are also homozygotes/compound heterozygotes for the pathogenic variant(s) in the gene associated with a hereditary nephrotic syndrome. NPHS2-related SRNS and WT1-associated glomerulopathy are the two forms of the disease with the most significant intra- and inter-family variability, with age-dependent penetrance reflecting defective variant type [32, 43]. Moreover, incomplete penetrance has been described in families with a WT1 pathogenic variant [65].
References
Holmberg C, Antikainen M, Rönnholm K, Ala Houhala M, Jalanko H. Management of congenital nephrotic syndrome of the Finnish type. Pediatr Nephrol. 1995;9:87–93.
Büscher AK, Weber S. Educational paper: the podocytopathies. Eur J Pediatr. 2012;171:1151–60.
Ahvenainen EK, Hallman N, Hjelt L. Nephrotic syndrome in newborn and young infants. Ann Paediatr Fenn. 1956;2:227–41.
Lenkkeri U, Männikkö M, McCready P, Lamerdin J, Gribouval O, Niaudet PM, et al. Structure of the gene for congenital nephrotic syndrome of the finnish type (NPHS1) and characterization of mutations. Am J Hum Genet. 1999;64:51–61.
Jalanko H. Congenital nephrotic syndrome. Pediatr Nephrol. 2009;24:2121–8.
Sadowski CE, Lovric S, Ashraf S, Pabst WL, Gee HY, Kohl S, et al. A single-gene cause in 29.5% of cases of steroid-resistant nephrotic syndrome. J Am Soc Nephrol. 2015;26:1279–89.
Trautmann A, Bodria M, Ozaltin F, Gheisari A, Melk A, Azocar M, et al. Spectrum of steroid-resistant and congenital nephrotic syndrome in children: the PodoNet registry cohort. Clin J Am Soc Nephrol. 2015;10:592–600.
Vivante A, Hildebrandt F. Exploring the genetic basis of early-onset chronic kidney disease. Nat Rev Nephrol. 2016;12:133–46.
Hinkes BG, Mucha B, Vlangos CN, Gbadegesin R, Liu J, Hasselbacher K, et al. Nephrotic syndrome in the first year of life: two thirds of cases are caused by mutations in 4 genes (NPHS1, NPHS2, WT1, and LAMB2). Pediatrics. 2007;119:e907–19.
Machuca E, Benoit G, Nevo F, Tête MJ, Gribouval O, Pawtowski A, et al. Genotype-phenotype correlations in non-Finnish congenital nephrotic syndrome. J Am Soc Nephrol. 2010;21:1209–17.
Sako M, Nakanishi K, Obana M, Yata N, Hoshii S, Takahashi S, et al. Analysis of NPHS1, NPHS2, ACTN4, and WT1 in Japanese patients with congenital nephrotic syndrome. Kidney Int. 2005;67:1248–55.
Lee JH, Han KH, Lee H, Kang HG, Moon KC, Shin JI, et al. Genetic basis of congenital and infantile nephrotic syndromes. Am J Kidney Dis. 2011;58:1042–3.
Li GM, Cao Q, Shen Q, Sun L, Zhai YH, Liu HM, et al. Gene mutation analysis in 12 Chinese children with congenital nephrotic syndrome. BMC Nephrol 2018;19:382.
Cil O, Besbas N, Duzova A, Topaloglu R, Peco-Antić A, Korkmaz E, et al. Genetic abnormalities and prognosis in patients with congenital and infantile nephrotic syndrome. Pediatr Nephrol. 2015;30:1279–87.
Hölttä T, Bonthuis M, Van Stralen KJ, Bjerre A, Topaloglu R, Ozaltin F, et al. Timing of renal replacement therapy does not influence survival and growth in children with congenital nephrotic syndrome caused by mutations in NPHS1: data from the ESPN/ERA-EDTA Registry. Pediatr Nephrol. 2016;31:2317–25.
Huttunen NP. Congenital nephrotic syndrome of Finnish type. Study of 75 patients. Arch Dis Child. 1976;51:344–8.
Colin E, Huynh Cong E, Mollet G, Guichet A, Gribouval O, Arrondel C, et al. Loss-of-function mutations in WDR73 are responsible for microcephaly and steroid-resistant nephrotic syndrome: Galloway–Mowat syndrome. Am J Hum Genet. 2014;95:637–48.
Braun DA, Rao J, Mollet G, Schapiro D, Daugeron MC, Tan W, et al. Mutations in KEOPS-complex genes cause nephrotic syndrome with primary microcephaly. Nat Genet. 2017;49:1529–38.
