Abstract
Dysfunctional mitochondria have been shown to enhance cancer cell proliferation, reduce apoptosis, and increase chemoresistance. Chemoresistance develops in nearly all patients with colorectal cancer, leading to a decrease in the therapeutic efficacies of anticancer agents. However, the effect of dynamin-related protein 1 (Drp1)-mediated mitochondrial fission on chemoresistance in colorectal cancer is unclear. Here, we found that the release of high-mobility group box 1 protein (HMGB1) in conditioned medium from dying cells by chemotherapeutic drugs and resistant cells, which triggered Drp1 phosphorylation via its receptor for advanced glycation end product (RAGE). RAGE signals ERK1/2 activation to phosphorylate Drp1 at residue S616 triggerring autophagy for chemoresistance and regrowth in the surviving cancer cells. Abolishment of Drp1 phosphorylation by HMGB1 inhibitor and RAGE blocker significantly enhance sensitivity to the chemotherapeutic treatment by suppressing autophagy. Furthermore, patients with high phospho-Drp1Ser616 are associated with high risk on developing tumor relapse, poor 5-year disease-free survival (DFS) and 5-year overall survival (OS) after neoadjuvant chemoradiotherapy (neoCRT) treatment in locally advanced rectal cancer (LARC). Moreover, patients with RAGE-G82S polymorphism (rs2070600) are associated with high phospho-Drp1Ser616 within tumor microenvironment. These findings suggest that the release of HMGB1 from dying cancer cells enhances chemoresistance and regrowth via RAGE-mediated ERK/Drp1 phosphorylation.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is one of the leading cause of death worldwide1, accounting for approximately 9.7% of all cancer cases and approximately 8.5% of cancer deaths. Currently, the major chemotherapy drugs for the treatment of CRC include oxaliplatin (OXP), 5-fluorouracil (5-FU) and irinotecan (CPT-11). However, a considerable proportion of CRC patients develop local recurrence and distant metastasis within 5 years after surgical treatment. The overall survival of CRC patients remains poor with a median of 12–18 months, and the response to chemotherapy is only 50%2. Moreover, most metastatic CRC patients develop resistance to OXP as the cancer progresses within 8 months3.
Mitochondria are organelles that provide most of the energy to nearly all cells because of their synthesis of ATP via oxidative phosphorylation. Mitochondria are key organelles for cellular homeostasis, which is regulated by the following dynamic networks: fusion and fission4. Fusion is mediated by Mitofusin-1 and Mitofusin-2 (Mfn1 and Mfn2) and optic atrophy 1 (Opa1) proteins located at the outer and inner mitochondrial membranes, respectively. Fission is mediated by dynamin-related protein 1 (Drp1), which is a cytosolic protein that is recruited to the surface of mitochondria during activation, leading to mitochondria fragmentation. Cumulative evidence has revealed that unbalanced mitochondrial dynamics dysregulate key cellular processes, potentially contributing to tumorigenesis5,6, including lung, bladder, breast, and colon cancers7,8,9,10. An imbalance in the expression of Drp1/Mfn is associated with excess mitochondrial fission and impaired mitochondrial fusion, which are important for the cell cycle progression5,7. Recently, the mitogen-activated protein kinase (MAPK) pathway has been shown to result in an increased mitochondrial fragmentation and promote tumor growth and chemoresistance via the phosphorylation of the mitochondrial protein Drp1 at serine 616 by extracellular signal-regulated kinase 2(ERK 2) in several cancers11,12,13,14. However, to date, the molecular mechanisms by which the dysregulated mitochondrial dynamics contribute to cancer cell survival remain unclear.
High-mobility group box 1 protein (HMGB1) is a highly conserved nuclear protein that functions as a chromatin-binding factor that bends DNA and promotes access to transcriptional protein assemblies on specific targets. In addition to its intra-nuclear role, HMGB1 functions as an extracellular signaling molecule. Released HMGB1 mediates diverse responses by binding to several receptors, including the receptor for advanced glycation end products (RAGEs) and toll-like receptors (TLR)-2 and 4, thereby triggering pleiotropic effects, such as cell proliferation, differentiation, death, inflammation, and immunity. Moreover, HMGB1 passively released from dying tumor cells after chemotherapy and radiotherapy or directly secreted from tumor cells promotes regrowth, proliferation and metastasis15,16. It may facilitate autophagy following cytotoxic insults for chemoresistance via its receptor RAGE through the MEK/ERK signaling pathway in colorectal cancer and lung adenocarcinoma17,18,19,20,21. Moreover, the expression of RAGE is closely associated with the invasion and metastasis of gastric cancer and colorectal cancer22,23. The germ-line single-nucleotide polymorphism (SNP) of RAGE with Gly82Ser (rs2070600), which are known to display increased ligand binding to enhance the downstream signaling pathway24,25, is associated with a significantly increased risk of several cancer types26. However, the mechanism and significance of HMGB1-mediated autophagy for chemoresistance remain unknown.
Here, we showed that ERK-mediated Drp1 phosphorylation is necessary for resisting chemotherapeutic cytotoxicity and reported for the first time that this mediation was associated with extracellular HMGB1 released from dying tumor cells. Extracellular HMGB1 promoted ERK activation-induced Drp1 phosphorylation via RAGE, which induced autophagy and acted upon surviving cancer cells to promote regrowth. Furthermore, patients with highly phosphorylated Drp1 proteins are associated with the increased risk on developing tumor relapse after neoadjuvant chemoradiotherapy (neoCRT) treatment in locally advanced rectal cancer (LARC). Moreover, patients with RAGE-G82S polymorphism (rs2070600) are positively associated with phospho-Drp1Ser616 after neoCRT treatment. Altogether, our findings showed that the HMGB1-induced Drp1-dependent mitochondrial dynamics via RAGE-ERK signaling pathway for autophagy and chemoresistance could be candidate therapeutic targets for new treatment strategies in colorectal cancer.
Materials and methods
Cell culture and reagents
The human colorectal cancer cell lines SW480, SW620, LoVo and LoVoOXR were cultivated in RPMI 1640 growth medium supplemented with 10% fetal bovine serum (Invitrogen, California, USA) at 37 °C under a humidified atmosphere of 5% CO2 and 95% air.
To prepare the conditioned medium (CM), the cells were washed and incubated with a serum-free medium for 2 h when sub-confluent. The medium was discarded, and the cells were incubated with a serum-free medium again. After 48 h, the CM was harvested and centrifuged to remove debris, filtered through a 0.22 μm filter, and stored at −20 °C until use.