Rosti RO, Dikoglu E, Zaki MS, Abdel-Salam G, Makhseed N, Sese JC, et al. Extending the mutation spectrum for Galloway-Mowat syndrome to include homozygous missense mutations in the WDR73 gene. Am J Med Genet A. 2016;170A:992–8.
Rötig A, Appelkvist EL, Geromel V, Chretien D, Kadhom N, Edery P, et al. Quinone-responsive multiple respiratory-chain dysfunction due to widespread coenzyme Q10 deficiency. Lancet. 2000;356:391–5.
Eroglu FK, Ozaltin F, Gönç N, Nalçacıoğlu H, Özçakar ZB, Yalnızoğlu D, et al. Response to early coenzyme Q10 supplementation is not sustained in CoQ10 deficiency caused by CoQ2 mutation. Pediatr Neurol. 2018;88:71–4.
López LC, Schuelke M, Quinzii CM, Kanki T, Rodenburg RJ, Naini A, et al. Leigh syndrome with nephropathy and CoQ10 deficiency due to decaprenyl diphosphate synthase subunit 2 (PDSS2) mutations. Am J Hum Genet. 2006;79:1125–9.
Slavotinek A, Kaylor J, Pierce H, Cahr M, DeWard SJ, Schneidman-Duhovny D, et al. CRB2 mutations produce a phenotype resembling congenital nephrosis, Finnish type, with cerebral ventriculomegaly and raised alpha-fetoprotein. Am J Hum Genet. 2015;96:162–9.
Prasad R, Hadjidemetriou I, Maharaj A, Meimaridou E, Buonocore F, Saleem M, et al. Sphingosine-1-phosphate lyase mutations cause primary adrenal insufficiency and steroid-resistant nephrotic syndrome. J Clin Investig. 2017;127:942–53.
Petrikin JE, Willig LK, Smith LD, Kingsmore SF. Rapid whole genome sequencing and precision neonatology. Semin Perinatol. 2015;39:623–31.
Borghesi A, Mencarelli MA, Memo L, Ferrero GB, Bartuli A, Genuardi M, et al. Intersociety policy statement on the use of whole-exome sequencing in the critically ill newborn infant. Italian J Pediatr. 2017;43:100.
Warejko JK, Tan W, Daga A, Schapiro D, Lawson JA, Shril S, et al. Whole exome sequencing of patients with steroid-resistant nephrotic syndrome. Clin J Am Soc Nephrol. 2018;13:53–62.
McCarthy HJ, Bierzynska A, Wherlock M, Ognjanovic M, Kerecuk L, Hegde S, et al. Simultaneous sequencing of 24 genes associated with steroid-resistant nephrotic syndrome. Clin J Am Soc Nephrol. 2013;8:637–48.
Wang F, Zhang Y, Mao J, Yu Z, Yi Z, Yu L, et al. Spectrum of mutations in Chinese children with steroid-resistant nephrotic syndrome. Pediatr Nephrol. 2017;32:1181–92.
Ng SB, Turner EH, Robertson PD, Flygare SD, Bigham AW, Lee C, et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature. 2009;461:272–6.
Groopman EE, Rasouly HM, Gharavi AG. Genomic medicine for kidney disease. Nat Rev Nephrol. 2018;14:83–104.
Lipska BS, Ranchin B, Iatropoulos P, Gellermann J, Melk A, Ozaltin F, et al. Genotype–phenotype associations in WT1 glomerulopathy. Kidney Int. 2014;85:1169–78.
Ukarapong S, Berkovitz G, McElreavey K, Bashamboo A, Bao Y. Early recognition of gonadal dysgenesis in congenital nephrotic syndrome. Clin Nephrol. 2016;86:341–4.
Paynter RA, Skibola DR, Skibola CF, Buffler PA, Wiemels JL, Smith MT. Accuracy of multiplexed Illumina platform-based single-nucleotide polymorphism genotyping compared between genomic and whole genome amplified DNA collected from multiple sources. Cancer Epidemiol Biomark Prev. 2006;15:2533–6.
Rajatileka S, Luyt K, El-Bokle M, Williams M, Kemp H, Molnár E, et al. Isolation of human genomic DNA for genetic analysis from premature neonates: a comparison between newborn dried blood spots, whole blood and umbilical cord tissue. BMC Genet. 2013;14:105.
van El CG, Cornel MC, ESHG Public and Professional Policy Committee. Genetic testing and common disorders in a public health framework. Eur J Hum Genet. 2011;19:377–81.