Antibodies and reagents
The antibodies used in this study were as follows: anti-β-actin (sc-8432, Santa Cruz, CA, USA), anti-ERK1/2 (sc-135900, Santa Cruz, CA, USA), anti-p-ERK1/2 (sc-7383, Santa Cruz, CA, USA), anti-MFN1 (sc-50330, Santa Cruz, CA, USA), anti-RAGE (sc-365154, Santa Cruz, CA, USA and ab89911, Abcam, Cambridge, UK), anti-MFN2 (sc-100560, Santa Cruz, CA, USA), anti-OPA1(sc-393296, Santa Cruz, CA, USA), anti-phospho-Drp1Ser616 (#3455, Cell Signaling Technology, MA, USA), anti-phospho-Drp1 Ser637 (#6319, Cell Signaling Technology, MA, USA), anti-LC3 (#3868, Cell Signaling Technology, MA, USA), anti-HMGB1 (#3935, Cell Signaling Technology, MA, USA and #651401, Biolegend, San Diego, CA, USA), and anti-p62 (#39749, Cell Signaling Technology, MA, USA). All secondary antibodies (HRP-conjugated anti-rabbit, anti-mouse and anti-goat) were purchased from Santa Cruz Biotechnology. The lentivirus carrying shRNAs against RAGE, HMGB1, Drp1, MFN1 and OPA1 were obtained from the National Core Facility for Manipulation of Gene Function by RNAi, miRNA, miRNA sponges, and CRISPR/Genomic Research Center, Academia Sinica. The recombinant HMGB1 proteins, HMGB1 inhibitor quercetin were purchased from Sigma (St. Louis, MO, USA). RAGE inhibitor FPS-ZM1 was purchased from Merck (Temecula, CA, USA).
Western blot analysis
The conditioned medium was collected by ultrafiltration using a Microcon filter (10,000-Da cutoff; Millipore, MA, USA) to purify the secreted protein. Either the total lysates (30 μg) or subcellular fractions (10 μg) were separated via 6–12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto a PVDF membrane (GE Healthcare, Amersham, UK)27. The membranes were blocked with 5% non-fat milk and probed with specific antibodies overnight at 4 °C. Horseradish peroxidase (HRP)-conjugated secondary antibodies (1:10,000, GE Healthcare, Amersham, UK) and the Immobilon Western Chemiluminescent HRP Substrate (Millipore, MA, USA) were used. A densitometric analysis of the western blots was performed using an AlphaImager2200 digital imaging system (Digital Imaging System, CA, USA). The digital images were processed using Adobe Photoshop 7.0. Each blot was stripped using Restore Western Blot Stripping Buffer (Pierce, IO, USA) to incubate with the other antibodies. The results were assessed using Image J software (NIH, MD, USA).
Assessment of cell growth and apoptosis, and mitochondria morphology
The cell growth was assessed using an MTT assay. Apoptosis and necrosis were assessed using the apoptosis/necrosis detection kit (Enzo Life Sciences, Plymouth Meeting, USA). According to the kit, the apoptotic and necrotic cells were labeled fluorescently by annexin V-EnzoGold and 7-AAD, respectively. The cells (2000) were observed to determine apoptosis or necrosis under a fluorescence microscope. The experiments were performed in triplicate. Lactate dehydrogenase C (LDH) and Terminal deoxynucleotidyl transferase (TdT)-mediated dUTP-biotin nick end labeling (TUNEL) were also performed.
Cells were stained with MitoTracker Red (MTR, Life Technologies #M7512). Cells were stained with the 100 nM MTR in completed medium for 30 min at 37 °C, after which the media was replaced for imaging.
Animal model
Athymic nude BALB/c mice (male, 4-weeks-old) were maintained according to the institutional guidelines approved by the China Medical University Institutional Animal Care and Use Committee. The serum HMGB1 level was examined on day 9. LoVo, LoVoOXR, SW480-vector, SW480-Drp1WT, SW480-Drp1S616E (1 × 106 cells/mouse) were suspended in Matrigel matrix with 50% serum-free DMEM, and injected subcutaneously into the flank of each mouse. The tumor volume, body weight and survival time were measured at various time intervals throughout the study. The tumor volumes were calculated according to the formula (width2 × length)/2. The mice were sacrificed at the termination of the experiments, and the serum and tumor tissues from representative mice were collected for lysis, subjected to the immunoblotting analysis and stained by immunohistochemistry.
Enzyme-linked immunosorbent assay (ELISA)
The mouse blood was rapidly centrifuged at 500 × g for 4 min at 4 °C. The supernatant was used as the serum. The HMGB1 in the serum was measured using ELISA kit (Shinotest, Tokyo, Japan, and Abnova, Taipei City, Taiwan). ELISA was performed according to manufacturer’s instructions. The experiment was repeated three times.
Patient selection, clinical staging, treatment, and pathological evaluation
Between 2006 and 2014, patients with locally advanced rectal cancer were treated at the China Medical University Hospital. Patients with biopsy-proven, locally advanced rectal cancer [cT3-4 or cN+ by endorectal ultrasonography (EUS), computed tomography (CT), or magnetic resonance imaging (MRI)] who were treated with preoperative chemoradiotherapy followed by radical resection at the China Medical University Hospital were the study cohorts. The study was reviewed and approved by the Internal Review Board (IRB) of the China Medical University Hospital [Protocol number: CMUH105-REC2-072]. Patients with concurrent distant metastasis or concurrent inflammatory bowel disease, hereditary colorectal cancer syndromes, concurrent malignancy, emergent surgery, prior history of radiotherapy to the pelvis, or prior history of malignancy were excluded. The tumors were staged according to the American Joint Committee on Cancer (AJCC) staging system.
The patients were treated with chemoradiotherapy at a median radiotherapy dose of 50.4 Gy and concurrent fluoropyrimidine-based chemotherapy (mainly single-agent infusional capecitabine). The age ranged from 31 to 90 years. Surgery, including low anterior resection, proctectomy with coloanal reconstruction, abdominoperineal resection, or multivisceral rectal resection using total mesorectal excision principles, was performed 6–8 weeks after the completion of chemoradiotherapy. Adjuvant chemotherapy was recommended for patients with metastatic lymph node(s) in the surgical specimen and consisted of infusional fluorouracil or capecitabine for a period of 4–6 months.
The pathological staging of the resected specimen was performed after resection in accordance with the guidelines of the College of American Pathologists, and the histopathological diagnosis was performed by pathologists specializing in gastrointestinal cancer. A pathological complete response (pCR) was defined as the absence of viable adenocarcinoma cells in the surgical specimens (ypT0N0). The complete response (cCR) classification included specimens with acellular mucin pools without viable tumor cells.
Construction of tissue microarray (TMA)
Tissue microarrays were constructed from biopsies from 106 pair-matched pre-neoCRT and post-neoCRT rectal cancer patients (Table S1). One hundred and six pre-neoCRT biopsies and 93 [13 tumor regression grade 4 (TRG 4)] post-neoCRT surgical specimens were evaluated. The areas of the tumor were marked on hematoxylin & eosin (H & E) stained slides. The corresponding area on the matching paraffin block (donor block) was then identified and marked. We used the AutoTiss 10C system (EverBio Technology, Inc., Taipei, Taiwan) to remove the tissue core from these areas on the donor blocks, which were placed into the recipient block in a precisely arrayed fashion. The diameter of the punches was 2 mm. A maximum of 60 punches were placed on one single block. The sample sections (cut via a microtome) were then mounted on capillary-gap slides (Dako, Hamburg, Germany) and baked overnight28.