Schoeb DS, Chernin G, Heeringa SF, Matejas V, Held S, Vega-Warner V, et al. Nineteen novel NPHS1 mutations in a worldwide cohort of patients with congenital nephrotic syndrome (CNS). Nephrol Dial Transplant. 2010;25:2970–6.
Bérody S, Heidet L, Gribouval O, Harambat J, Niaudet P, Baudouin V, et al. Treatment and outcome of congenital nephrotic syndrome. Nephrol Dial Transplant. 2019;34:458–67.
Koziell A, Grech V, Hussain S, Lee G, Lenkkeri U, Tryggvason K, et al. Genotype/phenotype correlations of NPHS1 and NPHS2 mutations in nephrotic syndrome advocate a functional inter-relationship in glomerular filtration. Hum Mol Genet. 2002;11:379–88.
Boyer O, Benoit G, Gribouval O, Nevo F, Pawtowski A, Bilge I, et al. Mutational analysis of the PLCE1 gene in steroid resistant nephrotic syndrome. J Med Genet. 2010;47:445–52.
Gilbert RD, Turner CL, Gibson J, Bass PS, Haq MR, Cross E, et al. Mutations in phospholipase C epsilon 1 are not sufficient to cause diffuse mesangial sclerosis. Kidney Int. 2009;75:415–9.
Tory K, Menyhárd DK, Woerner S, Nevo F, Gribouval O, Kerti A, et al. Mutation-dependent recessive inheritance of NPHS2-associated steroid-resistant nephrotic syndrome. Nat Genet. 2014;46:299–304.
Mikó Á, K Menyhárd D, Kaposi A, Antignac C, Tory K. The mutation-dependent pathogenicity of NPHS2p.R229Q: A guide for clinical assessment. Hum Mutat. 2018;39:1854–60.
Matejas V, Hinkes B, Alkandari F, Al-Gazali L, Annexstad E, Aytac MB, et al. Mutations in the human laminin beta2 (LAMB2) gene and the associated phenotypic spectrum. Hum Mutat. 2010;31:992–1002.
Lehnhardt A, Karnatz C, Ahlenstiel-Grunow T, Benz K, Benz MR, Budde K, et al. Clinical and molecular characterization of patients with heterozygous mutations in Wilms tumor suppressor gene 1. Clin J Am Soc Nephrol. 2015;10:825–31.
Ahn YH, Park EJ, Kang HG, Kim SH, Cho HY, Shin JI, et al. Genotype-phenotype analysis of pediatric patients with WT1 glomerulopathy. Pediatr Nephrol. 2017;32:81–9.
Heeringa SF, Chernin G, Chaki M, Zhou W, Sloan AJ, Ji Z, et al. COQ6 mutations in human patients produce nephrotic syndrome with sensorineural deafness. J Clin Investig. 2011;121:2013–24.
Vodopiutz J, Seidl R, Prayer D, Khan MI, Mayr JA, Streubel B, et al. WDR73 mutations cause infantile neurodegeneration and variable glomerular kidney disease. Hum Mutat. 2015;36:1021–8.
Braun DA, Shril S, Sinha A, Schneider R, Tan W, Ashraf S, et al. Mutations in WDR4 as a new cause of Galloway–Mowat syndrome. Am J Med Genet A. 2018;176:2460–5.
Braun DA, Lovric S, Schapiro D, Schneider R, Marquez J, Asif M, et al. Mutations in multiple components of the nuclear pore complex cause nephrotic syndrome. J Clin Investig. 2018;128:4313–28.
Lovric S, Goncalves S, Gee HY, Oskouian B, Srinivas H, Choi WI, et al. Mutations in sphingosine-1-phosphate lyase cause nephrosis with ichthyosis and adrenal insufficiency. J Clin Investig. 2017;127:912–28.
Janecke AR, Xu R, Steichen-Gersdorf E, Waldegger S, Entenmann A, Giner T, et al. Deficiency of the sphingosine-1-phosphate lyase SGPL1 is associated with congenital nephrotic syndrome and congenital adrenal calcifications. Hum Mutat. 2017;38:365–72.
Ebarasi L, Ashraf S, Bierzynska A, Gee HY, McCarthy HJ, Lovric S, et al. Defects of CRB2 cause steroid-resistant nephrotic syndrome. Am J Hum Genet. 2015;96:153–61.
Starr MC, Chang IJ, Finn LS, Sun A, Larson AA, Goebel J, et al. COQ2 nephropathy: a treatable cause of nephrotic syndrome in children. Pediatr Nephrol. 2018;33:1257–61.
Frishberg Y, Feinstein S, Rinat C, Becker-Cohen R, Lerer I, Raas-Rothschild A, et al. The heart of children with steroid-resistant nephrotic syndrome: is it all podocin? J Am Soc Nephrol. 2006;17:227–31.