Immunohistochemistry
The 3μm-thick TMA slides were individually stained with horseradish peroxidase-conjugated avidin biotin complex (ABC) from the Vectastain Elite ABC Kit (Vector Laboratories, Burlingame, CA) and NovaRed chromogen (Vector Laboratories, Burlingame, CA) and counterstained with hematoxylin. Positive staining for Drp1 and p-Drp1 was detected in the cytoplasm. The Drp1 or p-Drp1 staining patterns were evaluated in the sections according to the proportion of positively stained tumor cells under a microscope (OLYMPUS BX53, Tokyo, Japan). The intensity of the staining was evaluated, and five fields were used in the evaluation29. The pathologists estimated the number of immunopositive cells. The staining intensity was scored as follows: a score of 0 indicated no color; light red was scored as 1; purple-red was scored as 2; and dark-red was scored as 3. The percentage of positive cells was scored as follows: a score of 0 was assigned when the positive tumor cell proportion was 10%; 1 point was assigned when the positive cell proportion ranged from 10 to 25%; 2 points were assigned when the positive cell proportion ranged from 26 to 50%; and 3 points were assigned when the positive cell proportion ranged from 51% to 100. The sum of the two scores served as the staining score. Using score of 3 as a cutoff, the immunostains were defined as “low” for scores from 0 to 3 and as “high” for scores from 4 to 930.
RAGE germ-line single-nucleotide polymorphism (SNP) analysis
Genomic DNA from rectal cancer patients was extracted from the FFPE tissue using a QuickExtract™ FFPE DNA Extraction Kit (Epicenter, WI, USA). For the SNP genotyping, 10 ng to total genomic DNA were used for the PCR amplification using the following specific primers: 5′-RAGE G82S (5′- ATTTGGATCCCCGTCACTCTG -3′) and Biotin-3′- RAGE G82S (5′- GCCTGGCACCGGAAAATC -3′).
The biotinylated PCR products were immobilized into streptavidin-coated super paramagnetic Dynabeads M280-streptavidin. The sequencing primers and RAGE-G82S-Seq (5′-CGTGTCCTTCCCAAC-3′) were annealed to the DNA templates according to the manufacturer’s instructions (PSQk 96MA instrument, Qiagen).
Statistical analysis
The statistical analysis was performed using SAS statistical software, version PC 9.4 (SAS Institute, Cary, NC, USA). A two-sided p-value at a significance level of 0.05 was reported for all tests. The between-group comparisons were performed using Student’s t-test, Pearson chi-square test and Fisher’s exact test. A Cox regression analysis was performed to estimate the hazard ratios (HRs) and 95% confidence intervals (CIs) for the univariate and multivariate models. The influential factors affecting the survival rate of the patients with rectal cancer were adjusted in the Cox models—including sex (female versus male), age (<65 vs. ≥65), TRG (3–4 vs. 1–2), clinical response (good response vs. poor response), and pN stage (positive vs. negative). The Kaplan–Meier estimation method was used to assess the five-year overall survival and disease-free survival. The survival time was defined as the time from surgery to death.
Results
Mitochondrial fission protein Drp1 is hyperactivated upon cytotoxic insults and promotes chemoresistance
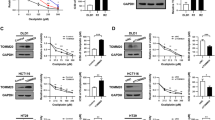
By comparing the mitochondria dynamics between LoVo and oxaliplatin (OXP)-resistant LoVo cells, we found that mitochondria morphology was dramatic changed in LoVoOXR cells (Fig. 1a). We then aim to identify the mitochondrial proteins that are involved in chemoresistance in CRC. By analyzing the LoVoWT and LoVoOXR cell lines, we found that the knockdown of the mitochondrial fusion proteins OPA1 and MFN1 did not sensitize the LoVoOXR cells to oxaliplatin (Fig. 1a). The knockdown efficiency was shown in Fig. S1A. Only knockdown of the mitochondrial fission protein Drp1 sensitized both the LoVoWT and LoVoOxR cells to OXP, suggesting that Drp1 might influence chemosensitivity in CRC (Fig. 1a). We then investigated the Drp1 expression and phosphorylated status following exposure to chemotherapeutic drugs. A strikingly elevated phosphorylation of activated Drp1 (Ser616) was observed following the chemotherapeutic drug OXP treatment (Fig. 1b) But there is no variation on mitochondrial proteins MFN1 and MFN2 upon OXP treatment, suggesting that only Drp 1 may participate in the chemoresistance via phosphorylation status. We then evaluated the phosphorylation status of Drp1 in well-characterized chemoresistant and metastatic colorectal cancer cell lines. By immunoblotting results, we found that Drp1 S616 phosphorylation were also significantly upregulated in LoVoOXR and SW620 cell lines. Moreover, the high levels of extracellular HMGB1 and phosphorylation of ERK1/2 were found in the OXP-resistant LoVoOXR (compared to LoVoWT) and metastatic SW620 (compared to SW480, Fig. 1c).
a LoVoWT and LoVoOXR were infected by lentivirus carrying shRNA against OPA1, Drp1, and MFN1. These cells were treated with oxaliplatin (OXP, 10 μM) for 48 h. The cell viability was analyzed using an MTT assay (n = 3). *p < 0.01 and **p < 0.01. b Mitochondria morphology was observed by MitoTracker Red staining. Scale bar = 10 μm. c LoVoWT cells were treated with diverse concentrations of OXP for 24 h. Cell lysates were analyzed by immunoblotting. Mitochondria morphology was observed by MitoTracker Red staining. Scale bar = 10 μm. d Cell lysate and medium were analyzed by immunoblotting. Quantification of these results is shown (n = 3). *p < 0.05, **p < 0.01 and ***p < 0.001. e SW480 and SW620 cells were treated with OXP (10 μM), CAP (10 μM), or CPT-11 (5 μM) for 24 h. Cell lysates were analyzed by immunoblotting. f LoVo and LoVoOxR cells with shNC and shDrp1 were treated with OXP (10 μM), CAP (10 μM), or CPT-11 (10 μM) for 48 h. Cell death was analyzed using an AnnexinV/PI assay (n = 3). *p < 0.05, **p < 0.01 and, ***p < 0.001. g LoVo and LoVoOxR cells carrying shNC and shDrp1 were treated with OXP (10 μM Oxaliplatin), CAP (10 μM), or CPT-11 (5 μM) for 48 h. The cell viability was analyzed using an MTT assay (n = 3). *p < 0.05 and **p < 0.01
Recent studies have shown that mitochondrial fission via Drp1 significantly promoted cell survival by increasing autophagy, which promoted resistance to apoptosis and induced chemoresistance10,11. The immunoblotting results showed that Drp1 and phospho-Drp1Ser616 were remarkably upregulated following diverse chemotherapeutic drug treatments, such as OXP, capecitabine (CAP, the pro-drug of 5-FU) and irinotecan (CPT-11) (Fig. 1d, Fig. S1B, C). Moreover, the amount of the mitochondrial autophagy adapter protein p62 was decreased and LC3-II was increased in response to chemotherapeutic drugs, implying that the chemotherapeutic drugs may induce autophagy to promote chemoresistance in CRC (Fig. 1d). Anticancer agents OXP, CAP, and CPT-11 also significantly promoted the HMGB1 expression in CRC (Fig. 1d).
To confirm the Drp1 phosphorylation promotes chemoresistance, we silenced Drp1 by lentivirus carrying shRNA against Drp1. Indeed, knockdown of Drp1 increased the chemosensitivity of LoVoOXR cells to chemotherapeutic drugs, promoting the cell death (Fig. 1e) and decreasing cell viability (Fig. 1f). Moreover, treatment with Drp1 inhibitor Mdivi-1 also increased the sensitivity to OXP either LoVoWT or LoVoOXR (Fig. S1D). These results indicated that Drp1-mediated mitochondrial fission notably promoted chemoresistance in CRC cells.