Ruf RG, Lichtenberger A, Karle SM, Haas JP, Anacleto FE, Schultheiss M, et al. Patients with mutations in NPHS2 (podocin) do not respond to standard steroid treatment of nephrotic syndrome. J Am Soc Nephrol. 2004;15:722–32.
Trautmann A, Schnaidt S, Lipska-Ziętkiewicz BS, Bodria M, Ozaltin F, Emma F, et al. Long-term outcome of steroid-resistant nephrotic syndrome in children. J Am Soc Nephrol. 2017;28:3055–65.
Lipska-Ziętkiewicz BS. WT1 Disorder. 2020 Apr 30. In: Adam MP, Ardinger HH, Pagon RA, et al., editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2020. Available from: https://www.ncbi.nlm.nih.gov/books/NBK556455/
Ruf RG, Schultheiss M, Lichtenberger A, Karle SM, Zalewski I, Mucha B, et al. Prevalence of WT1 mutations in a large cohort of patients with steroid-resistant and steroid-sensitive nephrotic syndrome. Kidney Int. 2004;66:564–70.
Hinkes B, Wiggins RC, Gbadegesin R, Vlangos CN, Seelow D, Nürnberg G, et al. Positional cloning uncovers mutations in PLCE1 responsible for a nephrotic syndrome variant that may be reversible. Nat Genet. 2006;38:1397–405.
Emma F, Montini G, Parikh SM, Salviati L. Mitochondrial dysfunction in inherited renal disease and acute kidney injury. Nat Rev Nephrol. 2016;12:267–80.
Parikh S, Goldstein A, Koenig MK, Scaglia F, Enns GM, Saneto R, et al. Diagnosis and management of mitochondrial disease: a consensus statement from the Mitochondrial Medicine Society. Genet Med. 2015;17:689–701.
Diomedi-Camassei F, Di Giandomenico S, Santorelli FM, Caridi G, Piemonte F, Montini G, et al. COQ2 nephropathy: a newly described inherited mitochondriopathy with primary renal involvement. J Am Soc Nephrol. 2007;18:2773–80.
Stańczyk M, Bałasz-Chmielewska I, Lipska-Ziętkiewicz B, Tkaczyk M. CoQ10-related sustained remission of proteinuria in a child with COQ6 glomerulopathy-a case report. Pediatr Nephrol. 2018;33:2383–7.
Fencl F, Malina M, Stará V, Zieg J, Mixová D, Seeman T, et al. Discordant expression of a new WT1 gene mutation in a family with monozygotic twins presenting with congenital nephrotic syndrome. Eur J Pediatr. 2012;171:121–4.
Acknowledgements
The authors gratefully acknowledge the support by the hereditary glomerulopathy working group of ERKNet, the European Reference Network for Rare Kidney Diseases, and the working group for inherited kidney disease of ESPN, the ESPN. We are indebted to Tanja Wlodkowski and Giulia Bassanese from the ERKNet Central Office, who performed the systematic literature search and evidence review that formed the basis of this document. Open Access Funding provided by Projekt DEAL.
Funding
The development of this consensus statement was made possible by a 5000 € grant from ERKNet, the European Reference Network for Rare Kidney Diseases. ERKNet is co-funded by the European Union within the framework of the Third Health Program “ERN-2016—Framework Partnership Agreement 2017–2021”. The funder took no influence on the contents of the document.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lipska-Ziętkiewicz, B.S., Ozaltin, F., Hölttä, T. et al. Genetic aspects of congenital nephrotic syndrome: a consensus statement from the ERKNet–ESPN inherited glomerulopathy working group. Eur J Hum Genet 28, 1368–1378 (2020). https://doi.org/10.1038/s41431-020-0642-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-020-0642-8
This article is cited by
-
Native nephrectomy in advanced pediatric kidney disease: indications, timing, and surgical approaches
Pediatric Nephrology (2024)
-
WT1-related disorders: more than Denys-Drash syndrome
Pediatric Nephrology (2024)
-
Genetic nephrotic syndrome associated with disturbed function of glomerular slit membrane and podocyte cytoskeleton in children
Clinical and Experimental Nephrology (2023)
-
IPNA clinical practice recommendations for the diagnosis and management of children with steroid-sensitive nephrotic syndrome
Pediatric Nephrology (2023)
-
Fanconi syndrome, nephrotic-range proteinuria, and hypoalbuminemia in a newborn—Occam’s razor or Hickam’s dictum? Answers
Pediatric Nephrology (2022)