Highly phosphorylated Drp1 may promote chemoresistant tumor growth through HMGB1/RAGE signaling pathway
To verify the role of Drp1 on chemoresistant CRC in vivo, we inoculated athymic nu/nu mice with LoVoWT and LoVoOXR. Tumors derived from LoVoWT grew more slowly compared with those derived from LoVoOXR (Fig. 2a). We observed that the LoVoOXR showed hyperactivated Drp1 levels in vivo (Fig. 2a). Moreover, the immunoblotting results showed that the phosphorylated Drp1Ser616 protein level was highly recruited to the mitochondrial fraction in the tumor derived from LoVoOXR cells (Fig. 2b). These results show that Drp1 is important for modulating chemoresistance via mitochondria fission in CRC cells in vivo.
a Athymic nude BALB/c mice were subcutaneously inoculated with LoVoWT and LoVoOXR cells into the flank of each mouse. (1 × 106 cells/ mice, n = 6). The tumor volume was measured at various time intervals throughout the study. Quantification of these results is shown (n = 6). *p < 0.05. b Xenografted tumors from representative mice were stained for phospho-Drp1Ser616 by immunohistochemistry. Bar: 10 μm. Xenografted tumors from representative mice were homogenized and isolated for the cytoplasmic and mitochondrial fraction analysis. Then, these fractions were analyzed by immunoblotting. Quantification of these results is shown (n = 3). *p < 0.05 and **p < 0.01. c Xenografted tumors from LoVoWT- and LoVoOXR-injected mice were analyzed by immunohistochemistry. Bar: 10 μm. Serum from LoVo-injected and LoVoOxR-injected mice was analyzed by ELISA. Quantification of these results is shown (n = 6). *p < 0.05. d Xenografted tumors from representative mice were collected for lysis and analyzed by immunoblotting. Quantification of these results is shown (n = 3). *p < 0.05 and **p < 0.01. These data were obtained from three independent experiments, and the values represent the means ± S.D
Moreover, immunohistochemistry analysis showed that the level of RAGE and the distribution of HMGB1 in LoVoOXR group was altered. Cytosolic translocation of HMGB1 was significantly increased in LoVoOXR group, suggesting that HMGB1 was translocated from nucleus into cytoplasm, and then secreted into plasma. Consistent with this observation, high serum HMGB1 was detected in LoVoOXR group (Fig. 2c). Moreover, the results of immunohistochemistry also showed that RAGE, the receptor of HMGB1, was greatly upregulated in LoVoOXR group (Fig. 2c). Immunoblotting results from resected tumor tissues also revealed high HMGB1 level, active ERK1/2 and phospho-Drp1S616 in LoVoOXR group (Fig. 2d). We also observed an increased level of LC3-II expression and decrease p62 level in LoVoOXR tumor comparison with control LoVoWT tumor (Fig. 2d). Moreover, the mitochondrial fusion protein MFN2 was not influenced between these two tumors. These results implied the critical role of HMGB1 in modulating the chemosensitivity via ERK-mediated Drp1 CRC in vitro and in vivo.
HMGB1 triggered ERK-mediated Drp1 phosphorylation via RAGE in CRC
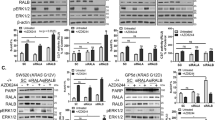
To confirm the relationship between HMGB1 and Drp1 phosphorylation, we treated the CRC cells with recombinant HMGB1 and observe its influence on ERK/Drp1 signaling. We found that ERK and Drp1 were activated by the HMGB1 recombinant protein in both SW480 and LoVo cell lines (Fig. 3a). Inhibition of RAGE by FPS-ZM131 blocked the HMGB1-induced Drp1 phosphorylation and ERK1/2 activation in dose-dependent manner (Fig. 3b). Furthermore, enforced expression of wt-RAGE sensitized the HMGB1-induced Drp1 phosphorylation (Fig. 3c). Similarly, knockdown of RAGE by lentivirus-based shRNA significantly alleviated HMGB1-induced Drp1 phosphorylation and ERK1/2 activation (Fig. 3d).
a SW480 and LoVo cells were treated with the recombinant human HMGB1 protein (0, 1, and 2 μg/mL) for 24 h. Cell lysate was analyzed by immunoblotting. Quantification of these results is shown (n = 3). ***p < 0.001. b SW480 cells were treated with the recombinant human HMGB1 protein (1 μg/mL) and RAGE inhibitor (10 μM FPS-ZM1) for 24 h. Cell lysate was analyzed by immunoblotting. Quantification of these results is shown (n = 3). **p < 0.01 and ***p < 0.001. c SW480 cells were transfected with the pcDNA3.1-vector, pcDNA3.1-wtRAGE for 24 h and then treated with the recombinant human HMGB1 protein (1 μg/mL) for 24 h. Cell lysate was analyzed by immunoblotting. Quantification of these results is shown (n = 3). **p < 0.01. d SW480 cells were silenced with lentivirus carrying shRNA again RAGE, and then treated with the recombinant human HMGB1 protein (1 μg/mL) for 24 h. Cell lysate was analyzed by immunoblotting. e LoVo cells were treated with recombinant human HMGB1 (1 μg/mL) for 24 h. Cell lysate was analyzed by immunoblotting. Quantification of these results is shown (n = 3). ***p < 0.001. Mitochondria morphology was observed by MitoTracker Red staining. Scale bar = 10 μm. f LoVo cells were treated with RAGE inhibitor (10 μM FPS-ZM1), Drp1 inhibitor (50 μM Mdivi-1) and recombinant human HMGB1 (1 μg/mL) for 1 h prior to addition of different dosages of OXP for 48 h. The cell viability was analyzed using an MTT assay (n = 3). *p < 0.05. g LoVo cells were treated with RAGE inhibitor (10 μM FPS-ZM1), Drp1 inhibitor (50 μM Mdivi-1) and recombinant human HMGB1 (1 μg/mL) for 1 h prior to addition of OXP (10 μM) for 48 h. The caspase-3 activity was analyzed (n = 3). **p < 0.01. These data were obtained from three independent experiments, and the values represent the means ± S.D
Furthermore, addition of HMGB1 dramatically influences the mitochondria morphology and increases conversion of LC3-I to LC3-II and decreases p62 level (Fig. 3e). Treatment with Drp1 inhibitor Mdivi-1, which is a small molecular inhibitor of Drp132, mitigates HMGB1-induced Drp1 phosphorylation, mitochondria morphology alteration, LC3 conversion and p62 levels decrease.
To identify whether HMGB1 was responsible for chemoresistance via Drp1 phosphorylation, we pre-treated with recombinant HMGB1 for 1 h prior to addition of OXP (Fig. 3f). Pre-incubation of HMGB1 significantly alleviated OXP-induced cell viability decrease. But blockade of RAGE by FPS-ZM1 or inhibition of Drp1 by Mdivi-1 neutralized the OXP-induced cell death. Addition of HMGB1 prior to OXP treatment also reduced the OXP-induced caspase-3 activity. However, the caspase-3 activity was decreased by FPS-ZM1 and Mdivi-1, suggesting that HMGB1 is required for chemoresistance to OXP via Drp1. Taken together, these results indicated that the extracellular HMGB1 promotes ERK activation-mediated Drp1 phosphorylation resulting in tumor growth and chemoresistance via RAGE.
Secretion of HMGB1 upon cytotoxic insults promotes Drp1 phosphorylation for chemoresistance via autophagy
To verify the HMGB1 secreted from dying cells can promote Drp1 phosphorylation, we challenges with diverse chemotherapeutic drugs and collected the medium for analysis. Indeed, extracellular HMGB1 secreted from dying tumor cells was detected in the medium following treatment with these anticancer agents (Fig. 4a). We then silenced HMGB1 using a lentivirus carrying shRNA against HMGB1 to observe the impact on Drp1 phosphorylation. As expected, the knockdown of HMGB1 clearly decreased phosphor-Drp1Ser616 (Fig. 4b). The silencing of HMGB1 also enhanced the sensitivity of colorectal cancer cells to OXP (Fig. 4b). Notably, the inhibition of HMGB1 secretion by quercetin33,34 significantly suppressed the CAP-induced ERK activation and Drp1 phosphorylation (Fig. 4c). The decrease in p62 and increase in LC3-II were also attenuated by quercetin, suggesting that CAP-induced autophagy was decreased as HMGB1/RAGE signaling was blocked. Moreover, blockade of MEK/ERK by MEK inhibitor (PD98059) significantly decreased Drp1 phosphorylation (Fig. 4d). These results suggested that the high level of HMGB1 secreted from dying and resistant cancer cells during chemotherapy may trigger Drp1 phosphorylation via RAGE/ERK.
a SW480 and SW620 cells were treated with OXP (10 μM), CAP (10 μM), or CPT-11 (5 μM) for 24 h. Conditioned medium were analyzed by immunoblotting. Quantification of these results is shown (n = 3). *p < 0.05, **p < 0.01 and ***p < 0.001. b SW480 cells carrying shNC, shHMGB1#1 and shHMGB1#2 were treated with various concentration of oxaliplatin for 48 h. The cell viability was analyzed using an MTT assay (n = 3). **p < 0.01. c SW480 cells were treated with CAP (10 μM) and HMGB1 inhibitor (Quercetin, 50 μM) for 24 h. Cell lysate was analyzed by immunoblotting. Quantification of these results is shown (n = 3). **p < 0.01 and ***p < 0.001. d SW480 cells were treated with OXP (10 μM) and PD98059 (10 μM) for 24 h. Cell lysate was analyzed by immunoblotting. e SW480 cells were transfected with the pcDNA3.1-vector, pcDNA3.1-wtRAGE and pcDNA3.1-RAGEΔCyto for 24 h and then treated with OXP (10 μM), CAP (10 μM), or CPT-11 (5 μM) for 48 h. The cell viability was analyzed using an MTT assay (n = 3). *p < 0.05. f SW480 cells carrying shNC and shRAGE were treated with OXP (10 μM Oxaliplatin), CAP (10 μM), or CPT-11 (5 μM) for 48 h. The caspase-3 activity was analyzed (n = 3). **p < 0.01. g SW480 cells carrying shNC and shRAGE were treated with OXP (10 μM), CAP (10 μM), or CPT-11 (5 μM) for 48 h. Cell death was analyzed using an AnnexinV/PI assay (n = 3). *p < 0.05 and ***p < 0.001. h SW480 cells were treated with CAP (10 μM) and RAGE inhibitor (FPS-ZM1, 10 μM) for 24 h. Cell lysate was analyzed by immunoblotting. Quantification of these results is shown (n = 3). **p < 0.01 and ***p < 0.001. i LoVo cells were pre-incubated with anti-RAGE or anti-HMGB1 neutralizing antibodies (2.5 μg/mL) for 30 min prior to addition of recombinant HMGB1 and OXP for 48 h. The cell viability was analyzed using an MTT assay (n = 3). *p < 0.05. These data were obtained from three independent experiments, and the values represent the means ± S.D
Moreover, overexpression of the wild-type RAGE increased the cell viability following the anticancer agent treatment (Fig. 4e). The dominant-negative of RAGE (RAGE-ΔCyto) had no influence on the cell viability following the cytotoxic insults (Fig. 4e). Consistent with these results, knockdown of RAGE increased caspase-3 activity (Fig. 4f) and cell death (Fig. 4g) following the chemotherapeutic drug treatments. Notably, the blockage of RAGE by the pharmacological inhibitor FPS-ZM1 also significantly suppressed the CAP-induced ERK activation and Drp1 phosphorylation (Fig. 4h). The decrease in p62 and increase in LC3-II were also attenuated by FPS-ZM1. Moreover, pretreatment with anti-RAGE or anti-HMGB1 neutralizing antibodies for 30 min prior to addition of recombinant HMGB1 and OXP significantly mitigated HMGB1-induced chemoresistance (Fig. 4i). Taken together, these results indicated that secretion of HMGB1 from dying tumor cells promotes ERK-mediated Drp1 phosphorylation, which contributes to autophagy and chemoresistance following chemotherapeutic treatment.
Inhibition of Drp1 alleviated chemotherapeutic drugs-induced autophagy and chemoresistance in vitro and in vivo
Consistent with previous results, we found that Drp1 inhibitor Mdivi-1 significantly alleviated the CAP-induced Drp1 phosphorylation, conversion of LC3 I to LC3II and decrease in p62 (Fig. 5a). To determine Drp1 phosphorylation is critical for drug resistance in tumor cells, we constructed a gain-of-function Drp1 containing both S616E (to mimic activating phosphorylation) and S637A (to block inhibitory phosphorylation) mutations. Enforced expression of Drp1S616E accelerated the CAP-induced p62 decrease and LC3-II conversion (Fig. 5b). These results demonstrate that hyperactivated Drp1 is essential for chemoresistance via mitochondrial fission and autophagy.
a SW480 and SW620 cells were treated with CAP (10 μM) and a Drp1 inhibitor (50 μM Mdivi-1) for 24 h. Cell lysate were analyzed by immunoblotting. b SW480 cells were transfected with a GFP vector, GFP-wtDrp1 or GFP-Drp1S616E for 24 h and then treated with CAP (10 μM Capecitabine) for 24 h. Cell lysate was analyzed by immunoblotting. Quantification of these results is shown (n = 3). *p < 0.05, **p < 0.01, and ***p < 0.001. c 1 × 106 cells were injected subcutaneously into the flank of each mouse. Tumor volume, body weight and survival time were measured at various time intervals throughout the study. Quantification of these results is shown (n = 5). **p < 0.01. d Xenografted tumors from representative mice were collected for lysis and analyzed by immunoblotting. Quantification of these results is shown (n = 3). *p < 0.05 and **p < 0.01. These data were obtained from three independent experiments, and the values represent the means ± S.D
To further explore the effect of the mitochondrial dynamics on cell survival in CRC cells, we inoculated athymic nude BALB/c mice with SW480 tumor cells that stably expressed hyperactivated Drp1S616E or wild-type Drp1. The growth of the Drp1S616E tumor cells was significantly faster than that of the wild-type Drp1–transfected tumors (Fig. 5c). We observed that increased autophagy in the resected tumor transfected with wild-type Drp1 and Drp1S616E in vivo (Fig. 5d). The level of p62 was dramatically decreased and the ratio of LC3-II/I was increased. But the Drp1S616E-transfected tumor cells displayed more autophagy (Fig. 5d). These results showed that Drp1-mediated mitochondrial fission notably promoted autophagy and chemoresistance in CRC cells.
Hyperactivated Drp-1 within tumor microenvironment is associated with poor 5-years disease-free survival (DFS) and 5-year overall survival in LARC patients
To determine the clinical relevance of these findings, we performed a combination of tissue analysis studies. We performed an immunohistochemistry analysis of Drp1 and phospho-Drp1Ser616 on 106 pair-matched pre-neoCRT biopsies and post-neoCRT surgical tissues in LARC patients using a tissue microarray (TMA, Fig. 6a). These patients were received the preoperative neoadjuvant chemoradiotherapy (neoCRT). The median dose of radiation was 50.4 Gy in 28 fractions. The concurrent chemotherapy was fluorouracil-based in 42% of the patients and capecitabine (CAP) in 58% of the patients. After neoCRT treatment, 12% (13/106) of the patients had a complete response, while 82% (93/106) of the patients had a partial response based on the tumor regression grade (TRG). The clinicopathological characteristics are presented in Table S1.
a The expression of phospho-Drp1S616 in the pre-neoCRT biopsies and metastatic lymph nodes. b High phospho-Drp1S616 within the tumor microenvironment of pre-neoCRT biopsies was associated with a poor 5-year DFS. c High phospho-Drp1S616 within the tumor microenvironment of pre-neoCRT biopsies was associated with a poor 5-year OS. d High phospho-Drp1S616 within the tumor microenvironment of post-neoCRT surgical tissues was associated with a poor 5-year DFS and 5-year OS. e High phospho-Drp1S616 within the tumor microenvironment of post-neoCRT surgical tissues was associated with a poor 5-year OS. f The proposed mechanism of HMGB1-mediated Drp1 phosphorylation via RAGE-ERK1/2 signaling to promote chemoresistance in colorectal cancer
Patients with high level of Drp1 have no significant correlation with these parameters. But patients with high level of phospho-Drp1Ser616 had high risk on tumor relapse within 5 years (p = 0.0073) (Table S1). Moreover, patients with high phospho-Drp1Ser616 showed increased risk on poor 5-years disease-free survival (DFS, p = 0.01, Fig. 6b) and 5-years overall survival (OS, p = 0.01, Fig. 6c) in LARC before neoCRT treatment (Table 1).
Furthermore, in the post-neoCRT surgical specimens (n = 93), we found that the patients with the high phospho-Drp1Ser616 still showed high risk on poor DFS (p = 0.0003, Fig. 6d) and 5-year OS (p = 0.002, Fig. 6e) in LARC. These results indicated that phospho-Drp1Ser616 was significantly associated with the risk of developing tumor relapse after neoCRT treatment, suggesting that phospho-Drp1Ser616 contributes to treatment resistance and metastasis in LARC.
Prognostic relevance of hyperactivated Drp-1 in LARC patients receiving neoadjuvant chemoradiotherapy (neoCRT) treatment
In the univariate analysis, the patients with high level of phospho-Drp1Ser616 had an increased risk on poor 5-year OS (HR = 3.164, 95% CI 1.238–8.089, p = 0.016, Table 2). In the multivariate analysis, the patients with high phospho-Drp1Ser616 in the tumor microenvironment had an increased risk on poor 5-year OS (HR = 3.821, 95% CI 1.459–10.008, p = 0.006, Table 2). These results indicated that phospho-Drp1Ser616 is a significant prognostic factor for survival outcome in LARC patients who treated with neoCRT. Moreover, after neoCRT treatment, we found that the high phospho-Drp1Ser616 was significantly observed in the metastatic lymph nodes (Fig. 6a), and associated with tumor relapse (Table S2).
Furthermore, we found a substantial proportion of RAGE-G82S polymorphism (36/106 [34%]) in patients with LARC. Patients with RAGE-G82S polymorphism did not correlated with high phospho-Drp1 Ser616 before neoCRT treatment (p = 0.84, Table 3). But patients with RAGE-G82S polymorphism displayed high phospho-Drp1 Ser616 after neoCRT treatment (p = 0.05, Table 3), suggesting that patients with RAGE-G82S polymorphisms may have increased Drp1 activation to promote chemoresistance and tumor regrowth after chemoradiotherapy treatment, leading to poor survival outcome in LARC. Taken together, these results demonstrated that HMGB1 released from dying cancer cells after neoCRT may enhance regrowth and metastasis of remnant cancer cells via RAGE-ERK-Drp1 activation.
Discussion
Most CRC patients develop resistance to chemotherapy drugs, such as oxaliplatin, which is a significant challenge in the treatment of colon cancer. This phenomenon has been attributed to different mechanisms, including dysfunctional membrane transport, resistance to apoptosis, and DNA repair mechanisms. In this study, we showed that Drp1-mediated mitochondria fission, which results in autophagy, is a significant contributor to drug resistance in CRC. Following chemotherapy treatment, the release of HMGB1 is induced, and HMGB1 can exert autocrine or paracrine effects that activate Drp1 and lead to chemoresistance and tumor recurrence (Fig. 6f). Moreover, the inhibition of mitochondria fission via Drp1, the blockage of the release of HMGB1 or the blockage of the HMGB1 receptor RAGE enhance the sensitivity of anti-cancer agents in CRC.
Cancer cells respond to chemotherapy in a variety of ways, ranging from the activation of survival pathways to the initiation of cell death. Consistent with previous studies that identified a cytoprotective role of autophagy, increased autophagy is observed following exposure to chemotherapy drugs35,36. Here, we reported that mitochondria fission by Drp1 hyperactivation is required for autophagy and chemoresistance. Because Drp1 has been shown to be involved in several cellular processes, such as mitochondrial energetics to sustain tumor development, stem cell maintenance, promotion of metastasis and cell proliferation, Drp1-driven mitochondrial fission has been considered a vital factor in the context of tumorigenesis. Several emerging studies have focused on the relationship between mitochondrial dynamics and cell fate determination12,13,37,38. Serasinghe et al. reported that ERK-mediated Drp1 S616 phosphorylation in mitochondrial division was necessary for RAS-induced transformation. The disruption of Drp1 activity was shown to promote mitochondrial fusion and inhibit tumor growth13. Moreover, recent studies have shown that mitochondrial fission via Drp1 significantly promoted cell survival by increasing autophagy, resulting in a resistance to apoptosis in HCC10 and chemoresistance in acute lymphoblastic leukemia (ALL)11. Cai et al. revealed that the remarkable elevation of the activating phosphorylation of Drp1 at S616 decreased the mitochondrial ROS levels, resulted in a pro-glycolytic metabolism shift and increased drug resistance11. Collectively, these results and our present findings indicate that Drp1-mediated mitochondrial dynamics are associated with increasing autophagy, resulting in resistance to cell death and chemoresistance.
HMGB1 has been reported as a DAMP for immunogenic cell death (ICD) during chemotherapy and radiotherapy, and its release from necrotic cancer cells may activate the innate immune system to promote host anticancer immunity. However, extracellular HMGB1 is known to act as a growth factor that activates mitogen-activated protein kinase (MAPK) and thereby enhances the regrowth, metastasis, and chemoresistance of remnant cancer cells that survived prior chemotherapy via RAGE15,19,39,40,41,42,43. Thus, the role of chemotherapy-associated HMGB1 is thought to depend on the balance of these different effects of HMGB1. Indeed, the knockdown of RAGE or HMGB1 increased cell death and restored the chemosensitivity of the cancer cells in vivo and in vitro21, indicating that this this knockdown may lead to the induction of cancer progression and drug resistance. The ability of released HMGB1 to promote chemoresistance in cancer cells, such as neuroblastoma44, lung adenocarcinoma18, prostate cancer45, CRC17, and leukemia46, is reportedly due to autophagy47. Consistently, we found that OXP, CAP, and CPT-11 promote the enhanced expression and secretion of HMGB1 in human colorectal cancer cells. HMGB1 has been reported to be a critical regulator of autophagy. Cytosolic HMGB1 is an inducer of starvation-mediated autophagy by binding to Beclin 1 in colorectal cancer48. In addition, extracellular HMGB1 induces autophagy via its receptor RAGE in colorectal and pancreatic cancer cells21,49. HMGB1-mediated autophagy modulates the sensitivity of colorectal cancer cells to Oxaliplatin via the MEK/ERK signaling pathway17. This hypothesis is supported by our findings in which HMGB1 was associated with autophagy and chemoresistance via RAGE/MEK/ERK/Drp1 in colorectal cancer. Moreover, we found that the expression of Drp1 and p-Drp1 was positively correlated with patients carrying germ-line single-nucleotide polymorphisms (SNPs) of RAGE with G82S (rs2070600) after neoCRT treatment. RAGE-G82S has been reported to exhibit potentially enhanced ligand-binding affinity, such as to HMGB124,50, which supports our finding in which HMGB1 triggered Drp1 phosphorylation via RAGE.
Our results suggested that patients with RAGE-G82S polymorphisms may have high affinity for the ligand, such as HMGB1, which activates ERK1/2 to phosphorylate Drp1 for mitochondria fission and autophagy. Eventually, Drp1-mediated mitochondria fission promotes chemoresistance and tumor regrowth, leading to poor survival outcome in LARC.
The data reported here indicate the potential of extracellular HMGB1 released from colorectal cancer cells to exert a paracrine effect on surviving cancer cells enabling them to resist chemoradiotherapy. Targeting the receptor RAGE and the downstream effector Drp1 in cancer cells may prevent or inhibit the development of drug resistance. In contrast, there is evidence that chemotherapeutic drug-induced HMGB1 can mediate the activation of innate immunity and tumor clearance. Thus, caution must be exercised because of the potential positive and negative aspects of HMGB1 expression during different phases of tumor development and treatment.
These results showed that the potential of extracellular HMGB1 released from colorectal cancer cells to exert a paracrine effect on surviving cancer cells enabling them to resist chemoradiotherapy via RAGE-ERK-Drp1 signaling pathway.
References
Siegel, R. L. et al. Colorectal cancer statistics, 2017. Cancer J. Clin. 67, 177–193 (2017).
Adlard, J. W., Richman, S. D., Seymour, M. T. & Quirke, P. Prediction of the response of colorectal cancer to systemic therapy. Lancet Oncol. 3, 75–82 (2002).
Goldberg, R. M. et al. A randomized controlled trial of fluorouracil plus leucovorin, irinotecan, and oxaliplatin combinations in patients with previously untreated metastatic colorectal cancer. J. Clin. Oncol. 22, 23–30 (2004).
Wang, K. C., Brooks, D. A., Botting, K. J. & Morrison, J. L. IGF-2R-mediated signaling results in hypertrophy of cultured cardiomyocytes from fetal sheep. Biol. Reprod. 86, 183 (2012).
Zhao, J. et al. Mitochondrial dynamics regulates migration and invasion of breast cancer cells. Oncogene 32, 4814–4824 (2013).
Grandemange, S., Herzig, S. & Martinou, J. C. Mitochondrial dynamics and cancer. Semin. Cancer Biol. 19, 50–56 (2009).
Rehman, J. et al. Inhibition of mitochondrial fission prevents cell cycle progression in lung cancer. Faseb. J. 26, 2175–2186 (2012).
Jin, B. et al. Anti-tumour efficacy of mitofusin-2 in urinary bladder carcinoma. Med. Oncol. 28(Suppl 1), S373–S380 (2011).
Inoue-Yamauchi, A. & Oda, H. Depletion of mitochondrial fission factor DRP1 causes increased apoptosis in human colon cancer cells. Biochem. Biophys. Res. Commun. 421, 81–85 (2012).
Huang, Q. et al. Increased mitochondrial fission promotes autophagy and hepatocellular carcinoma cell survival through the ROS-modulated coordinated regulation of the NFKB and TP53 pathways. Autophagy 12, 999–1014 (2016).
Cai, J. et al. ERK/Drp1-dependent mitochondrial fission is involved in the MSC-induced drug resistance of T-cell acute lymphoblastic leukemia cells. Cell Death Dis. 7, e2459 (2016).
Prieto, J. et al. Early ERK1/2 activation promotes DRP1-dependent mitochondrial fission necessary for cell reprogramming. Nat. Commun. 7, 11124 (2016).
Kashatus, J. A. et al. Erk2 phosphorylation of Drp1 promotes mitochondrial fission and MAPK-driven tumor growth. Mol. Cell 57, 537–551 (2015).
Fu, L. et al. SIRT4 inhibits malignancy progression of NSCLCs, through mitochondrial dynamics mediated by the ERK-Drp1 pathway. Oncogene 36, 2724–2736 (2017).
Luo, Y. et al. High mobility group box 1 released from necrotic cells enhances regrowth and metastasis of cancer cells that have survived chemotherapy. Eur. J. Cancer 49, 741–751 (2013).
Lee, H. et al. Diagnostic significance of serum HMGB1 in colorectal carcinomas. PLoS One 7, e34318 (2012).
Liu, W. et al. HMGB1-mediated autophagy modulates sensitivity of colorectal cancer cells to oxaliplatin via MEK/ERK signaling pathway. Cancer Biol. Ther. 16, 511–517 (2015).
Pan, B. et al. HMGB1-mediated autophagy promotes docetaxel resistance in human lung adenocarcinoma. Mol. Cancer 13, 165 (2014).
Amornsupak, K. et al. Cancer-associated fibroblasts induce high mobility group box 1 and contribute to resistance to doxorubicin in breast cancer cells. BMC Cancer 14, 955 (2014).
Zhang, R. et al. Interference with HMGB1 increases the sensitivity to chemotherapy drugs by inhibiting HMGB1-mediated cell autophagy and inducing cell apoptosis. Tumour Biol. 36, 8585–8592 (2015).
Huang, J. et al. HMGB1 promotes drug resistance in osteosarcoma. Cancer Res. 72, 230–238 (2012).
Kuniyasu, H. et al. Expression of receptors for advanced glycation end-products (RAGE) is closely associated with the invasive and metastatic activity of gastric cancer. J. Pathol. 196, 163–170 (2002).
Sasahira, T., Akama, Y., Fujii, K. & Kuniyasu, H. Expression of receptor for advanced glycation end products and HMGB1/amphoterin in colorectal adenomas. Virchows. Arch. 446, 411–415 (2005).
Hofmann, M. A. et al. RAGE and arthritis: the G82S polymorphism amplifies the inflammatory response. Genes Immun. 3, 123–135 (2002).
Huttunen, H. J. & Rauvala, H. Amphoterin as an extracellular regulator of cell motility: from discovery to disease. J. Intern. Med. 255, 351–366 (2004).
Xia, W. et al. Association of RAGE polymorphisms and cancer risk: a meta-analysis of 27 studies. Med. Oncol. 32, 442 (2015).
Lin, T. Y., Fan, C. W., Maa, M. C. & Leu, T. H. Lipopolysaccharide-promoted proliferation of Caco-2 cells is mediated by c-Src induction and ERK activation. Biomedicine 5, 5 (2015).
Chao, K. S. C. et al. Acquired immunity trumps ypN+ and TRG as the sole prognostic biomarker for locally advanced rectal cancer (LARC) treated with neoadjuvant chemoradiation therapy (NeoCRT). Int. J. Radiat. Oncol. Biol. Phys. 96, S108 (2016).
Wang, X. et al. RSF-1 overexpression determines cancer progression and drug resistance in cervical cancer. Biomedicine 8, 4 (2018).
Huang, C. Y. et al. Cytosolic high-mobility group box protein 1 (HMGB1) and/or PD-1 + TILs in the tumor microenvironment may be contributing prognostic biomarkers for patients with locally advanced rectal cancer who have undergone neoadjuvant chemoradiotherapy. Cancer Immunol. Immunother. 67, 551–562 (2018).
Zhan, D., Guo, L. & Zheng, L. Inhibition of the receptor for advanced glycation promotes proliferation and repair of human periodontal ligament fibroblasts in response to high glucose via the NF-kappaB signaling pathway. Arch. Oral Biol. 87, 86–93 (2017).
Tanaka, A. & Youle, R. J. A chemical inhibitor of DRP1 uncouples mitochondrial fission and apoptosis. Mol. Cell 29, 409–410 (2008).
Li, X. et al. Quercetin protects mice from ConA-induced hepatitis by inhibiting HMGB1-TLR expression and down-regulating the nuclear factor kappa B pathway. Inflammation 39, 96–106 (2016).
Tang, D. et al. Quercetin prevents LPS-induced high-mobility group box 1 release and proinflammatory function. Am. J. Respir. Cell Mol. Biol. 41, 651–660 (2009).
Kondo, Y., Kanzawa, T., Sawaya, R. & Kondo, S. The role of autophagy in cancer development and response to therapy. Nat. Rev. Cancer 5, 726–734 (2005).
Amaravadi, R. K. et al. Principles and current strategies for targeting autophagy for cancer treatment. Clin. Cancer Res. 17, 654–666 (2011).
Wieder, S. Y. et al. Activation of the mitochondrial fragmentation protein DRP1 correlates with BRAF(V600E) melanoma. J. Invest. Dermatol. 135, 2544–2547 (2015).
Serasinghe, M. N. et al. Mitochondrial division is requisite to RAS-induced transformation and targeted by oncogenic MAPK pathway inhibitors. Mol. Cell 57, 521–536 (2015).
Czura, C. J., Wang, H. & Tracey, K. J. Dual roles for HMGB1: DNA binding and cytokine. J. Endotoxin Res. 7, 315–321 (2001).
Bhawal, U. K. et al. Association of expression of receptor for advanced glycation end products and invasive activity of oral squamous cell carcinoma. Oncology 69, 246–255 (2005).
Taguchi, A. et al. Blockade of RAGE-amphoterin signalling suppresses tumour growth and metastases. Nature 405, 354–360 (2000).
Kuniyasu, H., Chihara, Y. & Kondo, H. Differential effects between amphoterin and advanced glycation end products on colon cancer cells. Int. J. Cancer 104, 722–727 (2003).
Kuniyasu, H., Chihara, Y. & Takahashi, T. Co-expression of receptor for advanced glycation end products and the ligand amphoterin associates closely with metastasis of colorectal cancer. Oncol. Rep. 10, 445–448 (2003).
Wang, L., Zhang, H., Sun, M., Yin, Z. & Qian, J. High mobility group box 1-mediated autophagy promotes neuroblastoma cell chemoresistance. Oncol. Rep. 34, 2969–2976 (2015).
Zhou, J. et al. HMGB1 induction of clusterin creates a chemoresistant niche in human prostate tumor cells. Sci. Rep. 5, 15085 (2015).
Liu, L. et al. HMGB1-induced autophagy promotes chemotherapy resistance in leukemia cells. Leukemia 25, 23–31 (2011).
Liu, L. et al. DAMP-mediated autophagy contributes to drug resistance. Autophagy 7, 112–114 (2011).
Tang, D. et al. Endogenous HMGB1 regulates autophagy. J. Cell. Biol. 190, 881–892 (2010).
Tang, D. et al. HMGB1 release and redox regulates autophagy and apoptosis in cancer cells. Oncogene 29, 5299–5310 (2010).
Osawa, M. et al. De-N-glycosylation or G82S mutation of RAGE sensitizes its interaction with advanced glycation endproducts. Biochim. Biophys. Acta 1770, 1468–1474 (2007).
Acknowledgements
The experiments and data analysis were performed using the Medical Research Core Facilities Center, Office of Research & Development at China Medical University, Taichung, Taiwan. We are grateful for the tissue microarray (TMA) support from the Translation Research Core, China Medical University Hospital. The lentivirus carrying shRNAs were obtained from the National Core Facility for Manipulation of Gene Function by RNAi, miRNA, miRNA sponges, and CRISPR/Genomic Research Center, Academia Sinica and supported by the National Core Facility Program for Biotechnology Grants of MOST (MOST 104-2319-B-001-001-). This work was supported by grants from the Ministry of Health and Welfare, Taiwan (MOHW107-TDU-B-212-123004), Health and welfare surcharge of tobacco products, China Medical University Hospital Cancer Research Center of Excellence (MOHW107-TDU-B-212-114024, Taiwan) and China Medical University Hospital, Taiwan (DMR-107-126).
Author information
Authors and Affiliations
Contributions
C.-Y.H. and S.-F.C. conducted and performed the experiments; W.T.-L.C., T.-W.K. and T.-W.C. participated in the collection and IHC evaluation of the LARC patients; Y.-S.Y. performed the statistical analysis; S.-F.C., C.-Y.H., William T.-L.C. and K.S.C.C. supervised this study; C.-Y.H., S.-F.C. and K.S.C.C. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Q. Chen
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, CY., Chiang, SF., Chen, W.TL. et al. HMGB1 promotes ERK-mediated mitochondrial Drp1 phosphorylation for chemoresistance through RAGE in colorectal cancer. Cell Death Dis 9, 1004 (2018). https://doi.org/10.1038/s41419-018-1019-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-1019-6
This article is cited by
-
Mitochondrial dynamics and colorectal cancer biology: mechanisms and potential targets
Cell Communication and Signaling (2024)
-
Mitochondria: a new intervention target for tumor invasion and metastasis
Molecular Medicine (2024)
-
Neoantigen-augmented iPSC cancer vaccine combined with radiotherapy promotes antitumor immunity in poorly immunogenic cancers
npj Vaccines (2024)
-
Mitochondrial function and gastrointestinal diseases
Nature Reviews Gastroenterology & Hepatology (2024)
-
Drp1: Focus on Diseases Triggered by the Mitochondrial Pathway
Cell Biochemistry and Biophysics (2024)